Lasse Lensu
Cross-modal learning for plankton recognition
Mar 17, 2026Abstract:This paper considers self-supervised cross-modal coordination as a strategy enabling utilization of multiple modalities and large volumes of unlabeled plankton data to build models for plankton recognition. Automated imaging instruments facilitate the continuous collection of plankton image data on a large scale. Current methods for automatic plankton image recognition rely primarily on supervised approaches, which require labeled training sets that are labor-intensive to collect. On the other hand, some modern plankton imaging instruments complement image information with optical measurement data, such as scatter and fluorescence profiles, which currently are not widely utilized in plankton recognition. In this work, we explore the possibility of using such measurement data to guide the learning process without requiring manual labeling. Inspired by the concepts behind Contrastive Language-Image Pre-training, we train encoders for both modalities using only binary supervisory information indicating whether a given image and profile originate from the same particle or from different particles. For plankton recognition, we employ a small labeled gallery of known plankton species combined with a $k$-NN classifier. This approach yields a recognition model that is inherently multimodal, i.e., capable of utilizing information extracted from both image and profile data. We demonstrate that the proposed method achieves high recognition accuracy while requiring only a minimal number of labeled images. Furthermore, we show that the approach outperforms an image-only self-supervised baseline. Code available at https://github.com/Jookare/cross-modal-plankton.
Self-Supervised Pretraining for Fine-Grained Plankton Recognition
Mar 14, 2025Abstract:Plankton recognition is an important computer vision problem due to plankton's essential role in ocean food webs and carbon capture, highlighting the need for species-level monitoring. However, this task is challenging due to its fine-grained nature and dataset shifts caused by different imaging instruments and varying species distributions. As new plankton image datasets are collected at an increasing pace, there is a need for general plankton recognition models that require minimal expert effort for data labeling. In this work, we study large-scale self-supervised pretraining for fine-grained plankton recognition. We first employ masked autoencoding and a large volume of diverse plankton image data to pretrain a general-purpose plankton image encoder. Then we utilize fine-tuning to obtain accurate plankton recognition models for new datasets with a very limited number of labeled training images. Our experiments show that self-supervised pretraining with diverse plankton data clearly increases plankton recognition accuracy compared to standard ImageNet pretraining when the amount of training data is limited. Moreover, the accuracy can be further improved when unlabeled target data is available and utilized during the pretraining.
Open-Set Plankton Recognition
Mar 14, 2025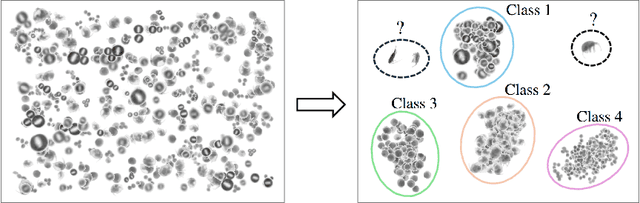
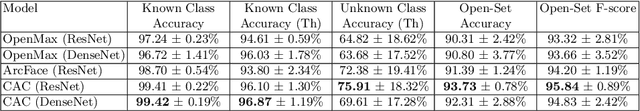
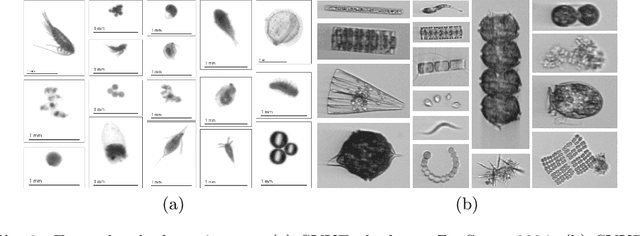
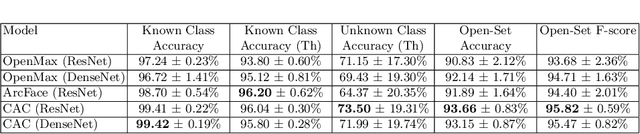
Abstract:This paper considers open-set recognition (OSR) of plankton images. Plankton include a diverse range of microscopic aquatic organisms that have an important role in marine ecosystems as primary producers and as a base of food webs. Given their sensitivity to environmental changes, fluctuations in plankton populations offer valuable information about oceans' health and climate change motivating their monitoring. Modern automatic plankton imaging devices enable the collection of large-scale plankton image datasets, facilitating species-level analysis. Plankton species recognition can be seen as an image classification task and is typically solved using deep learning-based image recognition models. However, data collection in real aquatic environments results in imaging devices capturing a variety of non-plankton particles and plankton species not present in the training set. This creates a challenging fine-grained OSR problem, characterized by subtle differences between taxonomically close plankton species. We address this challenge by conducting extensive experiments on three OSR approaches using both phyto- and zooplankton images analyzing also on the effect of the rejection thresholds for OSR. The results demonstrate that high OSR accuracy can be obtained promoting the use of these methods in operational plankton research. We have made the data publicly available to the research community.
DAPlankton: Benchmark Dataset for Multi-instrument Plankton Recognition via Fine-grained Domain Adaptation
Feb 08, 2024



Abstract:Plankton recognition provides novel possibilities to study various environmental aspects and an interesting real-world context to develop domain adaptation (DA) methods. Different imaging instruments cause domain shift between datasets hampering the development of general plankton recognition methods. A promising remedy for this is DA allowing to adapt a model trained on one instrument to other instruments. In this paper, we present a new DA dataset called DAPlankton which consists of phytoplankton images obtained with different instruments. Phytoplankton provides a challenging DA problem due to the fine-grained nature of the task and high class imbalance in real-world datasets. DAPlankton consists of two subsets. DAPlankton_LAB contains images of cultured phytoplankton providing a balanced dataset with minimal label uncertainty. DAPlankton_SEA consists of images collected from the Baltic Sea providing challenging real-world data with large intra-class variance and class imbalance. We further present a benchmark comparison of three widely used DA methods.
Log-Gaussian Gamma Processes for Training Bayesian Neural Networks in Raman and CARS Spectroscopies
Oct 12, 2023



Abstract:We propose an approach utilizing gamma-distributed random variables, coupled with log-Gaussian modeling, to generate synthetic datasets suitable for training neural networks. This addresses the challenge of limited real observations in various applications. We apply this methodology to both Raman and coherent anti-Stokes Raman scattering (CARS) spectra, using experimental spectra to estimate gamma process parameters. Parameter estimation is performed using Markov chain Monte Carlo methods, yielding a full Bayesian posterior distribution for the model which can be sampled for synthetic data generation. Additionally, we model the additive and multiplicative background functions for Raman and CARS with Gaussian processes. We train two Bayesian neural networks to estimate parameters of the gamma process which can then be used to estimate the underlying Raman spectrum and simultaneously provide uncertainty through the estimation of parameters of a probability distribution. We apply the trained Bayesian neural networks to experimental Raman spectra of phthalocyanine blue, aniline black, naphthol red, and red 264 pigments and also to experimental CARS spectra of adenosine phosphate, fructose, glucose, and sucrose. The results agree with deterministic point estimates for the underlying Raman and CARS spectral signatures.
Survey of Automatic Plankton Image Recognition: Challenges, Existing Solutions and Future Perspectives
May 19, 2023



Abstract:Planktonic organisms are key components of aquatic ecosystems and respond quickly to changes in the environment, therefore their monitoring is vital to understand the changes in the environment. Yet, monitoring plankton at appropriate scales still remains a challenge, limiting our understanding of functioning of aquatic systems and their response to changes. Modern plankton imaging instruments can be utilized to sample at high frequencies, enabling novel possibilities to study plankton populations. However, manual analysis of the data is costly, time consuming and expert based, making such approach unsuitable for large-scale application and urging for automatic solutions. The key problem related to the utilization of plankton datasets through image analysis is plankton recognition. Despite the large amount of research done, automatic methods have not been widely adopted for operational use. In this paper, a comprehensive survey on existing solutions for automatic plankton recognition is presented. First, we identify the most notable challenges that that make the development of plankton recognition systems difficult. Then, we provide a detailed description of solutions for these challenges proposed in plankton recognition literature. Finally, we propose a workflow to identify the specific challenges in new datasets and the recommended approaches to address them. For many of the challenges, applicable solutions exist. However, important challenges remain unsolved: 1) the domain shift between the datasets hindering the development of a general plankton recognition system that would work across different imaging instruments, 2) the difficulty to identify and process the images of previously unseen classes, and 3) the uncertainty in expert annotations that affects the training of the machine learning models for recognition. These challenges should be addressed in the future research.
Towards Phytoplankton Parasite Detection Using Autoencoders
Mar 15, 2023Abstract:Phytoplankton parasites are largely understudied microbial components with a potentially significant ecological impact on phytoplankton bloom dynamics. To better understand their impact, we need improved detection methods to integrate phytoplankton parasite interactions in monitoring aquatic ecosystems. Automated imaging devices usually produce high amount of phytoplankton image data, while the occurrence of anomalous phytoplankton data is rare. Thus, we propose an unsupervised anomaly detection system based on the similarity of the original and autoencoder-reconstructed samples. With this approach, we were able to reach an overall F1 score of 0.75 in nine phytoplankton species, which could be further improved by species-specific fine-tuning. The proposed unsupervised approach was further compared with the supervised Faster R-CNN based object detector. With this supervised approach and the model trained on plankton species and anomalies, we were able to reach the highest F1 score of 0.86. However, the unsupervised approach is expected to be more universal as it can detect also unknown anomalies and it does not require any annotated anomalous data that may not be always available in sufficient quantities. Although other studies have dealt with plankton anomaly detection in terms of non-plankton particles, or air bubble detection, our paper is according to our best knowledge the first one which focuses on automated anomaly detection considering putative phytoplankton parasites or infections.
Semi-Supervised Domain Adaptation for Weakly Labeled Semantic Video Object Segmentation
Jun 07, 2016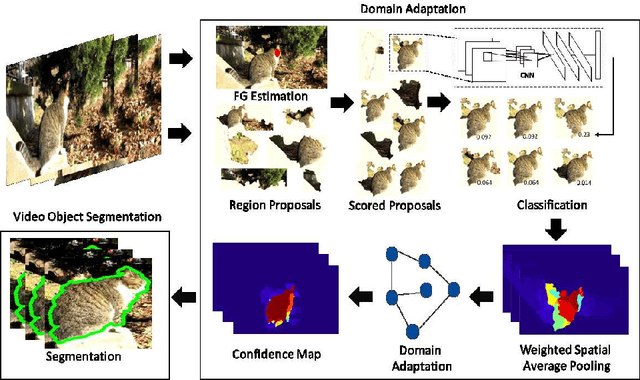
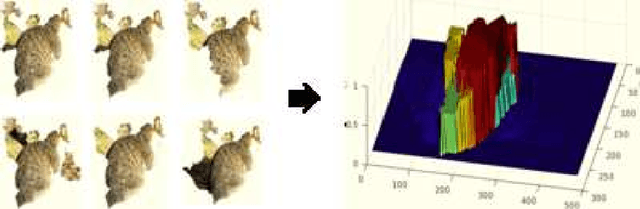
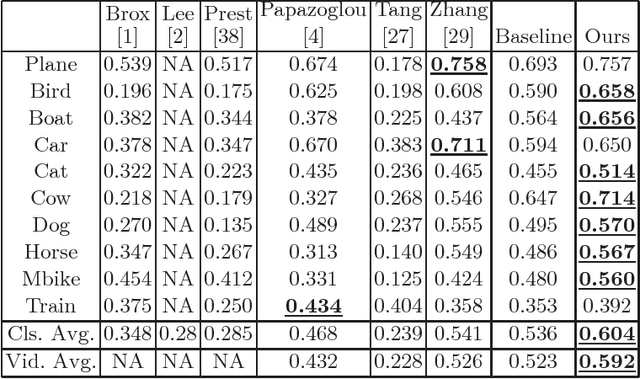


Abstract:Deep convolutional neural networks (CNNs) have been immensely successful in many high-level computer vision tasks given large labeled datasets. However, for video semantic object segmentation, a domain where labels are scarce, effectively exploiting the representation power of CNN with limited training data remains a challenge. Simply borrowing the existing pretrained CNN image recognition model for video segmentation task can severely hurt performance. We propose a semi-supervised approach to adapting CNN image recognition model trained from labeled image data to the target domain exploiting both semantic evidence learned from CNN, and the intrinsic structures of video data. By explicitly modeling and compensating for the domain shift from the source domain to the target domain, this proposed approach underpins a robust semantic object segmentation method against the changes in appearance, shape and occlusion in natural videos. We present extensive experiments on challenging datasets that demonstrate the superior performance of our approach compared with the state-of-the-art methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge