Kilian M Pohl
Diffusion Models for Computational Neuroimaging: A Survey
Feb 10, 2025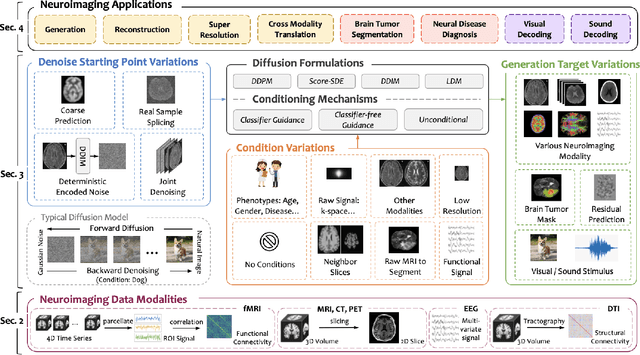

Abstract:Computational neuroimaging involves analyzing brain images or signals to provide mechanistic insights and predictive tools for human cognition and behavior. While diffusion models have shown stability and high-quality generation in natural images, there is increasing interest in adapting them to analyze brain data for various neurological tasks such as data enhancement, disease diagnosis and brain decoding. This survey provides an overview of recent efforts to integrate diffusion models into computational neuroimaging. We begin by introducing the common neuroimaging data modalities, follow with the diffusion formulations and conditioning mechanisms. Then we discuss how the variations of the denoising starting point, condition input and generation target of diffusion models are developed and enhance specific neuroimaging tasks. For a comprehensive overview of the ongoing research, we provide a publicly available repository at https://github.com/JoeZhao527/dm4neuro.
Self-Supervised Longitudinal Neighbourhood Embedding
Mar 09, 2021



Abstract:Longitudinal MRIs are often used to capture the gradual deterioration of brain structure and function caused by aging or neurological diseases. Analyzing this data via machine learning generally requires a large number of ground-truth labels, which are often missing or expensive to obtain. Reducing the need for labels, we propose a self-supervised strategy for representation learning named Longitudinal Neighborhood Embedding (LNE). Motivated by concepts in contrastive learning, LNE explicitly models the similarity between trajectory vectors across different subjects. We do so by building a graph in each training iteration defining neighborhoods in the latent space so that the progression direction of a subject follows the direction of its neighbors. This results in a smooth trajectory field that captures the global morphological change of the brain while maintaining the local continuity. We apply LNE to longitudinal T1w MRIs of two neuroimaging studies: a dataset composed of 274 healthy subjects, and Alzheimer's Disease Neuroimaging Initiative (ADNI, N=632). The visualization of the smooth trajectory vector field and superior performance on downstream tasks demonstrate the strength of the proposed method over existing self-supervised methods in extracting information associated with normal aging and in revealing the impact of neurodegenerative disorders. The code is available at \url{https://github.com/ouyangjiahong/longitudinal-neighbourhood-embedding.git}.
Representation Disentanglement for Multi-modal MR Analysis
Feb 23, 2021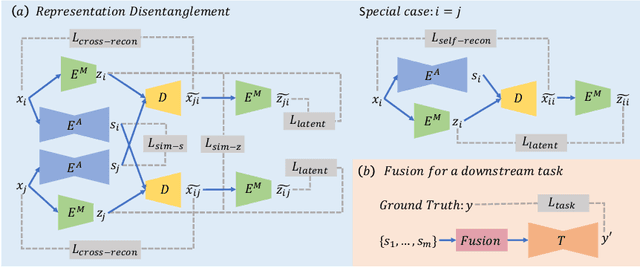
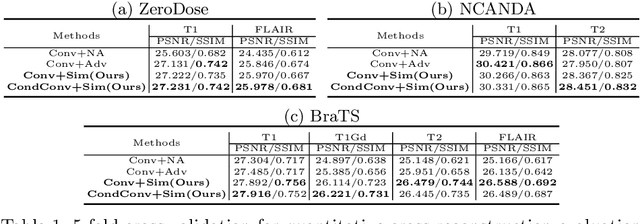
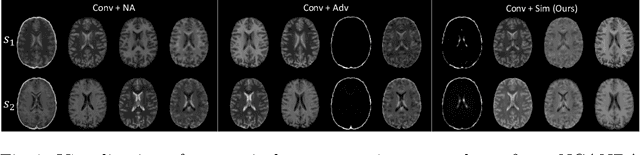
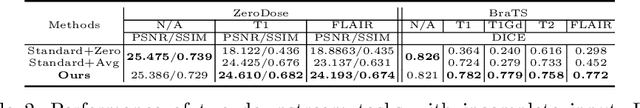
Abstract:Multi-modal MR images are widely used in neuroimaging applications to provide complementary information about the brain structures. Recent works have suggested that multi-modal deep learning analysis can benefit from explicitly disentangling anatomical (shape) and modality (appearance) representations from the images. In this work, we challenge existing strategies by showing that they do not naturally lead to representation disentanglement both in theory and in practice. To address this issue, we propose a margin loss that regularizes the similarity relationships of the representations across subjects and modalities. To enable a robust training, we further introduce a modified conditional convolution to design a single model for encoding images of all modalities. Lastly, we propose a fusion function to combine the disentangled anatomical representations as a set of modality-invariant features for downstream tasks. We evaluate the proposed method on three multi-modal neuroimaging datasets. Experiments show that our proposed method can achieve superior disentangled representations compared to existing disentanglement strategies. Results also indicate that the fused anatomical representation has great potential in the downstream task of zero-dose PET reconstruction and brain tumor segmentation.
Recurrent Neural Networks with Longitudinal Pooling and Consistency Regularization
Mar 31, 2020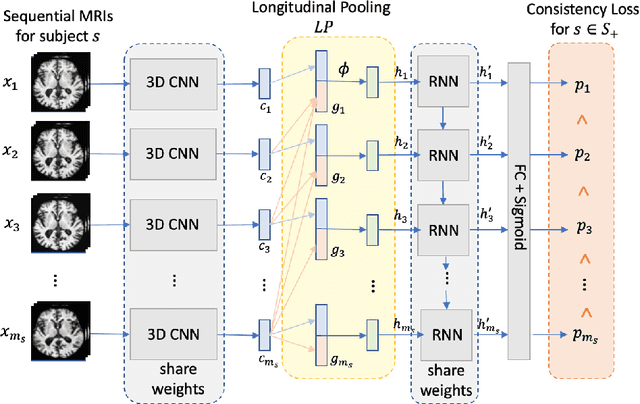
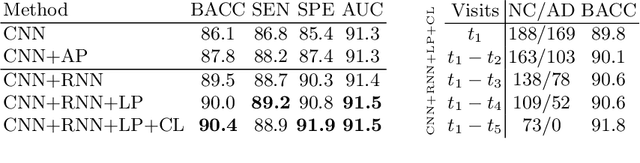
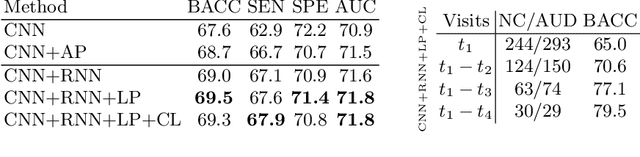

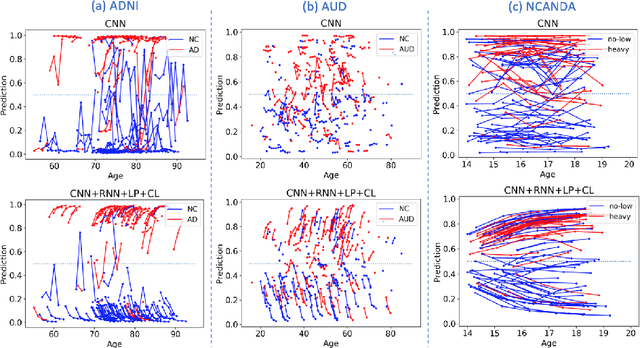

Abstract:Most neurological diseases are characterized by gradual deterioration of brain structure and function. To identify the impact of such diseases, studies have been acquiring large longitudinal MRI datasets and applied deep-learning to predict diagnosis label(s). These learning models apply Convolutional Neural Networks (CNN) to extract informative features from each time point of the longitudinal MRI and Recurrent Neural Networks (RNN) to classify each time point based on those features. However, they neglect the progressive nature of the disease, which may result in clinically implausible predictions across visits. In this paper, we propose a framework that injects the extracted features from CNNs at each time point to the RNN cells considering the dependencies across different time points in the longitudinal data. On the feature level, we propose a novel longitudinal pooling layer to couple features of a visit with those of proceeding ones. On the prediction level, we add a consistency regularization to the classification objective in line with the nature of the disease progression across visits. We evaluate the proposed method on the longitudinal structural MRIs from three neuroimaging datasets: Alzheimer's Disease Neuroimaging Initiative (ADNI, N=404), a dataset composed of 274 healthy controls and 329 patients with Alcohol Use Disorder (AUD), and 255 youths from the National Consortium on Alcohol and NeuroDevelopment in Adolescence (NCANDA). All three experiments show that our method is superior to the widely used methods. The code is available at https://github.com/ouyangjiahong/longitudinal-pooling.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge