Xiyu Ding
ProtoBERT-LoRA: Parameter-Efficient Prototypical Finetuning for Immunotherapy Study Identification
Mar 26, 2025Abstract:Identifying immune checkpoint inhibitor (ICI) studies in genomic repositories like Gene Expression Omnibus (GEO) is vital for cancer research yet remains challenging due to semantic ambiguity, extreme class imbalance, and limited labeled data in low-resource settings. We present ProtoBERT-LoRA, a hybrid framework that combines PubMedBERT with prototypical networks and Low-Rank Adaptation (LoRA) for efficient fine-tuning. The model enforces class-separable embeddings via episodic prototype training while preserving biomedical domain knowledge. Our dataset was divided as: Training (20 positive, 20 negative), Prototype Set (10 positive, 10 negative), Validation (20 positive, 200 negative), and Test (71 positive, 765 negative). Evaluated on test dataset, ProtoBERT-LoRA achieved F1-score of 0.624 (precision: 0.481, recall: 0.887), outperforming the rule-based system, machine learning baselines and finetuned PubMedBERT. Application to 44,287 unlabeled studies reduced manual review efforts by 82%. Ablation studies confirmed that combining prototypes with LoRA improved performance by 29% over stand-alone LoRA.
Classifying Long Clinical Documents with Pre-trained Transformers
May 14, 2021

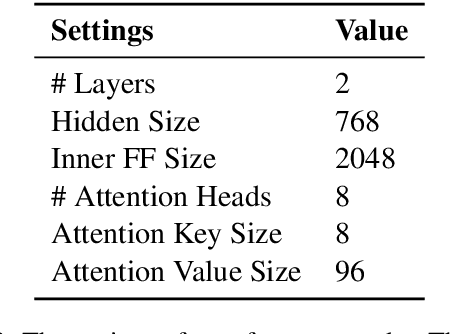

Abstract:Automatic phenotyping is a task of identifying cohorts of patients that match a predefined set of criteria. Phenotyping typically involves classifying long clinical documents that contain thousands of tokens. At the same time, recent state-of-art transformer-based pre-trained language models limit the input to a few hundred tokens (e.g. 512 tokens for BERT). We evaluate several strategies for incorporating pre-trained sentence encoders into document-level representations of clinical text, and find that hierarchical transformers without pre-training are competitive with task pre-trained models.
A machine learning methodology for real-time forecasting of the 2019-2020 COVID-19 outbreak using Internet searches, news alerts, and estimates from mechanistic models
Apr 08, 2020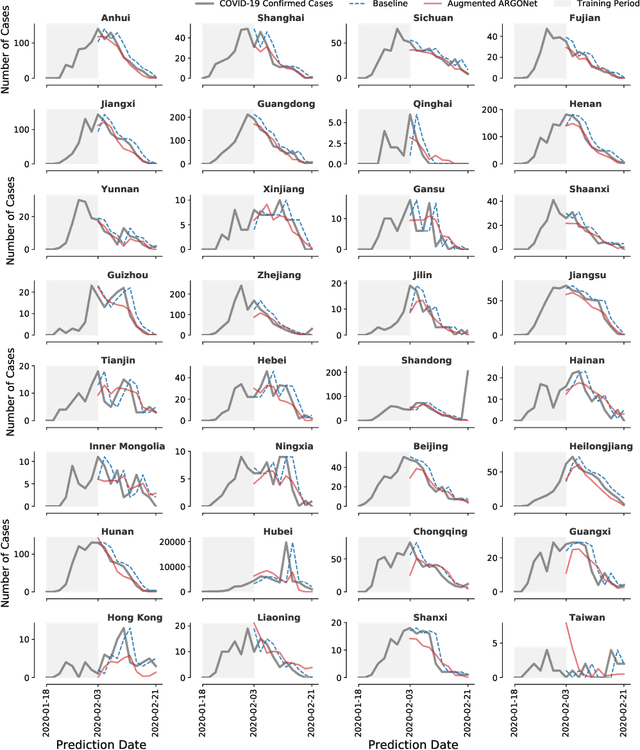
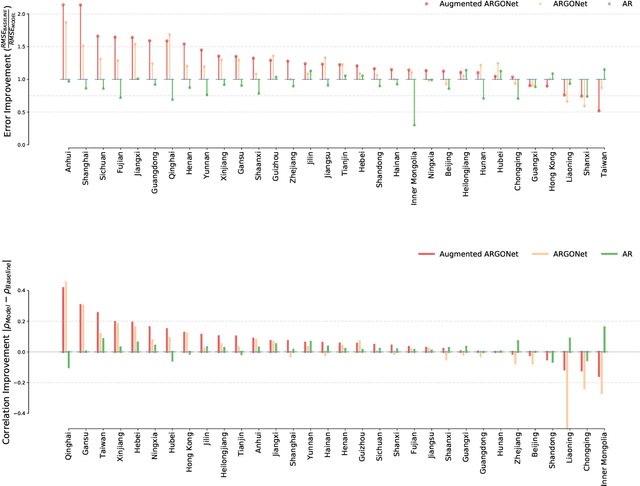
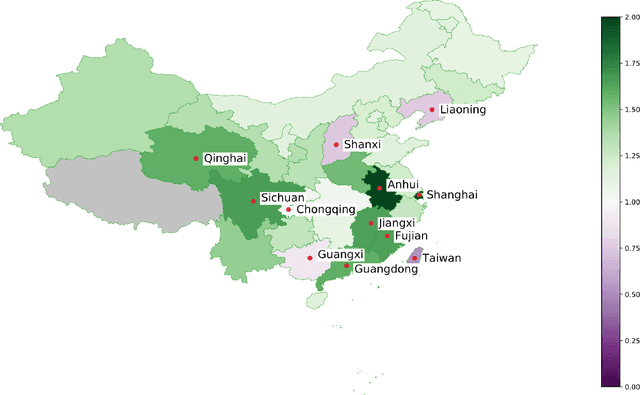
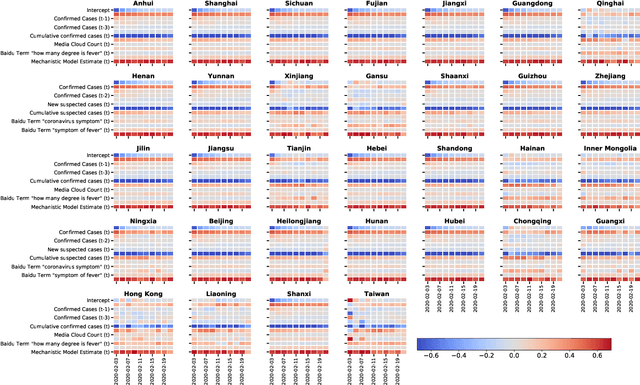
Abstract:We present a timely and novel methodology that combines disease estimates from mechanistic models with digital traces, via interpretable machine-learning methodologies, to reliably forecast COVID-19 activity in Chinese provinces in real-time. Specifically, our method is able to produce stable and accurate forecasts 2 days ahead of current time, and uses as inputs (a) official health reports from Chinese Center Disease for Control and Prevention (China CDC), (b) COVID-19-related internet search activity from Baidu, (c) news media activity reported by Media Cloud, and (d) daily forecasts of COVID-19 activity from GLEAM, an agent-based mechanistic model. Our machine-learning methodology uses a clustering technique that enables the exploitation of geo-spatial synchronicities of COVID-19 activity across Chinese provinces, and a data augmentation technique to deal with the small number of historical disease activity observations, characteristic of emerging outbreaks. Our model's predictive power outperforms a collection of baseline models in 27 out of the 32 Chinese provinces, and could be easily extended to other geographies currently affected by the COVID-19 outbreak to help decision makers.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge