Markus D. Herrmann
Characterizing the Interpretability of Attention Maps in Digital Pathology
Jul 02, 2024



Abstract:Interpreting machine learning model decisions is crucial for high-risk applications like healthcare. In digital pathology, large whole slide images (WSIs) are decomposed into smaller tiles and tile-derived features are processed by attention-based multiple instance learning (ABMIL) models to predict WSI-level labels. These networks generate tile-specific attention weights, which can be visualized as attention maps for interpretability. However, a standardized evaluation framework for these maps is lacking, questioning their reliability and ability to detect spurious correlations that can mislead models. We herein propose a framework to assess the ability of attention networks to attend to relevant features in digital pathology by creating artificial model confounders and using dedicated interpretability metrics. Models are trained and evaluated on data with tile modifications correlated with WSI labels, enabling the analysis of model sensitivity to artificial confounders and the accuracy of attention maps in highlighting them. Confounders are introduced either through synthetic tile modifications or through tile ablations based on their specific image-based features, with the latter being used to assess more clinically relevant scenarios. We also analyze the impact of varying confounder quantities at both the tile and WSI levels. Our results show that ABMIL models perform as desired within our framework. While attention maps generally highlight relevant regions, their robustness is affected by the type and number of confounders. Our versatile framework has the potential to be used in the evaluation of various methods and the exploration of image-based features driving model predictions, which could aid in biomarker discovery.
The NCI Imaging Data Commons as a platform for reproducible research in computational pathology
Mar 16, 2023



Abstract:Objective: Reproducibility is critical for translating machine learning-based (ML) solutions in computational pathology (CompPath) into practice. However, an increasing number of studies report difficulties in reproducing ML results. The NCI Imaging Data Commons (IDC) is a public repository of >120 cancer image collections, including >38,000 whole-slide images (WSIs), that is designed to be used with cloud-based ML services. Here, we explore the potential of the IDC to facilitate reproducibility of CompPath research. Materials and Methods: The IDC realizes the FAIR principles: All images are encoded according to the DICOM standard, persistently identified, discoverable via rich metadata, and accessible via open tools. Taking advantage of this, we implemented two experiments in which a representative ML-based method for classifying lung tumor tissue was trained and/or evaluated on different datasets from the IDC. To assess reproducibility, the experiments were run multiple times with independent but identically configured sessions of common ML services. Results: The AUC values of different runs of the same experiment were generally consistent and in the same order of magnitude as a similar, previously published study. However, there were occasional small variations in AUC values of up to 0.044, indicating a practical limit to reproducibility. Discussion and conclusion: By realizing the FAIR principles, the IDC enables other researchers to reuse exactly the same datasets. Cloud-based ML services enable others to run CompPath experiments in an identically configured computing environment without having to own high-performance hardware. The combination of both makes it possible to approach the reproducibility limit.
MITI Minimum Information guidelines for highly multiplexed tissue images
Aug 21, 2021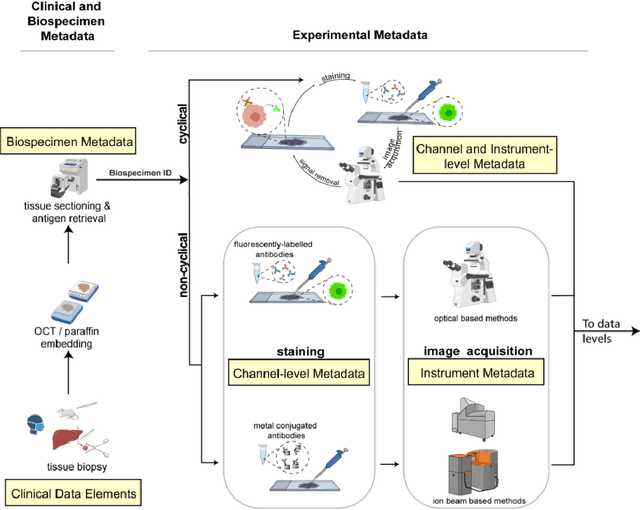
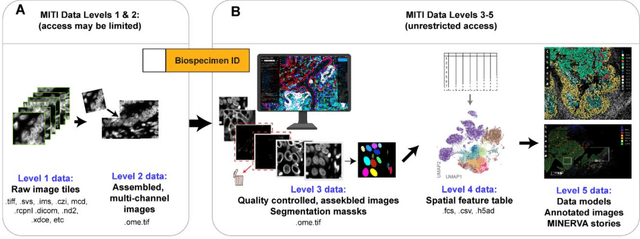
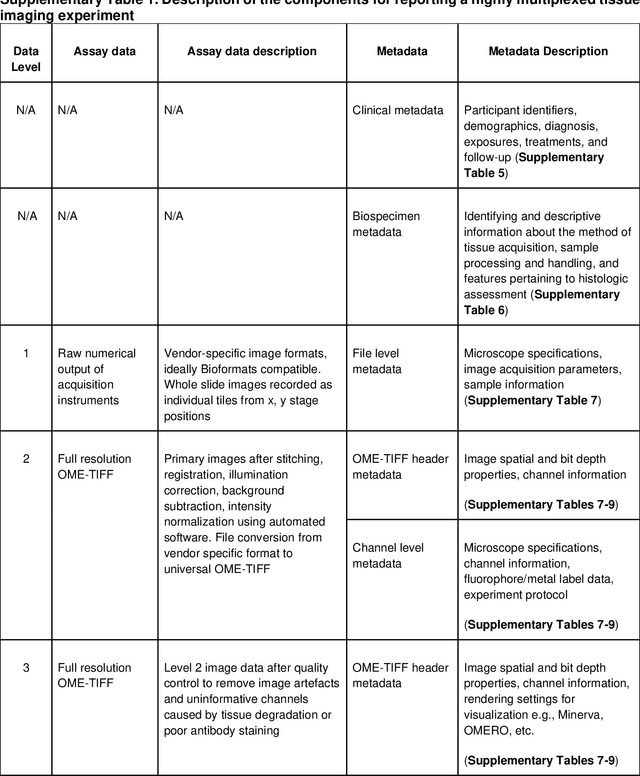
Abstract:The imminent release of atlases combining highly multiplexed tissue imaging with single cell sequencing and other omics data from human tissues and tumors creates an urgent need for data and metadata standards compliant with emerging and traditional approaches to histology. We describe the development of a Minimum Information about highly multiplexed Tissue Imaging (MITI) standard that draws on best practices from genomics and microscopy of cultured cells and model organisms.
Highdicom: A Python library for standardized encoding of image annotations and machine learning model outputs in pathology and radiology
Jun 14, 2021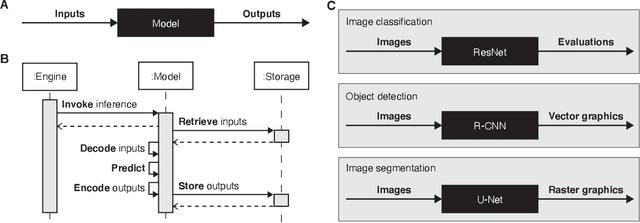
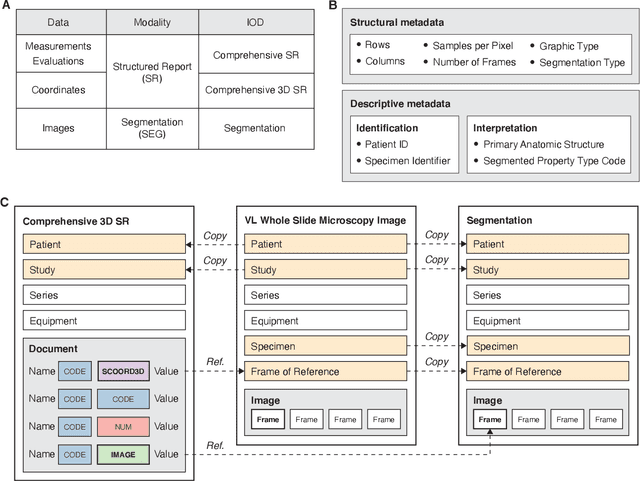
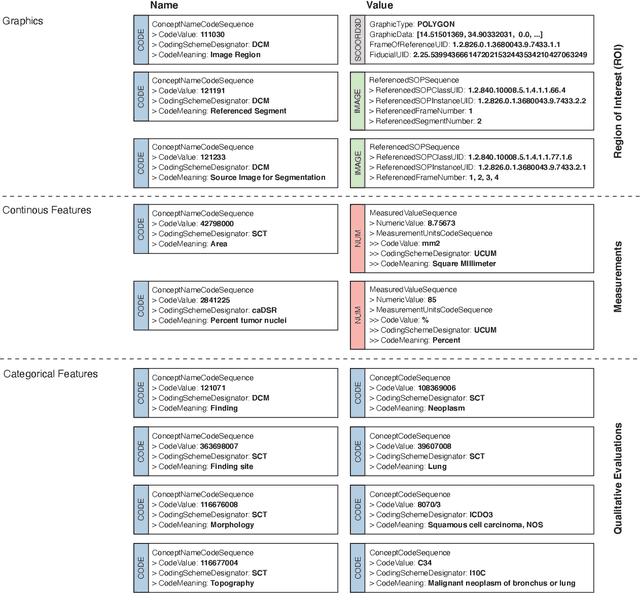
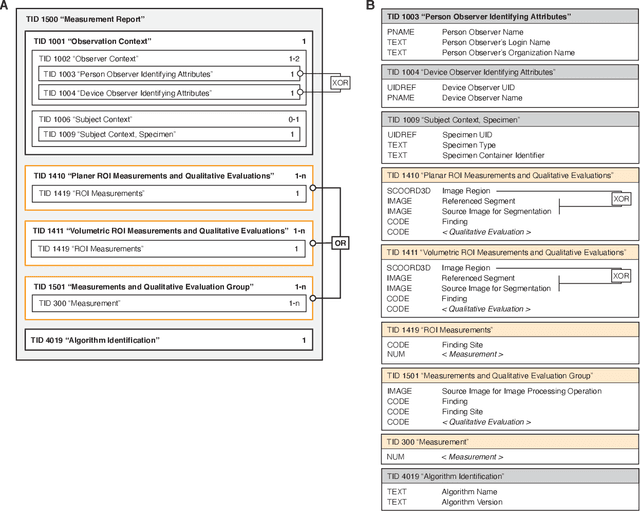
Abstract:Machine learning is revolutionizing image-based diagnostics in pathology and radiology. ML models have shown promising results in research settings, but their lack of interoperability has been a major barrier for clinical integration and evaluation. The DICOM a standard specifies Information Object Definitions and Services for the representation and communication of digital images and related information, including image-derived annotations and analysis results. However, the complexity of the standard represents an obstacle for its adoption in the ML community and creates a need for software libraries and tools that simplify working with data sets in DICOM format. Here we present the highdicom library, which provides a high-level application programming interface for the Python programming language that abstracts low-level details of the standard and enables encoding and decoding of image-derived information in DICOM format in a few lines of Python code. The highdicom library ties into the extensive Python ecosystem for image processing and machine learning. Simultaneously, by simplifying creation and parsing of DICOM-compliant files, highdicom achieves interoperability with the medical imaging systems that hold the data used to train and run ML models, and ultimately communicate and store model outputs for clinical use. We demonstrate through experiments with slide microscopy and computed tomography imaging, that, by bridging these two ecosystems, highdicom enables developers to train and evaluate state-of-the-art ML models in pathology and radiology while remaining compliant with the DICOM standard and interoperable with clinical systems at all stages. To promote standardization of ML research and streamline the ML model development and deployment process, we made the library available free and open-source.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge