Nicolas Paris
Terminologies augmented recurrent neural network model for clinical named entity recognition
May 15, 2019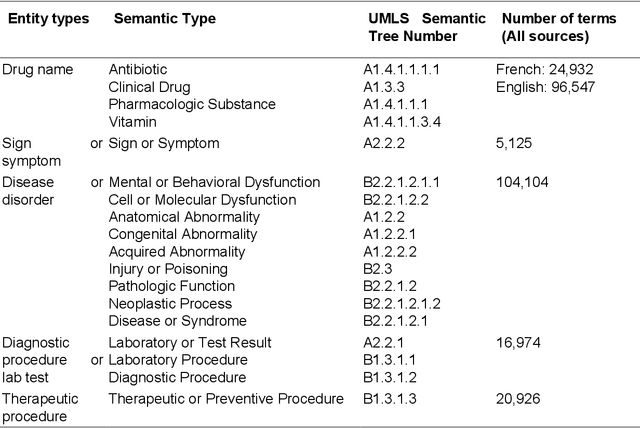
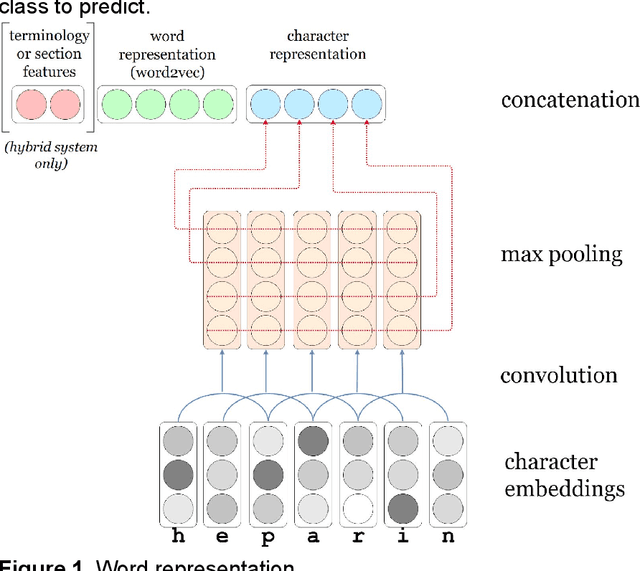
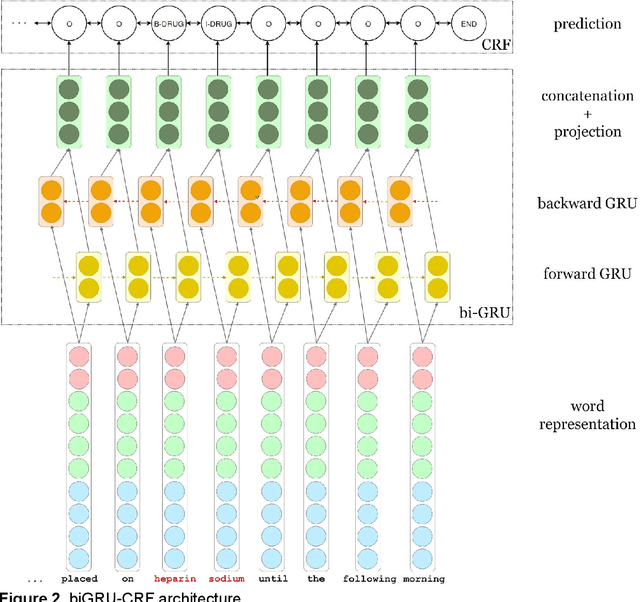
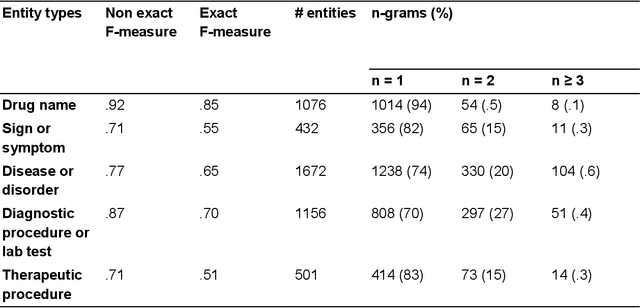
Abstract:We aimed to enhance the performance of a supervised model for clinical named-entity recognition (NER) using medical terminologies. In order to evaluate our system in French, we built a corpus for 5 types of clinical entities. We used a terminology-based system as baseline, built upon UMLS and SNOMED. Then, we evaluated a biGRU-CRF, and an hybrid system using the prediction of the terminology-based system as feature for the biGRU-CRF. In English, we evaluated the NER systems on the i2b2-2009 Medication Challenge for Drug name recognition, which contained 8,573 entities for 268 documents. In French, we built APcNER, a corpus of 147 documents annotated for 5 entities (drug name, sign or symptom, disease or disorder, diagnostic procedure or lab test and therapeutic procedure). We evaluated each NER systems using exact and partial match definition of F-measure for NER. The APcNER contains 4,837 entities which took 28 hours to annotate, the inter-annotator agreement was acceptable for Drug name in exact match (85%) and acceptable for other entity types in non-exact match (>70%). For drug name recognition on both i2b2-2009 and APcNER, the biGRU-CRF performed better that the terminology-based system, with an exact-match F-measure of 91.1% versus 73% and 81.9% versus 75% respectively. Moreover, the hybrid system outperformed the biGRU-CRF, with an exact-match F-measure of 92.2% versus 91.1% (i2b2-2009) and 88.4% versus 81.9% (APcNER). On APcNER corpus, the micro-average F-measure of the hybrid system on the 5 entities was 69.5% in exact match, and 84.1% in non-exact match. APcNER is a French corpus for clinical-NER of five type of entities which covers a large variety of document types. Extending supervised model with terminology allowed for an easy performance gain, especially in low regimes of entities, and established near state of the art results on the i2b2-2009 corpus.
BCMA-ES II: revisiting Bayesian CMA-ES
Apr 09, 2019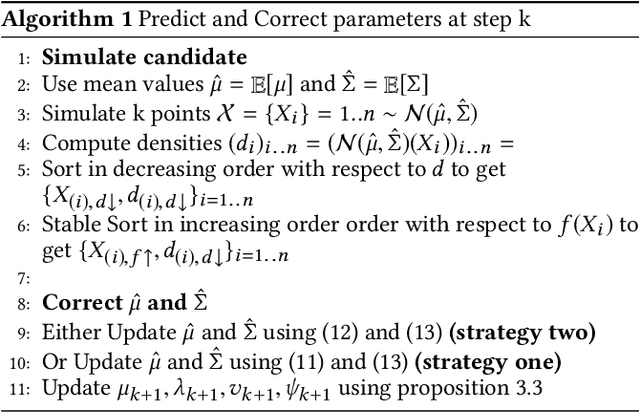
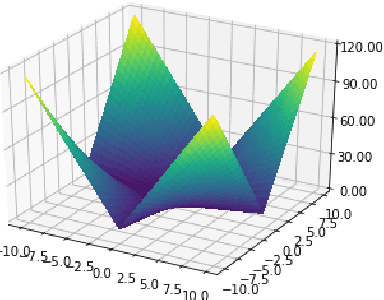
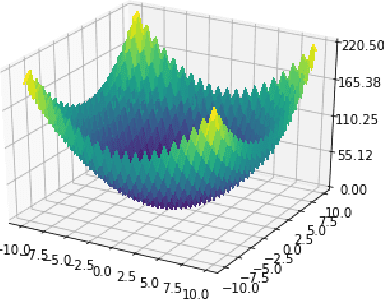
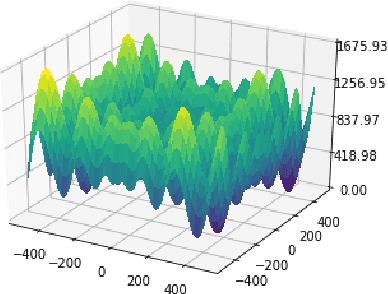
Abstract:This paper revisits the Bayesian CMA-ES and provides updates for normal Wishart. It emphasizes the difference between a normal and normal inverse Wishart prior. After some computation, we prove that the only difference relies surprisingly in the expected covariance. We prove that the expected covariance should be lower in the normal Wishart prior model because of the convexity of the inverse. We present a mixture model that generalizes both normal Wishart and normal inverse Wishart model. We finally present various numerical experiments to compare both methods as well as the generalized method.
Hybrid Approaches for our Participation to the n2c2 Challenge on Cohort Selection for Clinical Trials
Mar 19, 2019

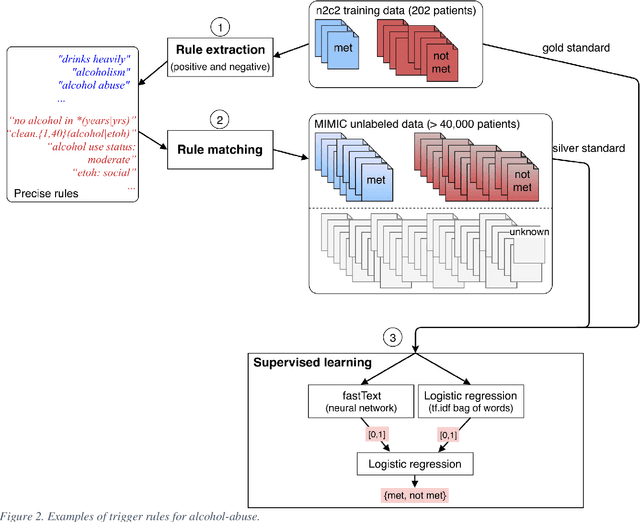
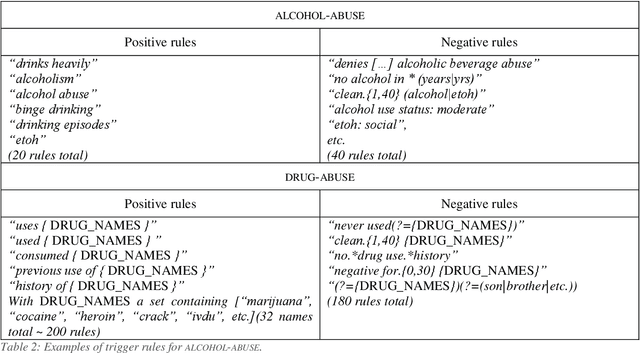
Abstract:Objective: Natural language processing can help minimize human intervention in identifying patients meeting eligibility criteria for clinical trials, but there is still a long way to go to obtain a general and systematic approach that is useful for researchers. We describe two methods taking a step in this direction and present their results obtained during the n2c2 challenge on cohort selection for clinical trials. Materials and Methods: The first method is a weakly supervised method using an unlabeled corpus (MIMIC) to build a silver standard, by producing semi-automatically a small and very precise set of rules to detect some samples of positive and negative patients. This silver standard is then used to train a traditional supervised model. The second method is a terminology-based approach where a medical expert selects the appropriate concepts, and a procedure is defined to search the terms and check the structural or temporal constraints. Results: On the n2c2 dataset containing annotated data about 13 selection criteria on 288 patients, we obtained an overall F1-measure of 0.8969, which is the third best result out of 45 participant teams, with no statistically significant difference with the best-ranked team. Discussion: Both approaches obtained very encouraging results and apply to different types of criteria. The weakly supervised method requires explicit descriptions of positive and negative examples in some reports. The terminology-based method is very efficient when medical concepts carry most of the relevant information. Conclusion: It is unlikely that much more annotated data will be soon available for the task of identifying a wide range of patient phenotypes. One must focus on weakly or non-supervised learning methods using both structured and unstructured data and relying on a comprehensive representation of the patients.
Solving WCSP by Extraction of Minimal Unsatisfiable Cores
Apr 19, 2013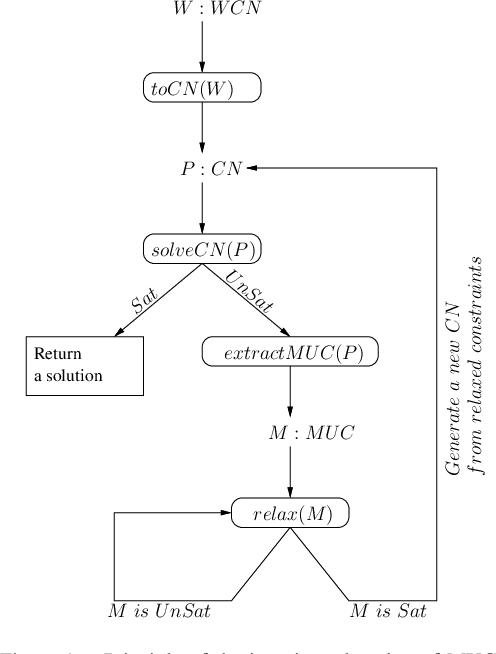
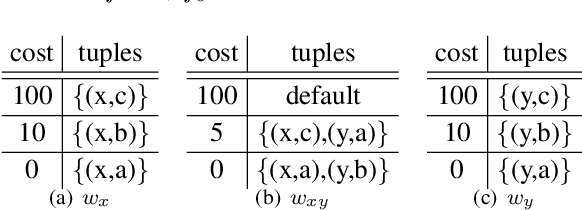
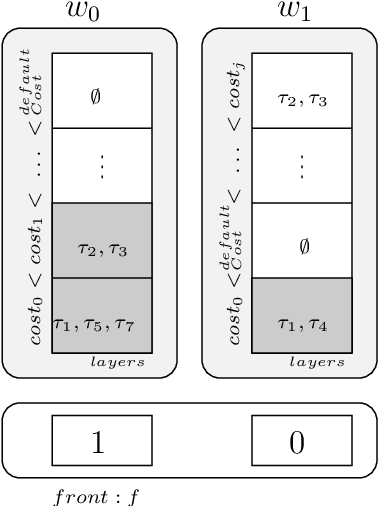

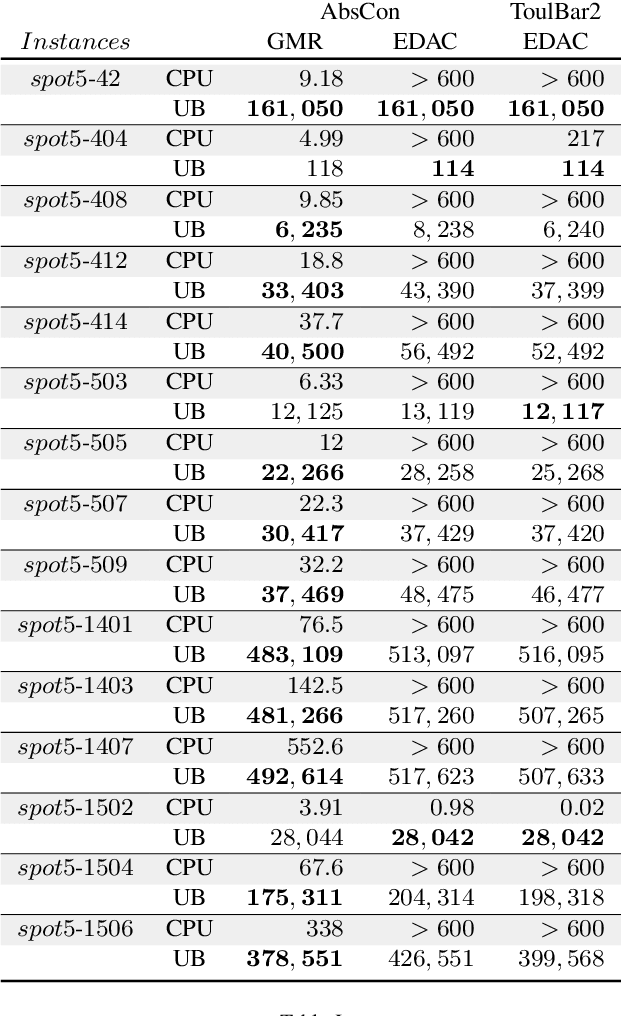
Abstract:Usual techniques to solve WCSP are based on cost transfer operations coupled with a branch and bound algorithm. In this paper, we focus on an approach integrating extraction and relaxation of Minimal Unsatisfiable Cores in order to solve this problem. We decline our approach in two ways: an incomplete, greedy, algorithm and a complete one.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge