Sydney Lim
Automated deep learning segmentation of high-resolution 7 T ex vivo MRI for quantitative analysis of structure-pathology correlations in neurodegenerative diseases
Mar 21, 2023



Abstract:Ex vivo MRI of the brain provides remarkable advantages over in vivo MRI for visualizing and characterizing detailed neuroanatomy, and helps to link microscale histology studies with morphometric measurements. However, automated segmentation methods for brain mapping in ex vivo MRI are not well developed, primarily due to limited availability of labeled datasets, and heterogeneity in scanner hardware and acquisition protocols. In this work, we present a high resolution dataset of 37 ex vivo post-mortem human brain tissue specimens scanned on a 7T whole-body MRI scanner. We developed a deep learning pipeline to segment the cortical mantle by benchmarking the performance of nine deep neural architectures. We then segment the four subcortical structures: caudate, putamen, globus pallidus, and thalamus; white matter hyperintensities, and the normal appearing white matter. We show excellent generalizing capabilities across whole brain hemispheres in different specimens, and also on unseen images acquired at different magnetic field strengths and different imaging sequence. We then compute volumetric and localized cortical thickness measurements across key regions, and link them with semi-quantitative neuropathological ratings. Our code, containerized executables, and the processed datasets are publicly available at: https://github.com/Pulkit-Khandelwal/upenn-picsl-brain-ex-vivo.
Improved Segmentation of Deep Sulci in Cortical Gray Matter Using a Deep Learning Framework Incorporating Laplace's Equation
Mar 03, 2023



Abstract:When developing tools for automated cortical segmentation, the ability to produce topologically correct segmentations is important in order to compute geometrically valid morphometry measures. In practice, accurate cortical segmentation is challenged by image artifacts and the highly convoluted anatomy of the cortex itself. To address this, we propose a novel deep learning-based cortical segmentation method in which prior knowledge about the geometry of the cortex is incorporated into the network during the training process. We design a loss function which uses the theory of Laplace's equation applied to the cortex to locally penalize unresolved boundaries between tightly folded sulci. Using an ex vivo MRI dataset of human medial temporal lobe specimens, we demonstrate that our approach outperforms baseline segmentation networks, both quantitatively and qualitatively.
Gray Matter Segmentation in Ultra High Resolution 7 Tesla ex vivo T2w MRI of Human Brain Hemispheres
Oct 14, 2021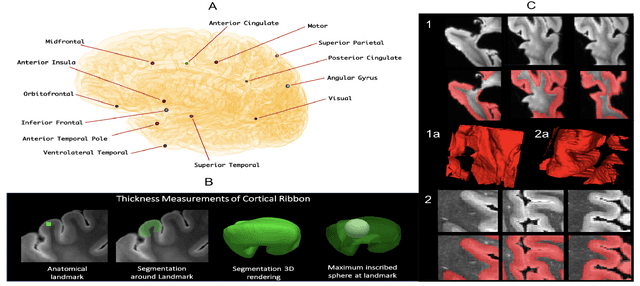
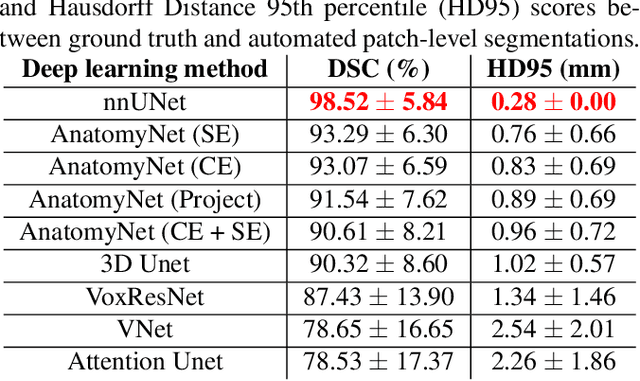
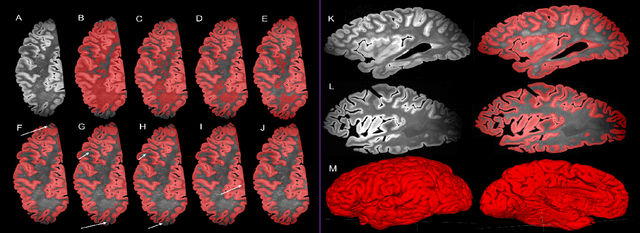
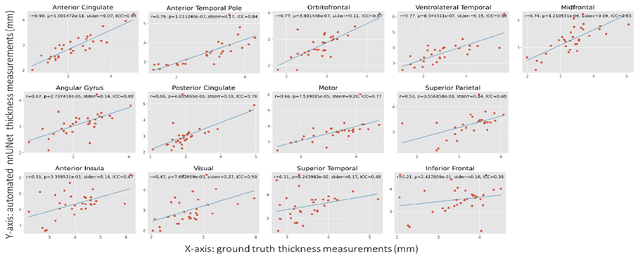
Abstract:Ex vivo MRI of the brain provides remarkable advantages over in vivo MRI for visualizing and characterizing detailed neuroanatomy. However, automated cortical segmentation methods in ex vivo MRI are not well developed, primarily due to limited availability of labeled datasets, and heterogeneity in scanner hardware and acquisition protocols. In this work, we present a high resolution 7 Tesla dataset of 32 ex vivo human brain specimens. We benchmark the cortical mantle segmentation performance of nine neural network architectures, trained and evaluated using manually-segmented 3D patches sampled from specific cortical regions, and show excellent generalizing capabilities across whole brain hemispheres in different specimens, and also on unseen images acquired at different magnetic field strength and imaging sequences. Finally, we provide cortical thickness measurements across key regions in 3D ex vivo human brain images. Our code and processed datasets are publicly available at https://github.com/Pulkit-Khandelwal/picsl-ex-vivo-segmentation.
Deep Label Fusion: A 3D End-to-End Hybrid Multi-Atlas Segmentation and Deep Learning Pipeline
Mar 19, 2021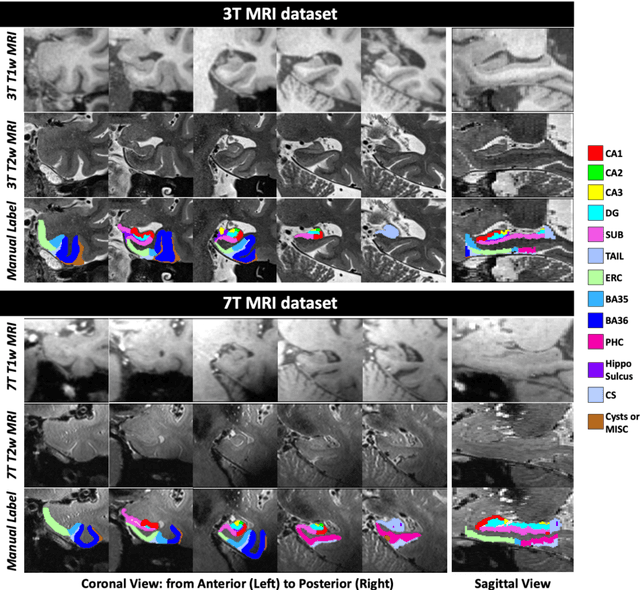
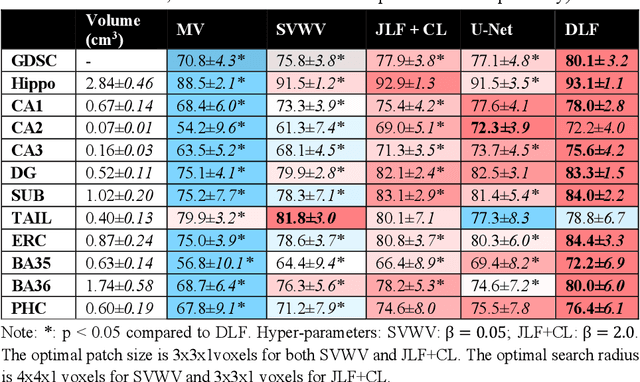
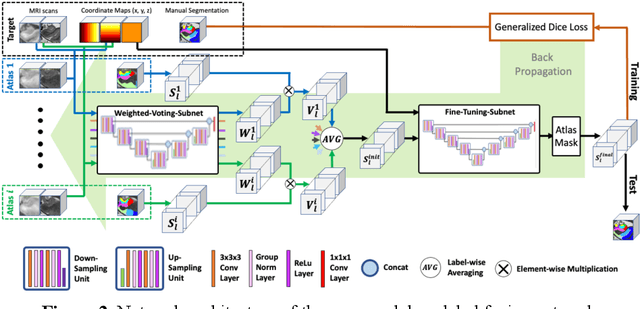

Abstract:Deep learning (DL) is the state-of-the-art methodology in various medical image segmentation tasks. However, it requires relatively large amounts of manually labeled training data, which may be infeasible to generate in some applications. In addition, DL methods have relatively poor generalizability to out-of-sample data. Multi-atlas segmentation (MAS), on the other hand, has promising performance using limited amounts of training data and good generalizability. A hybrid method that integrates the high accuracy of DL and good generalizability of MAS is highly desired and could play an important role in segmentation problems where manually labeled data is hard to generate. Most of the prior work focuses on improving single components of MAS using DL rather than directly optimizing the final segmentation accuracy via an end-to-end pipeline. Only one study explored this idea in binary segmentation of 2D images, but it remains unknown whether it generalizes well to multi-class 3D segmentation problems. In this study, we propose a 3D end-to-end hybrid pipeline, named deep label fusion (DLF), that takes advantage of the strengths of MAS and DL. Experimental results demonstrate that DLF yields significant improvements over conventional label fusion methods and U-Net, a direct DL approach, in the context of segmenting medial temporal lobe subregions using 3T T1-weighted and T2-weighted MRI. Further, when applied to an unseen similar dataset acquired in 7T, DLF maintains its superior performance, which demonstrates its good generalizability.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge