Neelam Sinha
Multi-Task Learning with Additive U-Net for Image Denoising and Classification
Feb 13, 2026Abstract:We investigate additive skip fusion in U-Net architectures for image denoising and denoising-centric multi-task learning (MTL). By replacing concatenative skips with gated additive fusion, the proposed Additive U-Net (AddUNet) constrains shortcut capacity while preserving fixed feature dimensionality across depth. This structural regularization induces controlled encoder-decoder information flow and stabilizes joint optimization. Across single-task denoising and joint denoising-classification settings, AddUNet achieves competitive reconstruction performance with improved training stability. In MTL, learned skip weights exhibit systematic task-aware redistribution: shallow skips favor reconstruction, while deeper features support discrimination. Notably, reconstruction remains robust even under limited classification capacity, indicating implicit task decoupling through additive fusion. These findings show that simple constraints on skip connections act as an effective architectural regularizer for stable and scalable multi-task learning without increasing model complexity.
Chest X-ray Classification using Deep Convolution Models on Low-resolution images with Uncertain Labels
Apr 12, 2025Abstract:Deep Convolutional Neural Networks have consistently proven to achieve state-of-the-art results on a lot of imaging tasks over the past years' majority of which comprise of high-quality data. However, it is important to work on low-resolution images since it could be a cheaper alternative for remote healthcare access where the primary need of automated pathology identification models occurs. Medical diagnosis using low-resolution images is challenging since critical details may not be easily identifiable. In this paper, we report classification results by experimenting on different input image sizes of Chest X-rays to deep CNN models and discuss the feasibility of classification on varying image sizes. We also leverage the noisy labels in the dataset by proposing a Randomized Flipping of labels techniques. We use an ensemble of multi-label classification models on frontal and lateral studies. Our models are trained on 5 out of the 14 chest pathologies of the publicly available CheXpert dataset. We incorporate techniques such as augmentation, regularization for model improvement and use class activation maps to visualize the neural network's decision making. Comparison with classification results on data from 200 subjects, obtained on the corresponding high-resolution images, reported in the original CheXpert paper, has been presented. For pathologies Cardiomegaly, Consolidation and Edema, we obtain 3% higher accuracy with our model architecture.
Graph Classification and Radiomics Signature for Identification of Tuberculous Meningitis
Apr 01, 2025
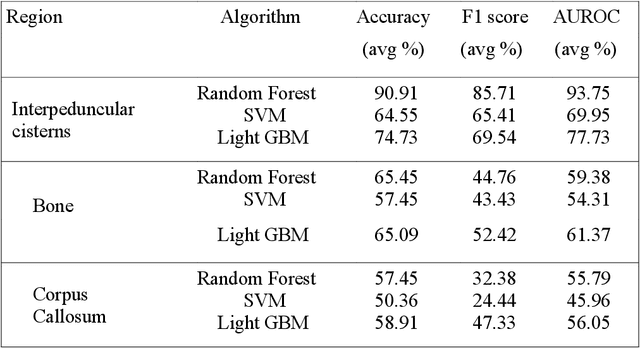


Abstract:Introduction: Tuberculous meningitis (TBM) is a serious brain infection caused by Mycobacterium tuberculosis, characterized by inflammation of the meninges covering the brain and spinal cord. Diagnosis often requires invasive lumbar puncture (LP) and cerebrospinal fluid (CSF) analysis. Objectives: This study aims to classify TBM patients using T1-weighted (T1w) non-contrast Magnetic Resonance Imaging (MRI) scans. We hypothesize that specific brain regions, such as the interpeduncular cisterns, bone, and corpus callosum, contain visual markers that can non-invasively distinguish TBM patients from healthy controls. We propose a novel Pixel-array Graphs Classifier (PAG-Classifier) that leverages spatial relationships between neighbouring 3D pixels in a graph-based framework to extract significant features through eigen decomposition. These features are then used to train machine learning classifiers for effective patient classification. We validate our approach using a radiomics-based methodology, classifying TBM patients based on relevant radiomics features. Results: We utilized an internal dataset consisting of 52 scans, 32 from confirmed TBM patients based on mycobacteria detection in CSF, and 20 from healthy individuals. We achieved a 5-fold cross-validated average F1 score of 85.71% for cistern regions with our PAG-Classifier and 92.85% with the radiomics features classifier, surpassing current state-of-the-art benchmarks by 15% and 22%, respectively. However, bone and corpus callosum regions showed poor classification effectiveness, with average F1 scores below 50%. Conclusion: Our study suggests that algorithms like the PAG-Classifier serve as effective tools for non-invasive TBM analysis, particularly by targeting the interpeduncular cistern. Findings indicate that the bone and corpus callosum regions lack distinctive patterns for differentiation.
Alzheimer's Disease Classification Using Retinal OCT: TransnetOCT and Swin Transformer Models
Mar 14, 2025Abstract:Retinal optical coherence tomography (OCT) images are the biomarkers for neurodegenerative diseases, which are rising in prevalence. Early detection of Alzheimer's disease using retinal OCT is a primary challenging task. This work utilizes advanced deep learning techniques to classify retinal OCT images of subjects with Alzheimer's disease (AD) and healthy controls (CO). The goal is to enhance diagnostic capabilities through efficient image analysis. In the proposed model, Raw OCT images have been preprocessed with ImageJ and given to various deep-learning models to evaluate the accuracy. The best classification architecture is TransNetOCT, which has an average accuracy of 98.18% for input OCT images and 98.91% for segmented OCT images for five-fold cross-validation compared to other models, and the Swin Transformer model has achieved an accuracy of 93.54%. The evaluation accuracy metric demonstrated TransNetOCT and Swin transformer models capability to classify AD and CO subjects reliably, contributing to the potential for improved diagnostic processes in clinical settings.
Persistent Homology for MCI Classification: A Comparative Analysis between Graph and Vietoris-Rips Filtrations
Oct 30, 2024



Abstract:Mild cognitive impairment (MCI), often linked to early neurodegeneration, is characterized by subtle cognitive declines and disruptions in brain connectivity. The present study offers a detailed analysis of topological changes associated with MCI, focusing on two subtypes: Early MCI and Late MCI. This analysis utilizes fMRI time series data from two distinct populations: the publicly available ADNI dataset (Western cohort) and the in-house TLSA dataset (Indian Urban cohort). Persistent Homology, a topological data analysis method, is employed with two distinct filtration techniques - Vietoris-Rips and graph filtration-for classifying MCI subtypes. For Vietoris-Rips filtration, inter-ROI Wasserstein distance matrices between persistent diagrams are used for classification, while graph filtration relies on the top ten most persistent homology features. Comparative analysis shows that the Vietoris-Rips filtration significantly outperforms graph filtration, capturing subtle variations in brain connectivity with greater accuracy. The Vietoris-Rips filtration method achieved the highest classification accuracy of 85.7\% for distinguishing between age and gender matched healthy controls and MCI, whereas graph filtration reached a maximum accuracy of 71.4\% for the same task. This superior performance highlights the sensitivity of Vietoris-Rips filtration in detecting intricate topological features associated with neurodegeneration. The findings underscore the potential of persistent homology, particularly when combined with the Wasserstein distance, as a powerful tool for early diagnosis and precise classification of cognitive impairments, offering valuable insights into brain connectivity changes in MCI.
Leveraging Persistent Homology for Differential Diagnosis of Mild Cognitive Impairment
Aug 28, 2024



Abstract:Mild cognitive impairment (MCI) is characterized by subtle changes in cognitive functions, often associated with disruptions in brain connectivity. The present study introduces a novel fine-grained analysis to examine topological alterations in neurodegeneration pertaining to six different brain networks of MCI subjects (Early/Late MCI). To achieve this, fMRI time series from two distinct populations are investigated: (i) the publicly accessible ADNI dataset and (ii) our in-house dataset. The study utilizes sliding window embedding to convert each fMRI time series into a sequence of 3-dimensional vectors, facilitating the assessment of changes in regional brain topology. Distinct persistence diagrams are computed for Betti descriptors of dimension-0, 1, and 2. Wasserstein distance metric is used to quantify differences in topological characteristics. We have examined both (i) ROI-specific inter-subject interactions and (ii) subject-specific inter-ROI interactions. Further, a new deep learning model is proposed for classification, achieving a maximum classification accuracy of 95% for the ADNI dataset and 85% for the in-house dataset. This methodology is further adapted for the differential diagnosis of MCI sub-types, resulting in a peak accuracy of 76.5%, 91.1% and 80% in classifying HC Vs. EMCI, HC Vs. LMCI and EMCI Vs. LMCI, respectively. We showed that the proposed approach surpasses current state-of-the-art techniques designed for classifying MCI and its sub-types using fMRI.
Non-linear Analysis Based ECG Classification of Cardiovascular Disorders
Aug 02, 2024



Abstract:Multi-channel ECG-based cardiac disorders detection has an impact on cardiac care and treatment. Limitations of existing methods included variation in ECG waveforms due to the location of electrodes, high non-linearity in the signal, and amplitude measurement in millivolts. The present study reports a non-linear analysis-based methodology that utilizes Recurrence plot visualization. The patterned occurrence of well-defined structures, such as the QRS complex, can be exploited effectively using Recurrence plots. This Recurrence-based method is applied to the publicly available Physikalisch-Technische Bundesanstalt (PTB) dataset from PhysioNet database, where we studied four classes of different cardiac disorders (Myocardial infarction, Bundle branch blocks, Cardiomyopathy, and Dysrhythmia) and healthy controls, achieving an impressive classification accuracy of 100%. Additionally, t-SNE plot visualizations of the latent space embeddings derived from Recurrence plots and Recurrence Quantification Analysis features reveal a clear demarcation between the considered cardiac disorders and healthy individuals, demonstrating the potential of this approach.
Classification of Alzheimer's Dementia vs. Healthy subjects by studying structural disparities in fMRI Time-Series of DMN
Jul 29, 2024



Abstract:Time series from different regions of interest (ROI) of default mode network (DMN) from Functional Magnetic Resonance Imaging (fMRI) can reveal significant differences between healthy and unhealthy people. Here, we propose the utility of an existing metric quantifying the lack/presence of structure in a signal called, "deviation from stochasticity" (DS) measure to characterize resting-state fMRI time series. The hypothesis is that differences in the level of structure in the time series can lead to discrimination between the subject groups. In this work, an autoencoder-based model is utilized to learn efficient representations of data by training the network to reconstruct its input data. The proposed methodology is applied on fMRI time series of 50 healthy individuals and 50 subjects with Alzheimer's Disease (AD), obtained from publicly available ADNI database. DS measure for healthy fMRI as expected turns out to be different compared to that of AD. Peak classification accuracy of 95% was obtained using Gradient Boosting classifier, using the DS measure applied on 100 subjects.
Analyzing Brain Tumor Connectomics using Graphs and Persistent Homology
Jul 25, 2024



Abstract:Recent advances in molecular and genetic research have identified a diverse range of brain tumor sub-types, shedding light on differences in their molecular mechanisms, heterogeneity, and origins. The present study performs whole-brain connectome analysis using diffusionweighted images. To achieve this, both graph theory and persistent homology - a prominent approach in topological data analysis are employed in order to quantify changes in the structural connectivity of the wholebrain connectome in subjects with brain tumors. Probabilistic tractography is used to map the number of streamlines connecting 84 distinct brain regions, as delineated by the Desikan-Killiany atlas from FreeSurfer. These streamline mappings form the connectome matrix, on which persistent homology based analysis and graph theoretical analysis are executed to evaluate the discriminatory power between tumor sub-types that include meningioma and glioma. A detailed statistical analysis is conducted on persistent homology-derived topological features and graphical features to identify the brain regions where differences between study groups are statistically significant (p < 0.05). For classification purpose, graph-based local features are utilized, achieving a highest accuracy of 88%. In classifying tumor sub-types, an accuracy of 80% is attained. The findings obtained from this study underscore the potential of persistent homology and graph theoretical analysis of the whole-brain connectome in detecting alterations in structural connectivity patterns specific to different types of brain tumors.
MRI Volume-Based Robust Brain Age Estimation Using Weight-Shared Spatial Attention in 3D CNNs
Jul 09, 2024



Abstract:Important applications of advancements in machine learning, are in the area of healthcare, more so for neurological disorder detection. A crucial step towards understanding the neurological status, is to estimate the brain age using structural MRI volumes, in order to measure its deviation from chronological age. Factors that contribute to brain age are best captured using a data-driven approach, such as deep learning. However, it places a huge demand on the availability of diverse datasets. In this work, we propose a robust brain age estimation paradigm that utilizes a 3D CNN model, by-passing the need for model-retraining across datasets. The proposed model consists of seven 3D CNN layers, with a shared spatial attention layer incorporated at each CNN layer followed by five dense layers. The novelty of the proposed method lies in the idea of spatial attention module, with shared weights across the CNN layers. This weight sharing ensures directed attention to specific brain regions, for localizing age-related features within the data, lending robustness. The proposed model, trained on ADNI dataset comprising 516 T1 weighted MRI volumes of healthy subjects, resulted in Mean Absolute Error (MAE) of 1.662 years, which is an improvement of 1.688 years over the state-of-the-art (SOTA) model, based on disjoint test samples from the same repository. To illustrate generalizability, the same pipeline was utilized on volumes from a publicly available source called OASIS3. From OASIS3, MRI volumes 890 healthy subjects were utilized resulting in MAE of 2.265 years. Due to diversity in acquisitions across multiple sites, races and genetic factors, traditional CNN models are not guaranteed to prioritize brain regions crucial for age estimation. In contrast, the proposed weight-shared spatial attention module, directs attention on specific regions, required for the estimation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge