Kieran Chin-Cheong
Automatic Classification of General Movements in Newborns
Nov 14, 2024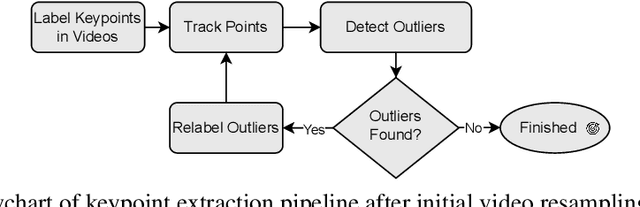
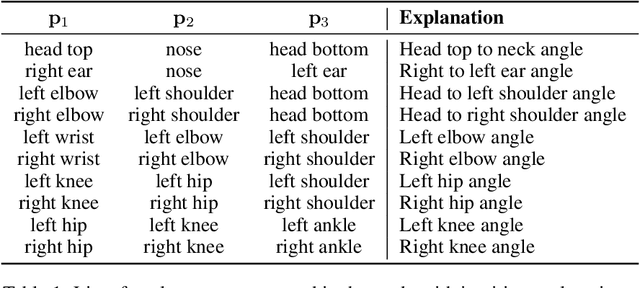

Abstract:General movements (GMs) are spontaneous, coordinated body movements in infants that offer valuable insights into the developing nervous system. Assessed through the Prechtl GM Assessment (GMA), GMs are reliable predictors for neurodevelopmental disorders. However, GMA requires specifically trained clinicians, who are limited in number. To scale up newborn screening, there is a need for an algorithm that can automatically classify GMs from infant video recordings. This data poses challenges, including variability in recording length, device type, and setting, with each video coarsely annotated for overall movement quality. In this work, we introduce a tool for extracting features from these recordings and explore various machine learning techniques for automated GM classification.
Interpretable and Intervenable Ultrasonography-based Machine Learning Models for Pediatric Appendicitis
Feb 28, 2023Abstract:Appendicitis is among the most frequent reasons for pediatric abdominal surgeries. With recent advances in machine learning, data-driven decision support could help clinicians diagnose and manage patients while reducing the number of non-critical surgeries. Previous decision support systems for appendicitis focused on clinical, laboratory, scoring and computed tomography data, mainly ignoring abdominal ultrasound, a noninvasive and readily available diagnostic modality. To this end, we developed and validated interpretable machine learning models for predicting the diagnosis, management and severity of suspected appendicitis using ultrasound images. Our models were trained on a dataset comprising 579 pediatric patients with 1709 ultrasound images accompanied by clinical and laboratory data. Our methodological contribution is the generalization of concept bottleneck models to prediction problems with multiple views and incomplete concept sets. Notably, such models lend themselves to interpretation and interaction via high-level concepts understandable to clinicians without sacrificing performance or requiring time-consuming image annotation when deployed.
On the Limitations of Multimodal VAEs
Oct 08, 2021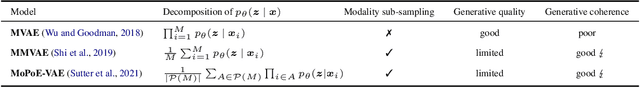
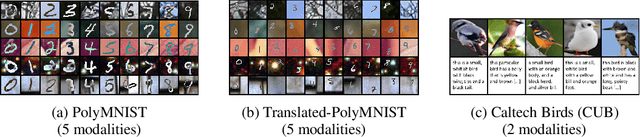

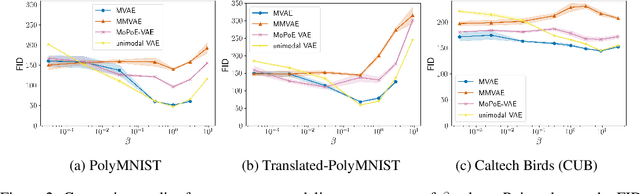
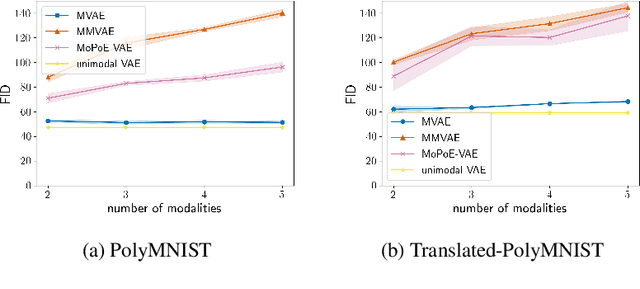
Abstract:Multimodal variational autoencoders (VAEs) have shown promise as efficient generative models for weakly-supervised data. Yet, despite their advantage of weak supervision, they exhibit a gap in generative quality compared to unimodal VAEs, which are completely unsupervised. In an attempt to explain this gap, we uncover a fundamental limitation that applies to a large family of mixture-based multimodal VAEs. We prove that the sub-sampling of modalities enforces an undesirable upper bound on the multimodal ELBO and thereby limits the generative quality of the respective models. Empirically, we showcase the generative quality gap on both synthetic and real data and present the tradeoffs between different variants of multimodal VAEs. We find that none of the existing approaches fulfills all desired criteria of an effective multimodal generative model when applied on more complex datasets than those used in previous benchmarks. In summary, we identify, formalize, and validate fundamental limitations of VAE-based approaches for modeling weakly-supervised data and discuss implications for real-world applications.
Deep Conditional Gaussian Mixture Model for Constrained Clustering
Jun 22, 2021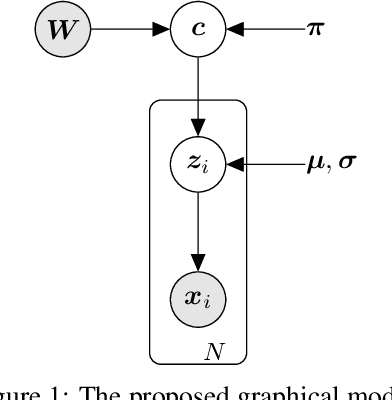
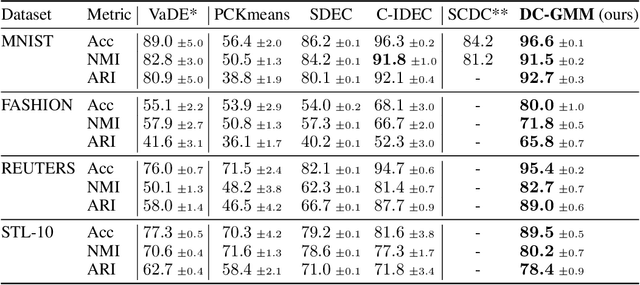
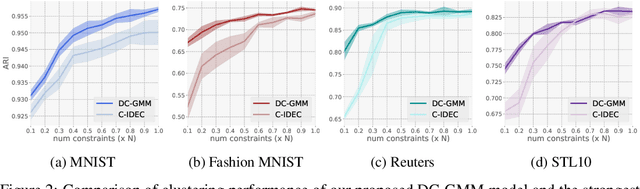
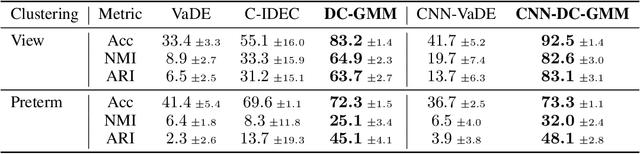
Abstract:Constrained clustering has gained significant attention in the field of machine learning as it can leverage prior information on a growing amount of only partially labeled data. Following recent advances in deep generative models, we propose a novel framework for constrained clustering that is intuitive, interpretable, and can be trained efficiently in the framework of stochastic gradient variational inference. By explicitly integrating domain knowledge in the form of probabilistic relations, our proposed model (DC-GMM) uncovers the underlying distribution of data conditioned on prior clustering preferences, expressed as pairwise constraints. These constraints guide the clustering process towards a desirable partition of the data by indicating which samples should or should not belong to the same cluster. We provide extensive experiments to demonstrate that DC-GMM shows superior clustering performances and robustness compared to state-of-the-art deep constrained clustering methods on a wide range of data sets. We further demonstrate the usefulness of our approach on two challenging real-world applications.
Generation of Differentially Private Heterogeneous Electronic Health Records
Jun 05, 2020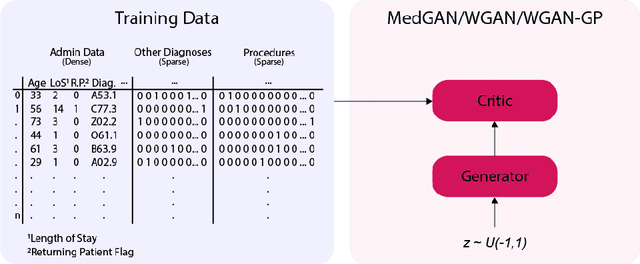
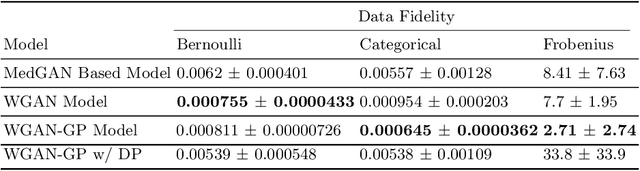
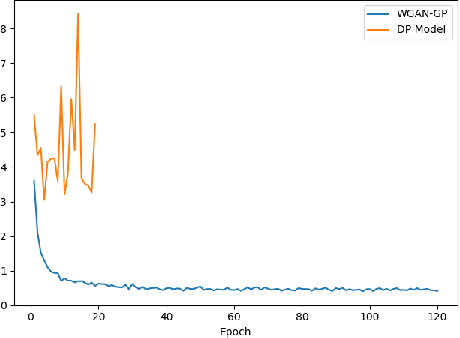
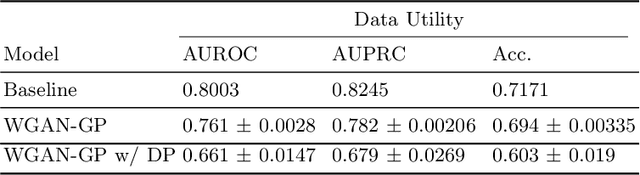
Abstract:Electronic Health Records (EHRs) are commonly used by the machine learning community for research on problems specifically related to health care and medicine. EHRs have the advantages that they can be easily distributed and contain many features useful for e.g. classification problems. What makes EHR data sets different from typical machine learning data sets is that they are often very sparse, due to their high dimensionality, and often contain heterogeneous (mixed) data types. Furthermore, the data sets deal with sensitive information, which limits the distribution of any models learned using them, due to privacy concerns. For these reasons, using EHR data in practice presents a real challenge. In this work, we explore using Generative Adversarial Networks to generate synthetic, heterogeneous EHRs with the goal of using these synthetic records in place of existing data sets for downstream classification tasks. We will further explore applying differential privacy (DP) preserving optimization in order to produce DP synthetic EHR data sets, which provide rigorous privacy guarantees, and are therefore shareable and usable in the real world. The performance (measured by AUROC, AUPRC and accuracy) of our model's synthetic, heterogeneous data is very close to the original data set (within 3 - 5% of the baseline) for the non-DP model when tested in a binary classification task. Using strong $(1, 10^{-5})$ DP, our model still produces data useful for machine learning tasks, albeit incurring a roughly 17% performance penalty in our tested classification task. We additionally perform a sub-population analysis and find that our model does not introduce any bias into the synthetic EHR data compared to the baseline in either male/female populations, or the 0-18, 19-50 and 51+ age groups in terms of classification performance for either the non-DP or DP variant.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge