Maria Valdés Hernández
Comparative evaluation of training strategies using partially labelled datasets for segmentation of white matter hyperintensities and stroke lesions in FLAIR MRI
Jan 28, 2026Abstract:White matter hyperintensities (WMH) and ischaemic stroke lesions (ISL) are imaging features associated with cerebral small vessel disease (SVD) that are visible on brain magnetic resonance imaging (MRI) scans. The development and validation of deep learning models to segment and differentiate these features is difficult because they visually confound each other in the fluid-attenuated inversion recovery (FLAIR) sequence and often appear in the same subject. We investigated six strategies for training a combined WMH and ISL segmentation model using partially labelled data. We combined privately held fully and partially labelled datasets with publicly available partially labelled datasets to yield a total of 2052 MRI volumes, with 1341 and 1152 containing ground truth annotations for WMH and ISL respectively. We found that several methods were able to effectively leverage the partially labelled data to improve model performance, with the use of pseudolabels yielding the best result.
Automated neuroradiological support systems for multiple cerebrovascular disease markers -- A systematic review and meta-analysis
Oct 22, 2024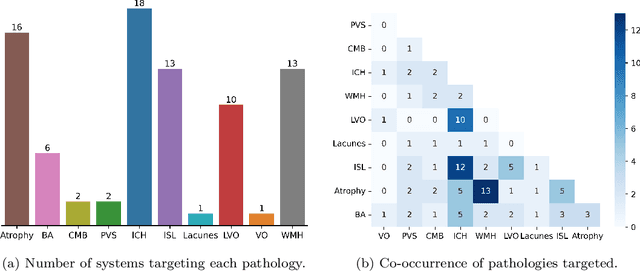
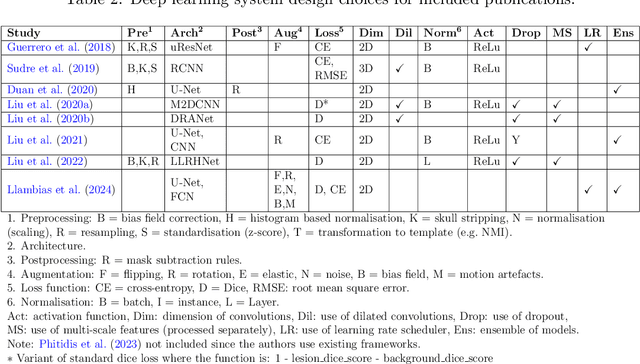
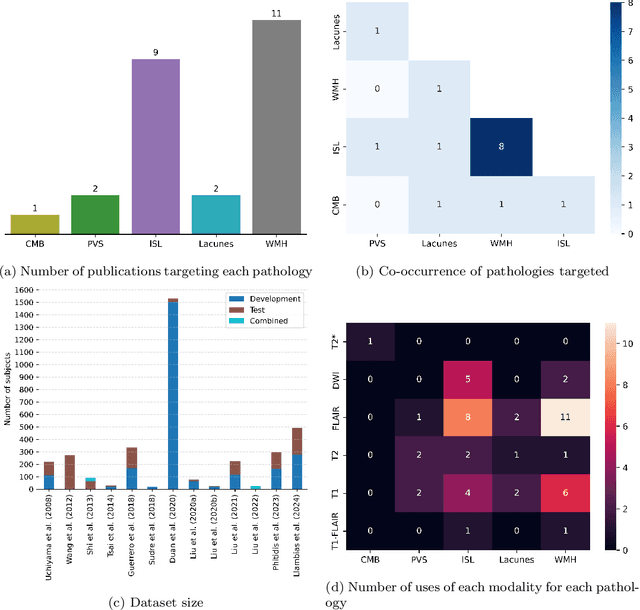
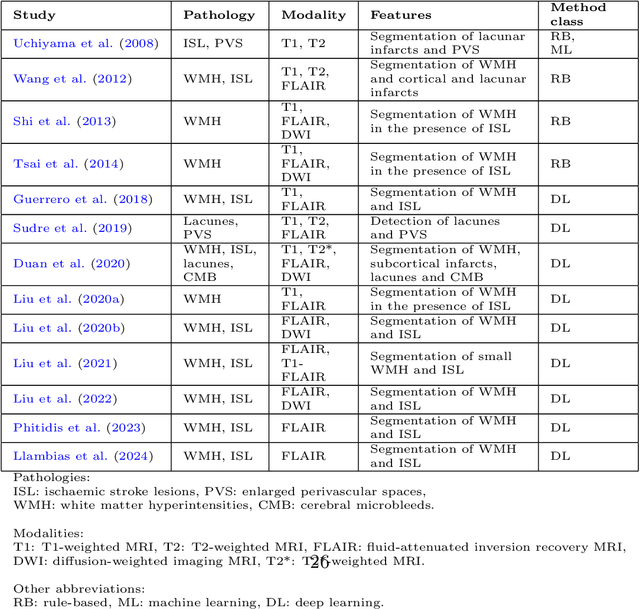
Abstract:Cerebrovascular diseases (CVD) can lead to stroke and dementia. Stroke is the second leading cause of death world wide and dementia incidence is increasing by the year. There are several markers of CVD that are visible on brain imaging, including: white matter hyperintensities (WMH), acute and chronic ischaemic stroke lesions (ISL), lacunes, enlarged perivascular spaces (PVS), acute and chronic haemorrhagic lesions, and cerebral microbleeds (CMB). Brain atrophy also occurs in CVD. These markers are important for patient management and intervention, since they indicate elevated risk of future stroke and dementia. We systematically reviewed automated systems designed to support radiologists reporting on these CVD imaging findings. We considered commercially available software and research publications which identify at least two CVD markers. In total, we included 29 commercial products and 13 research publications. Two distinct types of commercial support system were available: those which identify acute stroke lesions (haemorrhagic and ischaemic) from computed tomography (CT) scans, mainly for the purpose of patient triage; and those which measure WMH and atrophy regionally and longitudinally. In research, WMH and ISL were the markers most frequently analysed together, from magnetic resonance imaging (MRI) scans; lacunes and PVS were each targeted only twice and CMB only once. For stroke, commercially available systems largely support the emergency setting, whilst research systems consider also follow-up and routine scans. The systems to quantify WMH and atrophy are focused on neurodegenerative disease support, where these CVD markers are also of significance. There are currently no openly validated systems, commercially, or in research, performing a comprehensive joint analysis of all CVD markers (WMH, ISL, lacunes, PVS, haemorrhagic lesions, CMB, and atrophy).
GAN Augmentation: Augmenting Training Data using Generative Adversarial Networks
Oct 25, 2018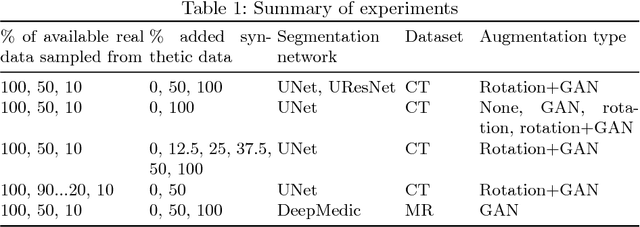
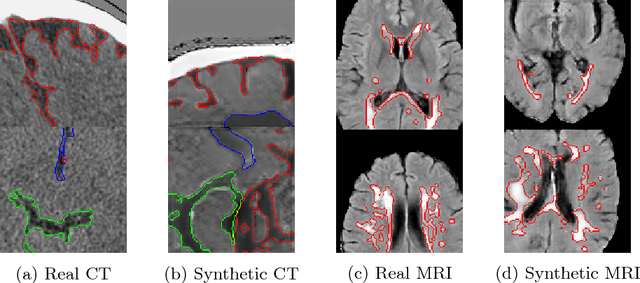
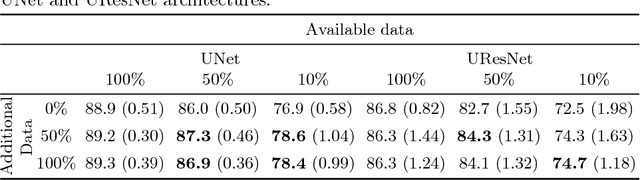
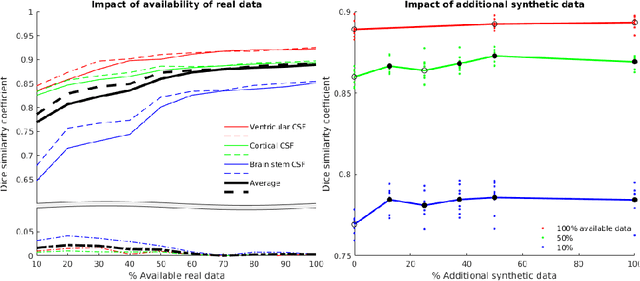
Abstract:One of the biggest issues facing the use of machine learning in medical imaging is the lack of availability of large, labelled datasets. The annotation of medical images is not only expensive and time consuming but also highly dependent on the availability of expert observers. The limited amount of training data can inhibit the performance of supervised machine learning algorithms which often need very large quantities of data on which to train to avoid overfitting. So far, much effort has been directed at extracting as much information as possible from what data is available. Generative Adversarial Networks (GANs) offer a novel way to unlock additional information from a dataset by generating synthetic samples with the appearance of real images. This paper demonstrates the feasibility of introducing GAN derived synthetic data to the training datasets in two brain segmentation tasks, leading to improvements in Dice Similarity Coefficient (DSC) of between 1 and 5 percentage points under different conditions, with the strongest effects seen fewer than ten training image stacks are available.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge