Nomi L. Harris
The Artificial Intelligence Ontology: LLM-assisted construction of AI concept hierarchies
Apr 03, 2024


Abstract:The Artificial Intelligence Ontology (AIO) is a systematization of artificial intelligence (AI) concepts, methodologies, and their interrelations. Developed via manual curation, with the additional assistance of large language models (LLMs), AIO aims to address the rapidly evolving landscape of AI by providing a comprehensive framework that encompasses both technical and ethical aspects of AI technologies. The primary audience for AIO includes AI researchers, developers, and educators seeking standardized terminology and concepts within the AI domain. The ontology is structured around six top-level branches: Networks, Layers, Functions, LLMs, Preprocessing, and Bias, each designed to support the modular composition of AI methods and facilitate a deeper understanding of deep learning architectures and ethical considerations in AI. AIO's development utilized the Ontology Development Kit (ODK) for its creation and maintenance, with its content being dynamically updated through AI-driven curation support. This approach not only ensures the ontology's relevance amidst the fast-paced advancements in AI but also significantly enhances its utility for researchers, developers, and educators by simplifying the integration of new AI concepts and methodologies. The ontology's utility is demonstrated through the annotation of AI methods data in a catalog of AI research publications and the integration into the BioPortal ontology resource, highlighting its potential for cross-disciplinary research. The AIO ontology is open source and is available on GitHub (https://github.com/berkeleybop/artificial-intelligence-ontology) and BioPortal (https://bioportal.bioontology.org/ontologies/AIO).
MapperGPT: Large Language Models for Linking and Mapping Entities
Oct 05, 2023Abstract:Aligning terminological resources, including ontologies, controlled vocabularies, taxonomies, and value sets is a critical part of data integration in many domains such as healthcare, chemistry, and biomedical research. Entity mapping is the process of determining correspondences between entities across these resources, such as gene identifiers, disease concepts, or chemical entity identifiers. Many tools have been developed to compute such mappings based on common structural features and lexical information such as labels and synonyms. Lexical approaches in particular often provide very high recall, but low precision, due to lexical ambiguity. As a consequence of this, mapping efforts often resort to a labor intensive manual mapping refinement through a human curator. Large Language Models (LLMs), such as the ones employed by ChatGPT, have generalizable abilities to perform a wide range of tasks, including question-answering and information extraction. Here we present MapperGPT, an approach that uses LLMs to review and refine mapping relationships as a post-processing step, in concert with existing high-recall methods that are based on lexical and structural heuristics. We evaluated MapperGPT on a series of alignment tasks from different domains, including anatomy, developmental biology, and renal diseases. We devised a collection of tasks that are designed to be particularly challenging for lexical methods. We show that when used in combination with high-recall methods, MapperGPT can provide a substantial improvement in accuracy, beating state-of-the-art (SOTA) methods such as LogMap.
Gene Set Summarization using Large Language Models
May 25, 2023Abstract:Molecular biologists frequently interpret gene lists derived from high-throughput experiments and computational analysis. This is typically done as a statistical enrichment analysis that measures the over- or under-representation of biological function terms associated with genes or their properties, based on curated assertions from a knowledge base (KB) such as the Gene Ontology (GO). Interpreting gene lists can also be framed as a textual summarization task, enabling the use of Large Language Models (LLMs), potentially utilizing scientific texts directly and avoiding reliance on a KB. We developed SPINDOCTOR (Structured Prompt Interpolation of Natural Language Descriptions of Controlled Terms for Ontology Reporting), a method that uses GPT models to perform gene set function summarization as a complement to standard enrichment analysis. This method can use different sources of gene functional information: (1) structured text derived from curated ontological KB annotations, (2) ontology-free narrative gene summaries, or (3) direct model retrieval. We demonstrate that these methods are able to generate plausible and biologically valid summary GO term lists for gene sets. However, GPT-based approaches are unable to deliver reliable scores or p-values and often return terms that are not statistically significant. Crucially, these methods were rarely able to recapitulate the most precise and informative term from standard enrichment, likely due to an inability to generalize and reason using an ontology. Results are highly nondeterministic, with minor variations in prompt resulting in radically different term lists. Our results show that at this point, LLM-based methods are unsuitable as a replacement for standard term enrichment analysis and that manual curation of ontological assertions remains necessary.
Structured prompt interrogation and recursive extraction of semantics (SPIRES): A method for populating knowledge bases using zero-shot learning
Apr 05, 2023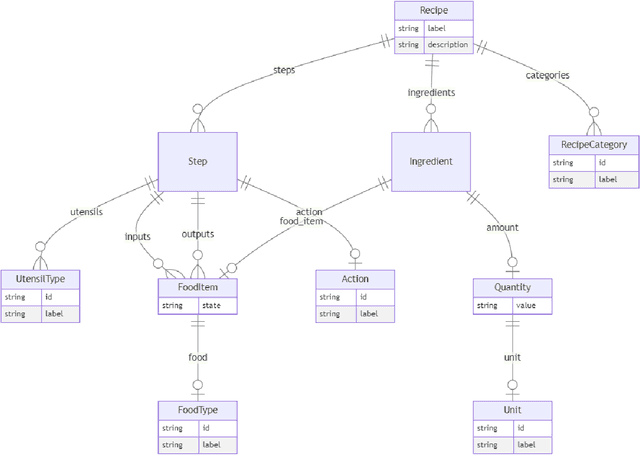
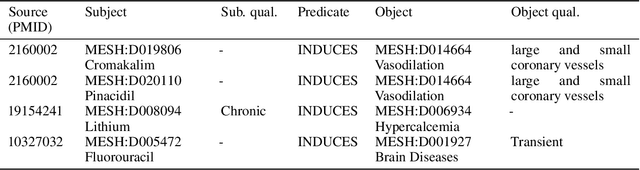
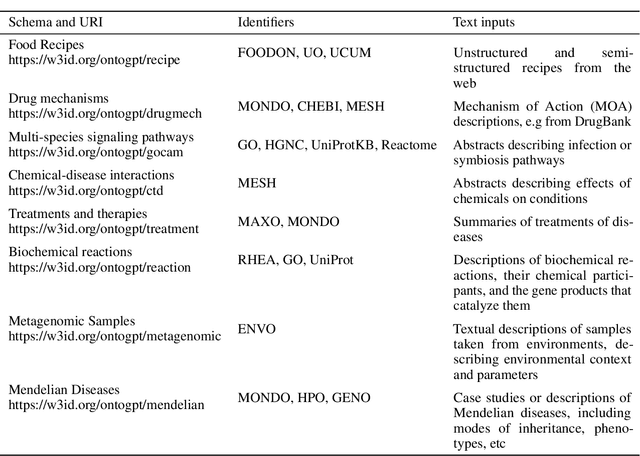
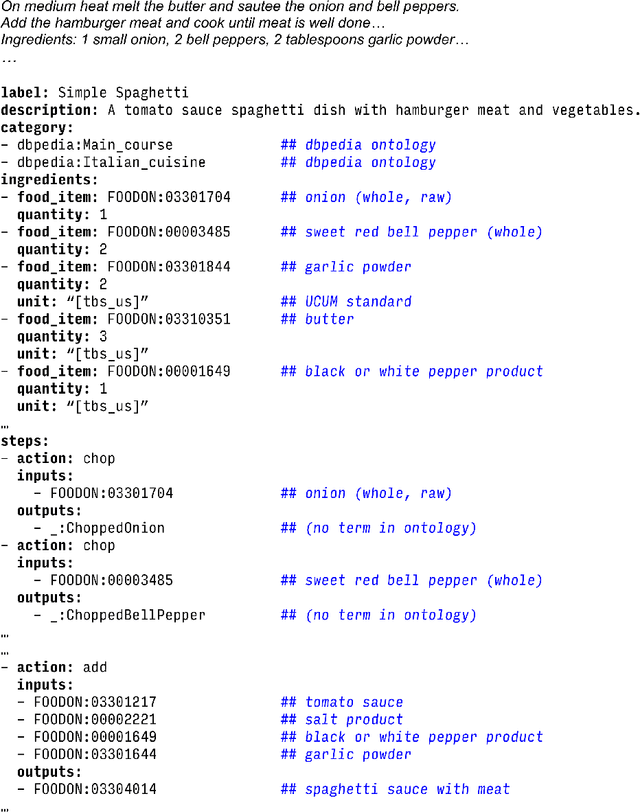

Abstract:Creating knowledge bases and ontologies is a time consuming task that relies on a manual curation. AI/NLP approaches can assist expert curators in populating these knowledge bases, but current approaches rely on extensive training data, and are not able to populate arbitrary complex nested knowledge schemas. Here we present Structured Prompt Interrogation and Recursive Extraction of Semantics (SPIRES), a Knowledge Extraction approach that relies on the ability of Large Language Models (LLMs) to perform zero-shot learning (ZSL) and general-purpose query answering from flexible prompts and return information conforming to a specified schema. Given a detailed, user-defined knowledge schema and an input text, SPIRES recursively performs prompt interrogation against GPT-3+ to obtain a set of responses matching the provided schema. SPIRES uses existing ontologies and vocabularies to provide identifiers for all matched elements. We present examples of use of SPIRES in different domains, including extraction of food recipes, multi-species cellular signaling pathways, disease treatments, multi-step drug mechanisms, and chemical to disease causation graphs. Current SPIRES accuracy is comparable to the mid-range of existing Relation Extraction (RE) methods, but has the advantage of easy customization, flexibility, and, crucially, the ability to perform new tasks in the absence of any training data. This method supports a general strategy of leveraging the language interpreting capabilities of LLMs to assemble knowledge bases, assisting manual knowledge curation and acquisition while supporting validation with publicly-available databases and ontologies external to the LLM. SPIRES is available as part of the open source OntoGPT package: https://github.com/ monarch-initiative/ontogpt.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge