David A. Clunie
PixelMed Publishing LLC, Bangor, PA, USA
From Data to Diagnosis: A Large, Comprehensive Bone Marrow Dataset and AI Methods for Childhood Leukemia Prediction
Sep 19, 2025Abstract:Leukemia diagnosis primarily relies on manual microscopic analysis of bone marrow morphology supported by additional laboratory parameters, making it complex and time consuming. While artificial intelligence (AI) solutions have been proposed, most utilize private datasets and only cover parts of the diagnostic pipeline. Therefore, we present a large, high-quality, publicly available leukemia bone marrow dataset spanning the entire diagnostic process, from cell detection to diagnosis. Using this dataset, we further propose methods for cell detection, cell classification, and diagnosis prediction. The dataset comprises 246 pediatric patients with diagnostic, clinical and laboratory information, over 40 000 cells with bounding box annotations and more than 28 000 of these with high-quality class labels, making it the most comprehensive dataset publicly available. Evaluation of the AI models yielded an average precision of 0.96 for the cell detection, an area under the curve of 0.98, and an F1-score of 0.61 for the 33-class cell classification, and a mean F1-score of 0.90 for the diagnosis prediction using predicted cell counts. While the proposed approaches demonstrate their usefulness for AI-assisted diagnostics, the dataset will foster further research and development in the field, ultimately contributing to more precise diagnoses and improved patient outcomes.
Medical Image De-Identification Benchmark Challenge
Jul 31, 2025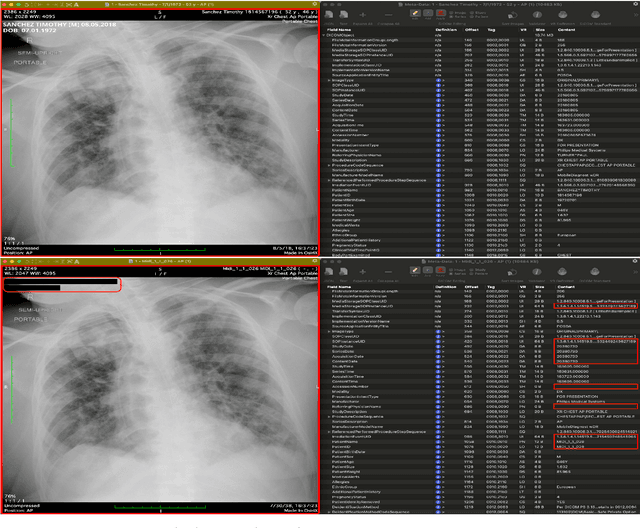
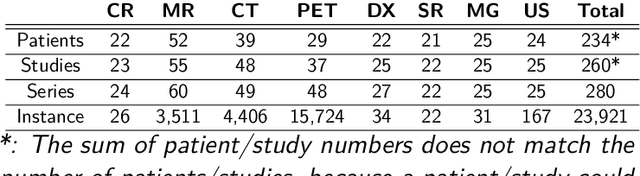
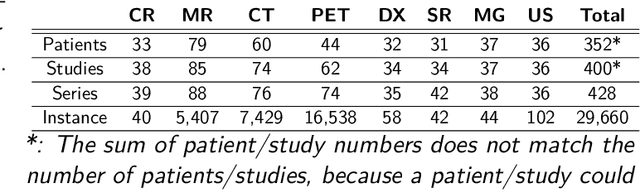
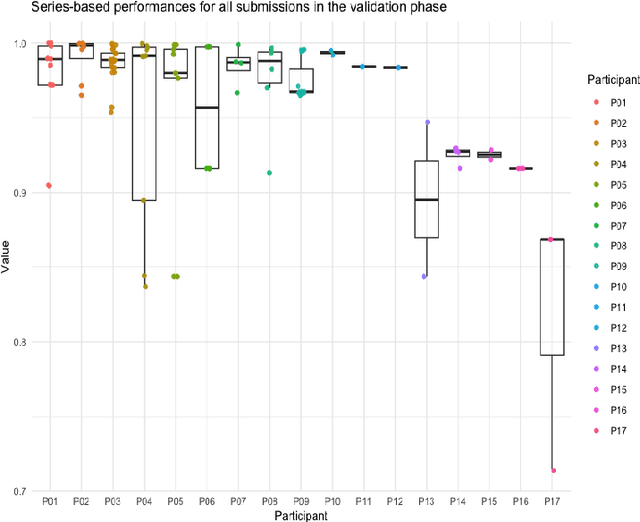
Abstract:The de-identification (deID) of protected health information (PHI) and personally identifiable information (PII) is a fundamental requirement for sharing medical images, particularly through public repositories, to ensure compliance with patient privacy laws. In addition, preservation of non-PHI metadata to inform and enable downstream development of imaging artificial intelligence (AI) is an important consideration in biomedical research. The goal of MIDI-B was to provide a standardized platform for benchmarking of DICOM image deID tools based on a set of rules conformant to the HIPAA Safe Harbor regulation, the DICOM Attribute Confidentiality Profiles, and best practices in preservation of research-critical metadata, as defined by The Cancer Imaging Archive (TCIA). The challenge employed a large, diverse, multi-center, and multi-modality set of real de-identified radiology images with synthetic PHI/PII inserted. The MIDI-B Challenge consisted of three phases: training, validation, and test. Eighty individuals registered for the challenge. In the training phase, we encouraged participants to tune their algorithms using their in-house or public data. The validation and test phases utilized the DICOM images containing synthetic identifiers (of 216 and 322 subjects, respectively). Ten teams successfully completed the test phase of the challenge. To measure success of a rule-based approach to image deID, scores were computed as the percentage of correct actions from the total number of required actions. The scores ranged from 97.91% to 99.93%. Participants employed a variety of open-source and proprietary tools with customized configurations, large language models, and optical character recognition (OCR). In this paper we provide a comprehensive report on the MIDI-B Challenge's design, implementation, results, and lessons learned.
Report of the Medical Image De-Identification Task Group -- Best Practices and Recommendations
Apr 01, 2023Abstract:This report addresses the technical aspects of de-identification of medical images of human subjects and biospecimens, such that re-identification risk of ethical, moral, and legal concern is sufficiently reduced to allow unrestricted public sharing for any purpose, regardless of the jurisdiction of the source and distribution sites. All medical images, regardless of the mode of acquisition, are considered, though the primary emphasis is on those with accompanying data elements, especially those encoded in formats in which the data elements are embedded, particularly Digital Imaging and Communications in Medicine (DICOM). These images include image-like objects such as Segmentations, Parametric Maps, and Radiotherapy (RT) Dose objects. The scope also includes related non-image objects, such as RT Structure Sets, Plans and Dose Volume Histograms, Structured Reports, and Presentation States. Only de-identification of publicly released data is considered, and alternative approaches to privacy preservation, such as federated learning for artificial intelligence (AI) model development, are out of scope, as are issues of privacy leakage from AI model sharing. Only technical issues of public sharing are addressed.
The NCI Imaging Data Commons as a platform for reproducible research in computational pathology
Mar 16, 2023



Abstract:Objective: Reproducibility is critical for translating machine learning-based (ML) solutions in computational pathology (CompPath) into practice. However, an increasing number of studies report difficulties in reproducing ML results. The NCI Imaging Data Commons (IDC) is a public repository of >120 cancer image collections, including >38,000 whole-slide images (WSIs), that is designed to be used with cloud-based ML services. Here, we explore the potential of the IDC to facilitate reproducibility of CompPath research. Materials and Methods: The IDC realizes the FAIR principles: All images are encoded according to the DICOM standard, persistently identified, discoverable via rich metadata, and accessible via open tools. Taking advantage of this, we implemented two experiments in which a representative ML-based method for classifying lung tumor tissue was trained and/or evaluated on different datasets from the IDC. To assess reproducibility, the experiments were run multiple times with independent but identically configured sessions of common ML services. Results: The AUC values of different runs of the same experiment were generally consistent and in the same order of magnitude as a similar, previously published study. However, there were occasional small variations in AUC values of up to 0.044, indicating a practical limit to reproducibility. Discussion and conclusion: By realizing the FAIR principles, the IDC enables other researchers to reuse exactly the same datasets. Cloud-based ML services enable others to run CompPath experiments in an identically configured computing environment without having to own high-performance hardware. The combination of both makes it possible to approach the reproducibility limit.
Highdicom: A Python library for standardized encoding of image annotations and machine learning model outputs in pathology and radiology
Jun 14, 2021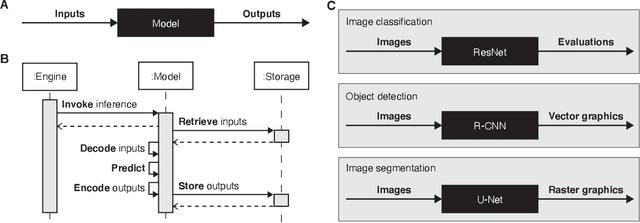
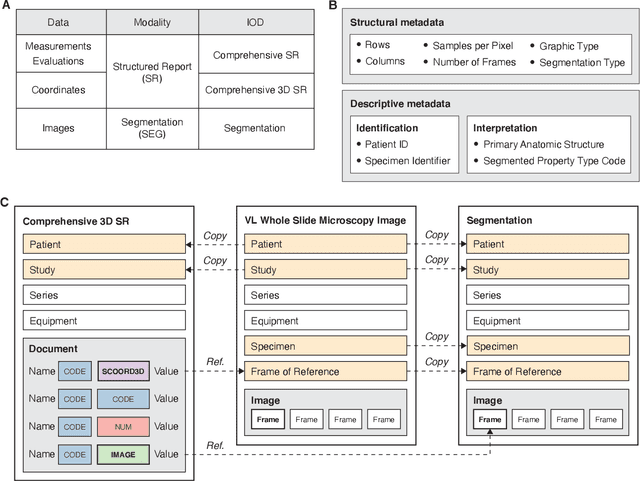
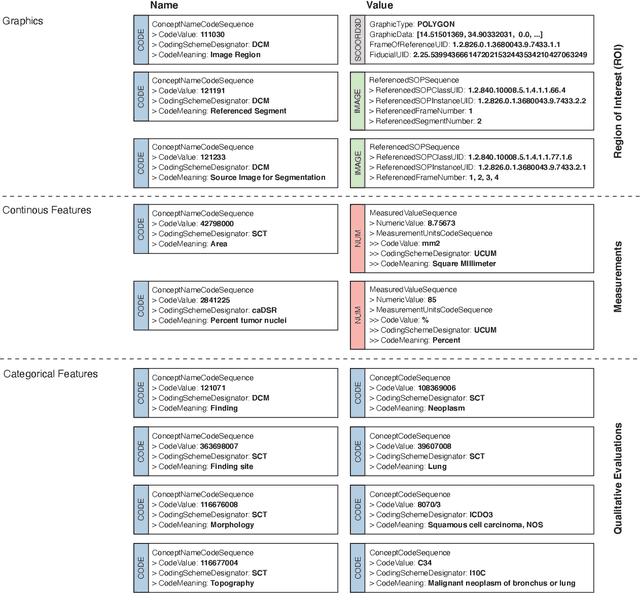
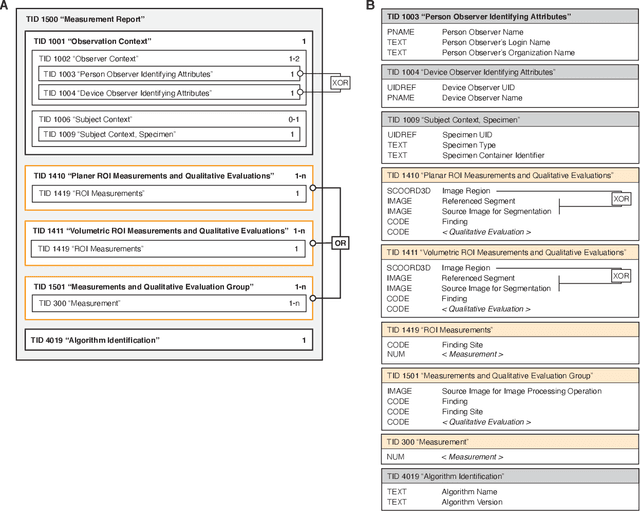
Abstract:Machine learning is revolutionizing image-based diagnostics in pathology and radiology. ML models have shown promising results in research settings, but their lack of interoperability has been a major barrier for clinical integration and evaluation. The DICOM a standard specifies Information Object Definitions and Services for the representation and communication of digital images and related information, including image-derived annotations and analysis results. However, the complexity of the standard represents an obstacle for its adoption in the ML community and creates a need for software libraries and tools that simplify working with data sets in DICOM format. Here we present the highdicom library, which provides a high-level application programming interface for the Python programming language that abstracts low-level details of the standard and enables encoding and decoding of image-derived information in DICOM format in a few lines of Python code. The highdicom library ties into the extensive Python ecosystem for image processing and machine learning. Simultaneously, by simplifying creation and parsing of DICOM-compliant files, highdicom achieves interoperability with the medical imaging systems that hold the data used to train and run ML models, and ultimately communicate and store model outputs for clinical use. We demonstrate through experiments with slide microscopy and computed tomography imaging, that, by bridging these two ecosystems, highdicom enables developers to train and evaluate state-of-the-art ML models in pathology and radiology while remaining compliant with the DICOM standard and interoperable with clinical systems at all stages. To promote standardization of ML research and streamline the ML model development and deployment process, we made the library available free and open-source.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge