Narisu Bai
Spatial Context-Aware Self-Attention Model For Multi-Organ Segmentation
Dec 16, 2020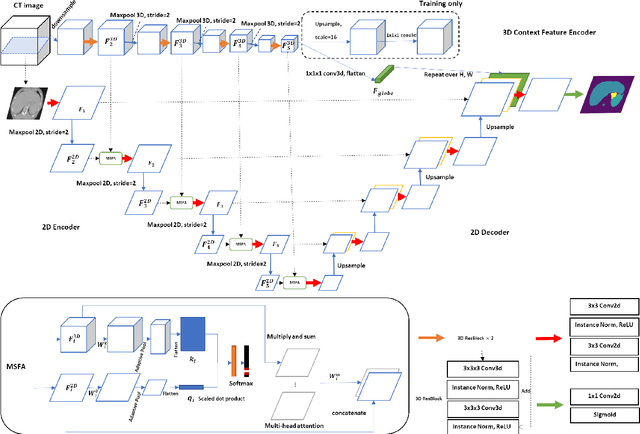

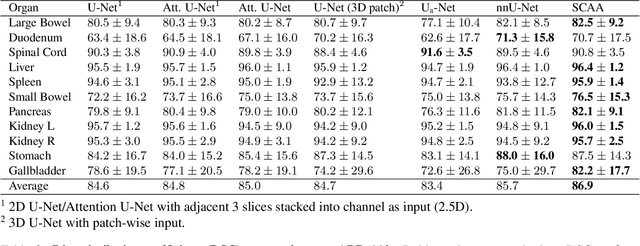
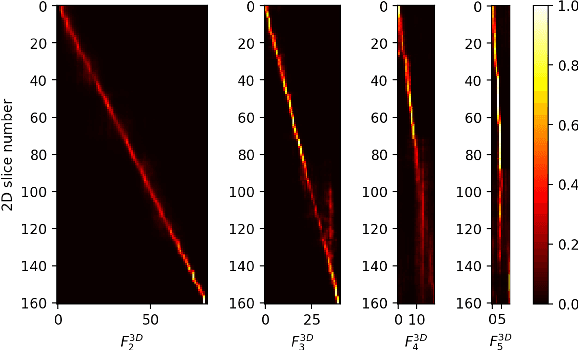
Abstract:Multi-organ segmentation is one of most successful applications of deep learning in medical image analysis. Deep convolutional neural nets (CNNs) have shown great promise in achieving clinically applicable image segmentation performance on CT or MRI images. State-of-the-art CNN segmentation models apply either 2D or 3D convolutions on input images, with pros and cons associated with each method: 2D convolution is fast, less memory-intensive but inadequate for extracting 3D contextual information from volumetric images, while the opposite is true for 3D convolution. To fit a 3D CNN model on CT or MRI images on commodity GPUs, one usually has to either downsample input images or use cropped local regions as inputs, which limits the utility of 3D models for multi-organ segmentation. In this work, we propose a new framework for combining 3D and 2D models, in which the segmentation is realized through high-resolution 2D convolutions, but guided by spatial contextual information extracted from a low-resolution 3D model. We implement a self-attention mechanism to control which 3D features should be used to guide 2D segmentation. Our model is light on memory usage but fully equipped to take 3D contextual information into account. Experiments on multiple organ segmentation datasets demonstrate that by taking advantage of both 2D and 3D models, our method is consistently outperforms existing 2D and 3D models in organ segmentation accuracy, while being able to directly take raw whole-volume image data as inputs.
AttentionAnatomy: A unified framework for whole-body organs at risk segmentation using multiple partially annotated datasets
Jan 13, 2020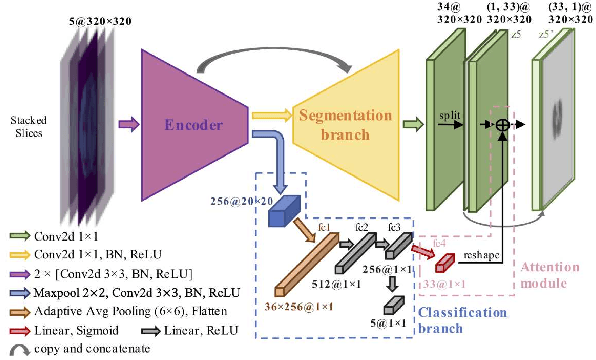
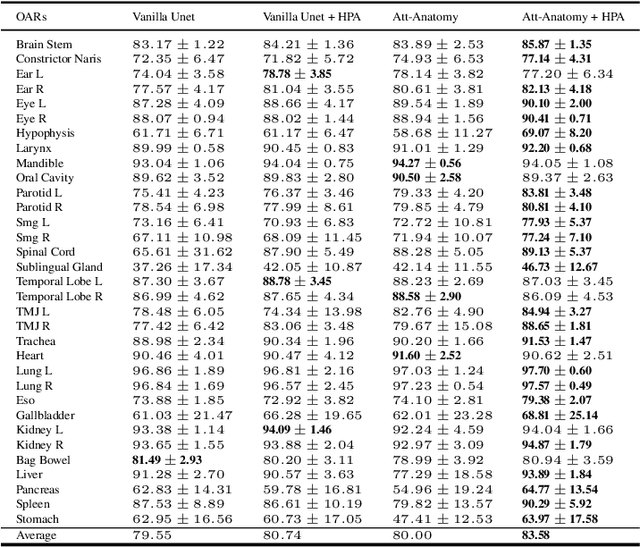
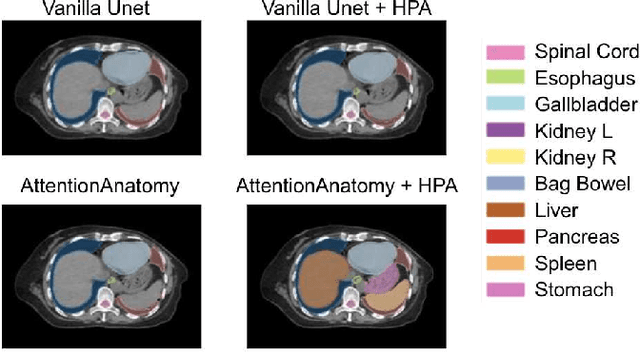
Abstract:Organs-at-risk (OAR) delineation in computed tomography (CT) is an important step in Radiation Therapy (RT) planning. Recently, deep learning based methods for OAR delineation have been proposed and applied in clinical practice for separate regions of the human body (head and neck, thorax, and abdomen). However, there are few researches regarding the end-to-end whole-body OARs delineation because the existing datasets are mostly partially or incompletely annotated for such task. In this paper, our proposed end-to-end convolutional neural network model, called \textbf{AttentionAnatomy}, can be jointly trained with three partially annotated datasets, segmenting OARs from whole body. Our main contributions are: 1) an attention module implicitly guided by body region label to modulate the segmentation branch output; 2) a prediction re-calibration operation, exploiting prior information of the input images, to handle partial-annotation(HPA) problem; 3) a new hybrid loss function combining batch Dice loss and spatially balanced focal loss to alleviate the organ size imbalance problem. Experimental results of our proposed framework presented significant improvements in both S{\o}rensen-Dice coefficient (DSC) and 95\% Hausdorff distance compared to the baseline model.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge