Long Xie
for the Alzheimer's Disease Neuroimaging Initiative
Development of an isotropic segmentation model for medial temporal lobe subregions on anisotropic MRI atlas using implicit neural representation
Aug 24, 2025

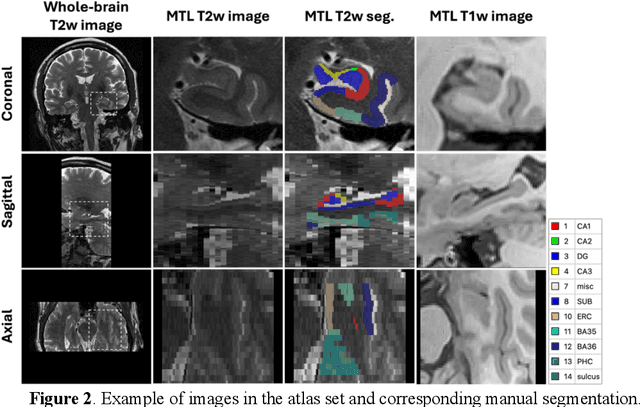
Abstract:Imaging biomarkers in magnetic resonance imaging (MRI) are important tools for diagnosing and tracking Alzheimer's disease (AD). As medial temporal lobe (MTL) is the earliest region to show AD-related hallmarks, brain atrophy caused by AD can first be observed in the MTL. Accurate segmentation of MTL subregions and extraction of imaging biomarkers from them are important. However, due to imaging limitations, the resolution of T2-weighted (T2w) MRI is anisotropic, which makes it difficult to accurately extract the thickness of cortical subregions in the MTL. In this study, we used an implicit neural representation method to combine the resolution advantages of T1-weighted and T2w MRI to accurately upsample an MTL subregion atlas set from anisotropic space to isotropic space, establishing a multi-modality, high-resolution atlas set. Based on this atlas, we developed an isotropic MTL subregion segmentation model. In an independent test set, the cortical subregion thickness extracted using this isotropic model showed higher significance than an anisotropic method in distinguishing between participants with mild cognitive impairment and cognitively unimpaired (CU) participants. In longitudinal analysis, the biomarkers extracted using isotropic method showed greater stability in CU participants. This study improved the accuracy of AD imaging biomarkers without increasing the amount of atlas annotation work, which may help to more accurately quantify the relationship between AD and brain atrophy and provide more accurate measures for disease tracking.
Nearly isotropic segmentation for medial temporal lobe subregions in multi-modality MRI
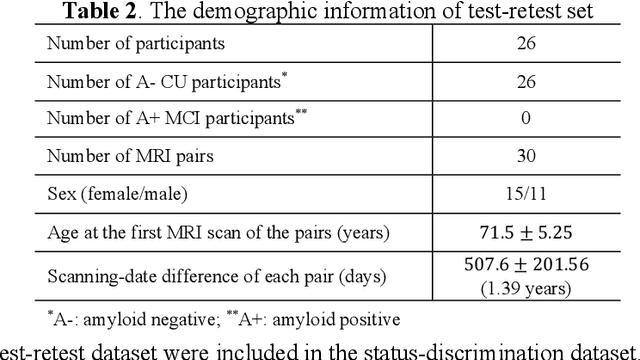
Apr 25, 2025Abstract:Morphometry of medial temporal lobe (MTL) subregions in brain MRI is sensitive biomarker to Alzheimers Disease and other related conditions. While T2-weighted (T2w) MRI with high in-plane resolution is widely used to segment hippocampal subfields due to its higher contrast in hippocampus, its lower out-of-plane resolution reduces the accuracy of subregion thickness measurements. To address this issue, we developed a nearly isotropic segmentation pipeline that incorporates image and label upsampling and high-resolution segmentation in T2w MRI. First, a high-resolution atlas was created based on an existing anisotropic atlas derived from 29 individuals. Both T1-weighted and T2w images in the atlas were upsampled from their original resolution to a nearly isotropic resolution 0.4x0.4x0.52mm3 using a non-local means approach. Manual segmentations within the atlas were also upsampled to match this resolution using a UNet-based neural network, which was trained on a cohort consisting of both high-resolution ex vivo and low-resolution anisotropic in vivo MRI with manual segmentations. Second, a multi-modality deep learning-based segmentation model was trained within this nearly isotropic atlas. Finally, experiments showed the nearly isotropic subregion segmentation improved the accuracy of cortical thickness as an imaging biomarker for neurodegeneration in T2w MRI.
Regional Deep Atrophy: a Self-Supervised Learning Method to Automatically Identify Regions Associated With Alzheimer's Disease Progression From Longitudinal MRI
Apr 10, 2023Abstract:Longitudinal assessment of brain atrophy, particularly in the hippocampus, is a well-studied biomarker for neurodegenerative diseases, such as Alzheimer's disease (AD). In clinical trials, estimation of brain progressive rates can be applied to track therapeutic efficacy of disease modifying treatments. However, most state-of-the-art measurements calculate changes directly by segmentation and/or deformable registration of MRI images, and may misreport head motion or MRI artifacts as neurodegeneration, impacting their accuracy. In our previous study, we developed a deep learning method DeepAtrophy that uses a convolutional neural network to quantify differences between longitudinal MRI scan pairs that are associated with time. DeepAtrophy has high accuracy in inferring temporal information from longitudinal MRI scans, such as temporal order or relative inter-scan interval. DeepAtrophy also provides an overall atrophy score that was shown to perform well as a potential biomarker of disease progression and treatment efficacy. However, DeepAtrophy is not interpretable, and it is unclear what changes in the MRI contribute to progression measurements. In this paper, we propose Regional Deep Atrophy (RDA), which combines the temporal inference approach from DeepAtrophy with a deformable registration neural network and attention mechanism that highlights regions in the MRI image where longitudinal changes are contributing to temporal inference. RDA has similar prediction accuracy as DeepAtrophy, but its additional interpretability makes it more acceptable for use in clinical settings, and may lead to more sensitive biomarkers for disease monitoring in clinical trials of early AD.
Improved Segmentation of Deep Sulci in Cortical Gray Matter Using a Deep Learning Framework Incorporating Laplace's Equation
Mar 03, 2023



Abstract:When developing tools for automated cortical segmentation, the ability to produce topologically correct segmentations is important in order to compute geometrically valid morphometry measures. In practice, accurate cortical segmentation is challenged by image artifacts and the highly convoluted anatomy of the cortex itself. To address this, we propose a novel deep learning-based cortical segmentation method in which prior knowledge about the geometry of the cortex is incorporated into the network during the training process. We design a loss function which uses the theory of Laplace's equation applied to the cortex to locally penalize unresolved boundaries between tightly folded sulci. Using an ex vivo MRI dataset of human medial temporal lobe specimens, we demonstrate that our approach outperforms baseline segmentation networks, both quantitatively and qualitatively.
Deep Label Fusion: A 3D End-to-End Hybrid Multi-Atlas Segmentation and Deep Learning Pipeline
Mar 19, 2021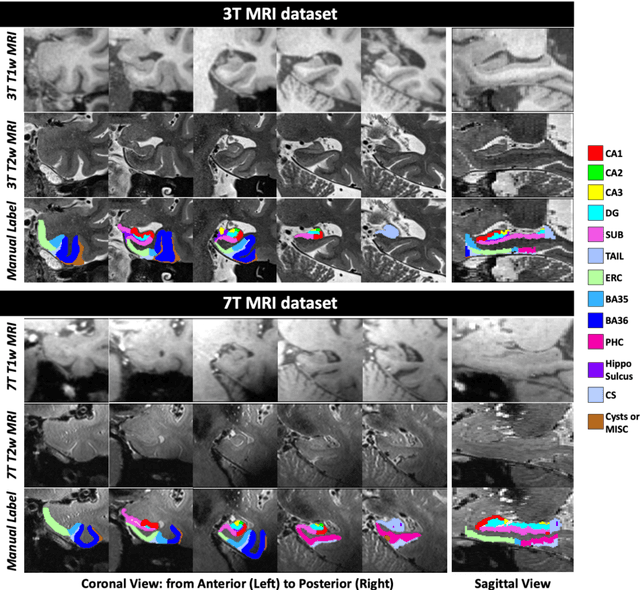
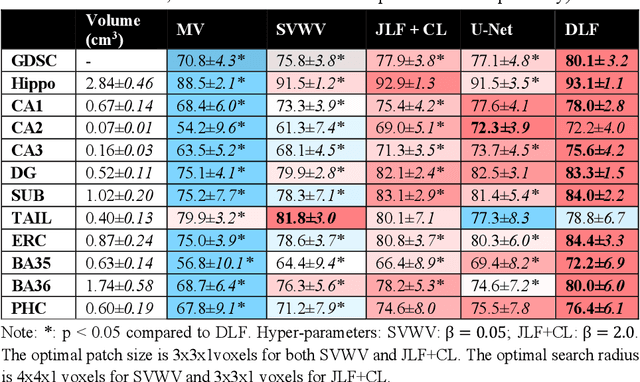
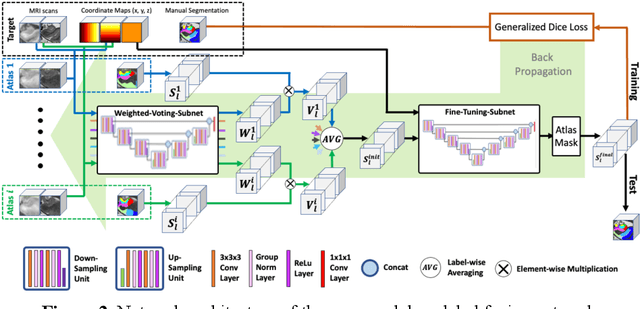

Abstract:Deep learning (DL) is the state-of-the-art methodology in various medical image segmentation tasks. However, it requires relatively large amounts of manually labeled training data, which may be infeasible to generate in some applications. In addition, DL methods have relatively poor generalizability to out-of-sample data. Multi-atlas segmentation (MAS), on the other hand, has promising performance using limited amounts of training data and good generalizability. A hybrid method that integrates the high accuracy of DL and good generalizability of MAS is highly desired and could play an important role in segmentation problems where manually labeled data is hard to generate. Most of the prior work focuses on improving single components of MAS using DL rather than directly optimizing the final segmentation accuracy via an end-to-end pipeline. Only one study explored this idea in binary segmentation of 2D images, but it remains unknown whether it generalizes well to multi-class 3D segmentation problems. In this study, we propose a 3D end-to-end hybrid pipeline, named deep label fusion (DLF), that takes advantage of the strengths of MAS and DL. Experimental results demonstrate that DLF yields significant improvements over conventional label fusion methods and U-Net, a direct DL approach, in the context of segmenting medial temporal lobe subregions using 3T T1-weighted and T2-weighted MRI. Further, when applied to an unseen similar dataset acquired in 7T, DLF maintains its superior performance, which demonstrates its good generalizability.
DeepAtrophy: Teaching a Neural Network to Differentiate Progressive Changes from Noise on Longitudinal MRI in Alzheimer's Disease
Oct 24, 2020



Abstract:Volume change measures derived from longitudinal MRI (e.g. hippocampal atrophy) are a well-studied biomarker of disease progression in Alzheimer's Disease (AD) and are used in clinical trials to track the therapeutic efficacy of disease-modifying treatments. However, longitudinal MRI change measures can be confounded by non-biological factors, such as different degrees of head motion and susceptibility artifact between pairs of MRI scans. We hypothesize that deep learning methods applied directly to pairs of longitudinal MRI scans can be trained to differentiate between biological changes and non-biological factors better than conventional approaches based on deformable image registration. To achieve this, we make a simplifying assumption that biological factors are associated with time (i.e. the hippocampus shrinks overtime in the aging population) whereas non-biological factors are independent of time. We then formulate deep learning networks to infer the temporal order of same-subject MRI scans input to the network in arbitrary order; as well as to infer ratios between interscan intervals for two pairs of same-subject MRI scans. In the test dataset, these networks perform better in tasks of temporal ordering (89.3%) and interscan interval inference (86.1%) than a state-of-the-art deformation-based morphometry method ALOHA (76.6% and 76.1% respectively) (Das et al., 2012). Furthermore, we derive a disease progression score from the network that is able to detect a group difference between 58 preclinical AD and 75 beta-amyloid-negative cognitively normal individuals within one year, compared to two years for ALOHA. This suggests that deep learning can be trained to differentiate MRI changes due to biological factors (tissue loss) from changes due to non-biological factors, leading to novel biomarkers that are more sensitive to longitudinal changes at the earliest stages of AD.
Barnes-Hut Approximation for Point SetGeodesic Shooting
Jul 10, 2019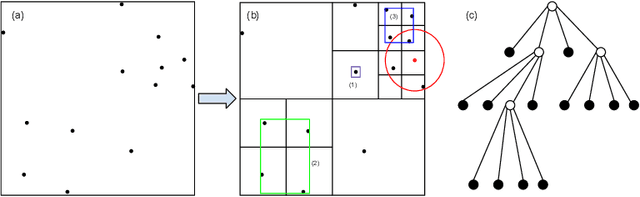

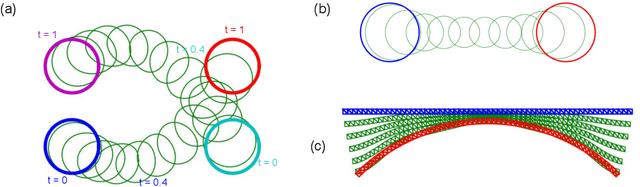

Abstract:Geodesic shooting has been successfully applied to diffeo-morphic registration of point sets. Exact computation of the geodesicshooting between point sets, however, requiresO(N2) calculations each time step on the number of points in the point set. We proposean approximation approach based on the Barnes-Hut algorithm to speedup point set geodesic shooting. This approximation can reduce the al-gorithm complexity toO(N b+N logN). The evaluation of the proposedmethod in both simulated images and the medial temporal lobe thick-ness analysis demonstrates a comparable accuracy to the exact point set geodesic shooting while offering up to 3-fold speed up. This improvementopens up a range of clinical research studies and practical problems towhich the method can be effectively applied.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge