Pierre Decazes
Deep evidential fusion with uncertainty quantification and contextual discounting for multimodal medical image segmentation
Sep 12, 2023
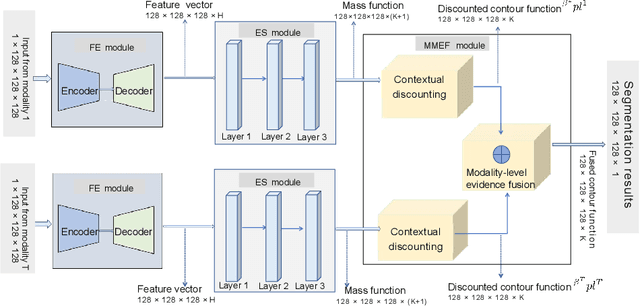
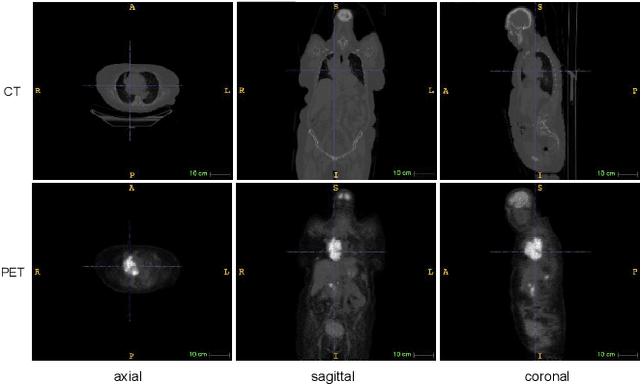
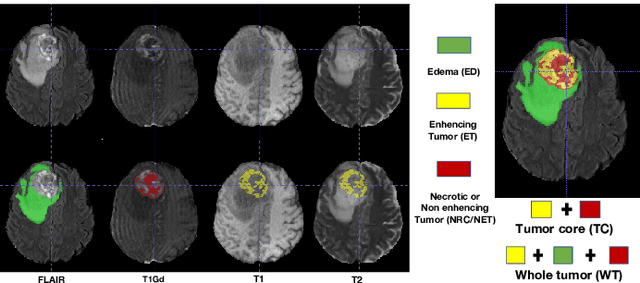
Abstract:Single-modality medical images generally do not contain enough information to reach an accurate and reliable diagnosis. For this reason, physicians generally diagnose diseases based on multimodal medical images such as, e.g., PET/CT. The effective fusion of multimodal information is essential to reach a reliable decision and explain how the decision is made as well. In this paper, we propose a fusion framework for multimodal medical image segmentation based on deep learning and the Dempster-Shafer theory of evidence. In this framework, the reliability of each single modality image when segmenting different objects is taken into account by a contextual discounting operation. The discounted pieces of evidence from each modality are then combined by Dempster's rule to reach a final decision. Experimental results with a PET-CT dataset with lymphomas and a multi-MRI dataset with brain tumors show that our method outperforms the state-of-the-art methods in accuracy and reliability.
Lymphoma segmentation from 3D PET-CT images using a deep evidential network
Jan 31, 2022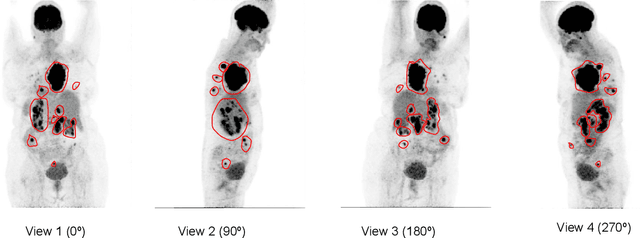
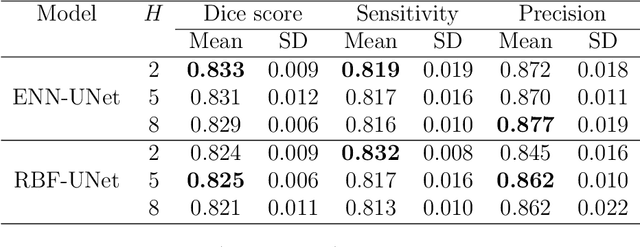
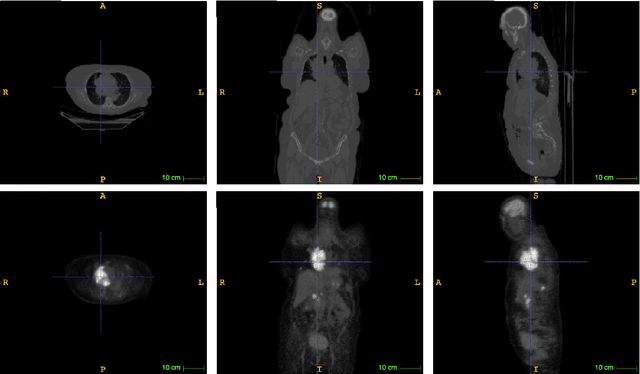
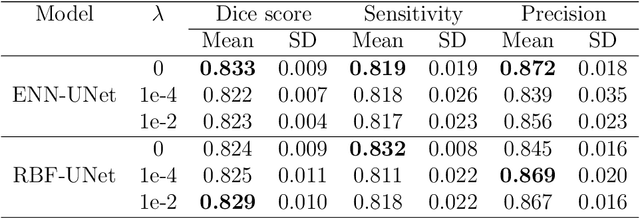
Abstract:An automatic evidential segmentation method based on Dempster-Shafer theory and deep learning is proposed to segment lymphomas from three-dimensional Positron Emission Tomography (PET) and Computed Tomography (CT) images. The architecture is composed of a deep feature-extraction module and an evidential layer. The feature extraction module uses an encoder-decoder framework to extract semantic feature vectors from 3D inputs. The evidential layer then uses prototypes in the feature space to compute a belief function at each voxel quantifying the uncertainty about the presence or absence of a lymphoma at this location. Two evidential layers are compared, based on different ways of using distances to prototypes for computing mass functions. The whole model is trained end-to-end by minimizing the Dice loss function. The proposed combination of deep feature extraction and evidential segmentation is shown to outperform the baseline UNet model as well as three other state-of-the-art models on a dataset of 173 patients.
AI-Based Detection, Classification and Prediction/Prognosis in Medical Imaging: Towards Radiophenomics
Nov 01, 2021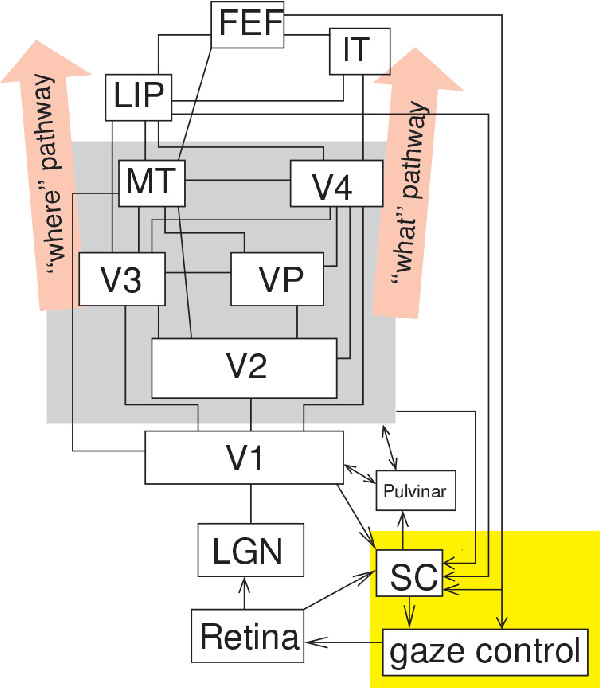
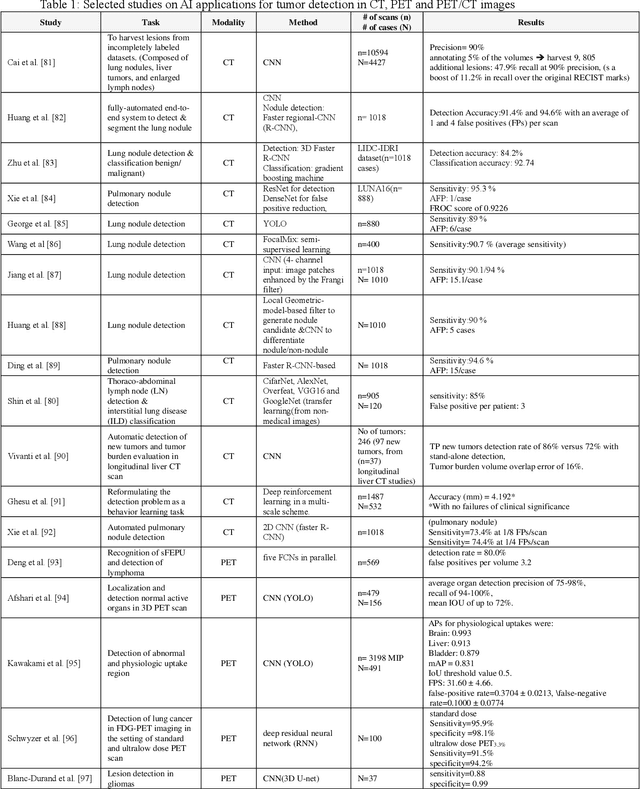

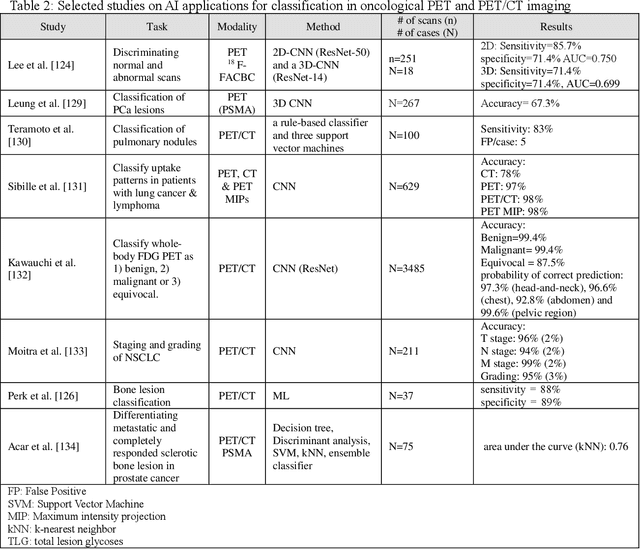
Abstract:Artificial intelligence (AI) techniques have significant potential to enable effective, robust and automated image phenotyping including identification of subtle patterns. AI-based detection searches the image space to find the regions of interest based on patterns and features. There is a spectrum of tumor histologies from benign to malignant that can be identified by AI-based classification approaches using image features. The extraction of minable information from images gives way to the field of radiomics and can be explored via explicit (handcrafted/engineered) and deep radiomics frameworks. Radiomics analysis has the potential to be utilized as a noninvasive technique for the accurate characterization of tumors to improve diagnosis and treatment monitoring. This work reviews AI-based techniques, with a special focus on oncological PET and PET/CT imaging, for different detection, classification, and prediction/prognosis tasks. We also discuss needed efforts to enable the translation of AI techniques to routine clinical workflows, and potential improvements and complementary techniques such as the use of natural language processing on electronic health records and neuro-symbolic AI techniques.
Deep PET/CT fusion with Dempster-Shafer theory for lymphoma segmentation
Aug 11, 2021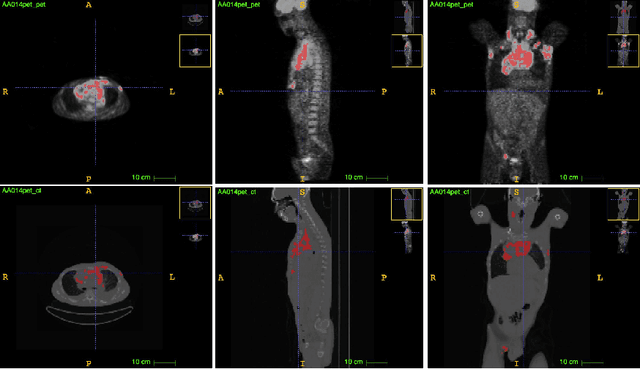
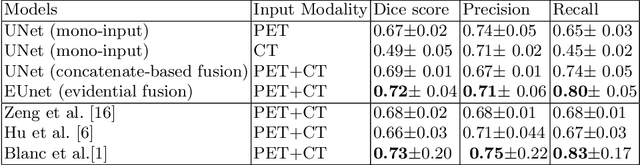
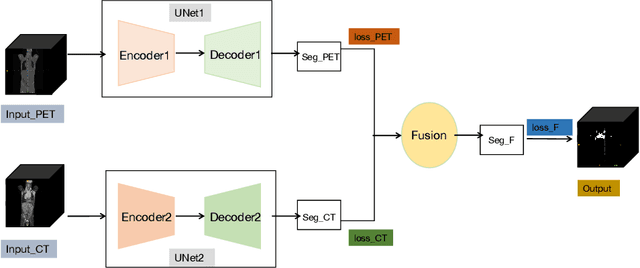
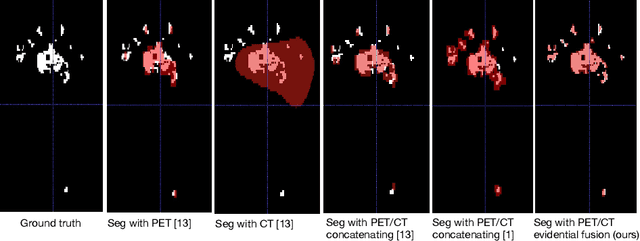
Abstract:Lymphoma detection and segmentation from whole-body Positron Emission Tomography/Computed Tomography (PET/CT) volumes are crucial for surgical indication and radiotherapy. Designing automatic segmentation methods capable of effectively exploiting the information from PET and CT as well as resolving their uncertainty remain a challenge. In this paper, we propose an lymphoma segmentation model using an UNet with an evidential PET/CT fusion layer. Single-modality volumes are trained separately to get initial segmentation maps and an evidential fusion layer is proposed to fuse the two pieces of evidence using Dempster-Shafer theory (DST). Moreover, a multi-task loss function is proposed: in addition to the use of the Dice loss for PET and CT segmentation, a loss function based on the concordance between the two segmentation is added to constrain the final segmentation. We evaluate our proposal on a database of polycentric PET/CT volumes of patients treated for lymphoma, delineated by the experts. Our method get accurate segmentation results with Dice score of 0.726, without any user interaction. Quantitative results show that our method is superior to the state-of-the-art methods.
Evidential segmentation of 3D PET/CT images
Apr 27, 2021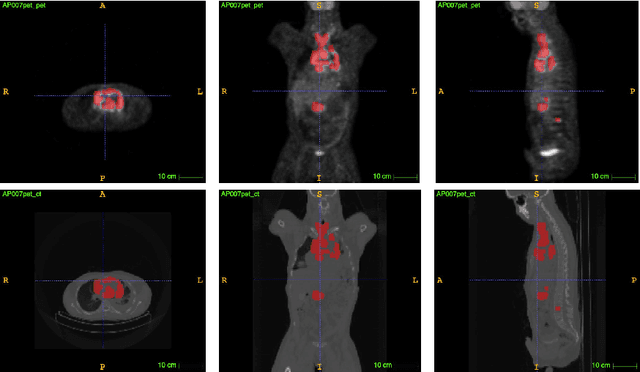
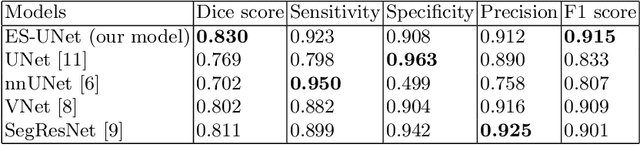
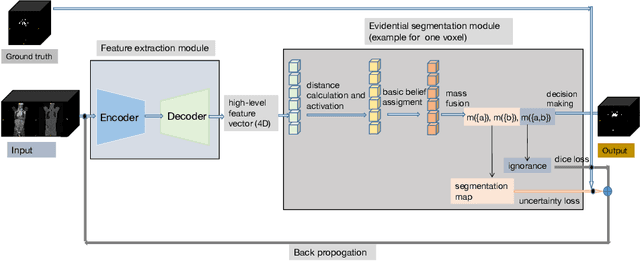
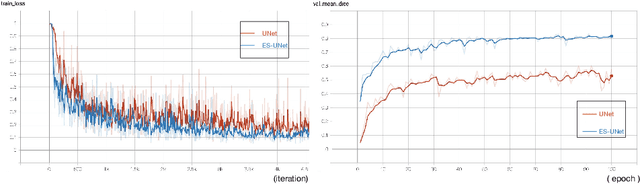
Abstract:PET and CT are two modalities widely used in medical image analysis. Accurately detecting and segmenting lymphomas from these two imaging modalities are critical tasks for cancer staging and radiotherapy planning. However, this task is still challenging due to the complexity of PET/CT images, and the computation cost to process 3D data. In this paper, a segmentation method based on belief functions is proposed to segment lymphomas in 3D PET/CT images. The architecture is composed of a feature extraction module and an evidential segmentation (ES) module. The ES module outputs not only segmentation results (binary maps indicating the presence or absence of lymphoma in each voxel) but also uncertainty maps quantifying the classification uncertainty. The whole model is optimized by minimizing Dice and uncertainty loss functions to increase segmentation accuracy. The method was evaluated on a database of 173 patients with diffuse large b-cell lymphoma. Quantitative and qualitative results show that our method outperforms the state-of-the-art methods.
RADIOGAN: Deep Convolutional Conditional Generative adversarial Network To Generate PET Images
Mar 19, 2020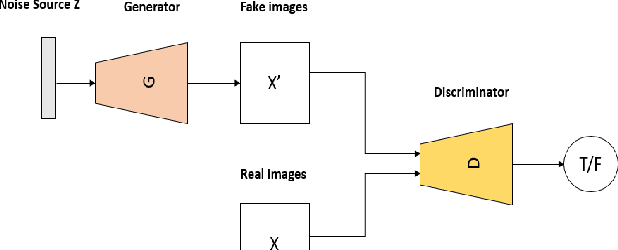
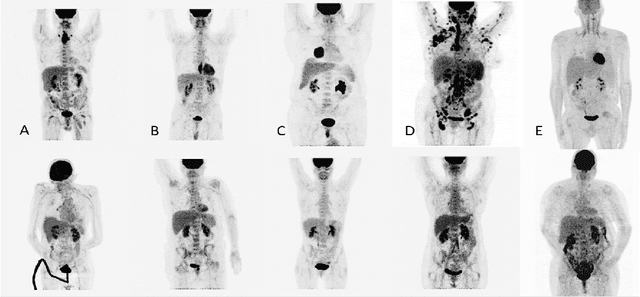
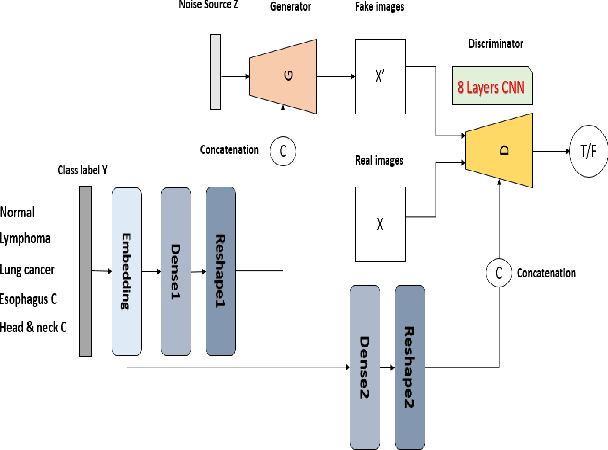
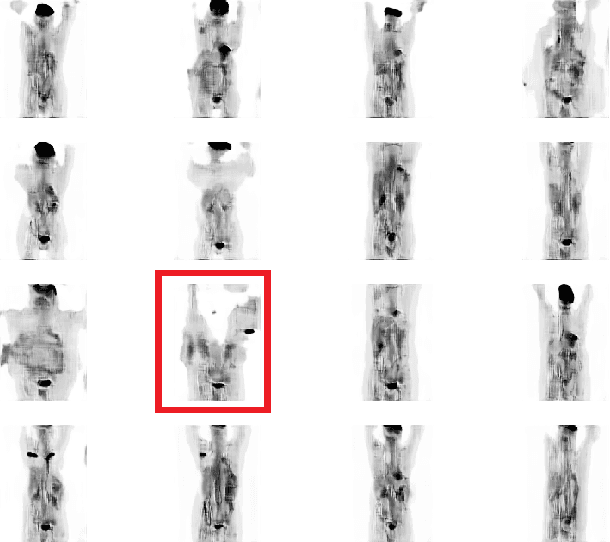
Abstract:One of the most challenges in medical imaging is the lack of data. It is proven that classical data augmentation methods are useful but still limited due to the huge variation in images. Using generative adversarial networks (GAN) is a promising way to address this problem, however, it is challenging to train one model to generate different classes of lesions. In this paper, we propose a deep convolutional conditional generative adversarial network to generate MIP positron emission tomography image (PET) which is a 2D image that represents a 3D volume for fast interpretation, according to different lesions or non lesion (normal). The advantage of our proposed method consists of one model that is capable of generating different classes of lesions trained on a small sample size for each class of lesion, and showing a very promising results. In addition, we show that a walk through a latent space can be used as a tool to evaluate the images generated.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge