Keyoumars Ashkan
From Pre- to Intra-operative MRI: Predicting Brain Shift in Temporal Lobe Resection for Epilepsy Surgery
Feb 03, 2026Abstract:Introduction: In neurosurgery, image-guided Neurosurgery Systems (IGNS) highly rely on preoperative brain magnetic resonance images (MRI) to assist surgeons in locating surgical targets and determining surgical paths. However, brain shift invalidates the preoperative MRI after dural opening. Updated intraoperative brain MRI with brain shift compensation is crucial for enhancing the precision of neuronavigation systems and ensuring the optimal outcome of surgical interventions. Methodology: We propose NeuralShift, a U-Net-based model that predicts brain shift entirely from pre-operative MRI for patients undergoing temporal lobe resection. We evaluated our results using Target Registration Errors (TREs) computed on anatomical landmarks located on the resection side and along the midline, and DICE scores comparing predicted intraoperative masks with masks derived from intraoperative MRI. Results: Our experimental results show that our model can predict the global deformation of the brain (DICE of 0.97) with accurate local displacements (achieve landmark TRE as low as 1.12 mm), compensating for large brain shifts during temporal lobe removal neurosurgery. Conclusion: Our proposed model is capable of predicting the global deformation of the brain during temporal lobe resection using only preoperative images, providing potential opportunities to the surgical team to increase safety and efficiency of neurosurgery and better outcomes to patients. Our contributions will be publicly available after acceptance in https://github.com/SurgicalDataScienceKCL/NeuralShift.
From artificial to organic: Rethinking the roots of intelligence for digital health
Dec 23, 2025Abstract:The term artificial implies an inherent dichotomy from the natural or organic. However, AI, as we know it, is a product of organic ingenuity: designed, implemented, and iteratively improved by human cognition. The very principles that underpin AI systems, from neural networks to decision-making algorithms, are inspired by the organic intelligence embedded in human neurobiology and evolutionary processes. The path from organic to artificial intelligence in digital health is neither mystical nor merely a matter of parameter count, it is fundamentally about organization and adaption. Thus, the boundaries between artificial and organic are far less distinct than the nomenclature suggests.
Overcoming challenges of translating deep-learning models for glioblastoma: the ZGBM consortium
May 07, 2024

Abstract:Objective: To report imaging protocol and scheduling variance in routine care of glioblastoma patients in order to demonstrate challenges of integrating deep-learning models in glioblastoma care pathways. Additionally, to understand the most common imaging studies and image contrasts to inform the development of potentially robust deep-learning models. Methods: MR imaging data were analysed from a random sample of five patients from the prospective cohort across five participating sites of the ZGBM consortium. Reported clinical and treatment data alongside DICOM header information were analysed to understand treatment pathway imaging schedules. Results: All sites perform all structural imaging at every stage in the pathway except for the presurgical study, where in some sites only contrast-enhanced T1-weighted imaging is performed. Diffusion MRI is the most common non-structural imaging type, performed at every site. Conclusion: The imaging protocol and scheduling varies across the UK, making it challenging to develop machine-learning models that could perform robustly at other centres. Structural imaging is performed most consistently across all centres. Advances in knowledge: Successful translation of deep-learning models will likely be based on structural post-treatment imaging unless there is significant effort made to standardise non-structural or peri-operative imaging protocols and schedules.
Machine Learning and Glioblastoma: Treatment Response Monitoring Biomarkers in 2021
Apr 15, 2021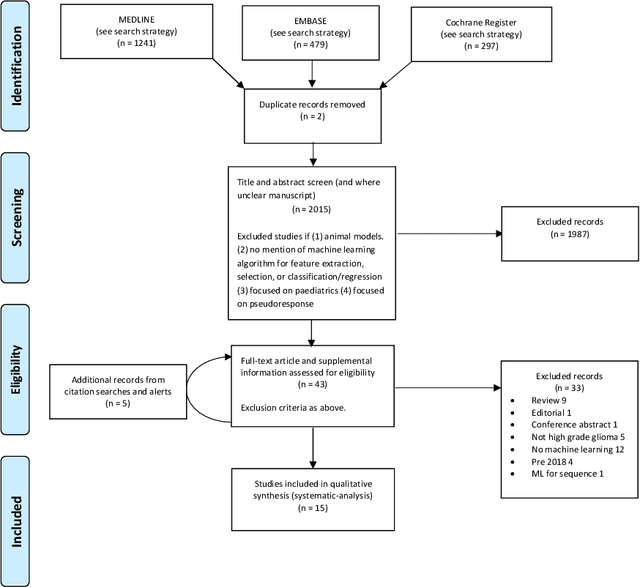
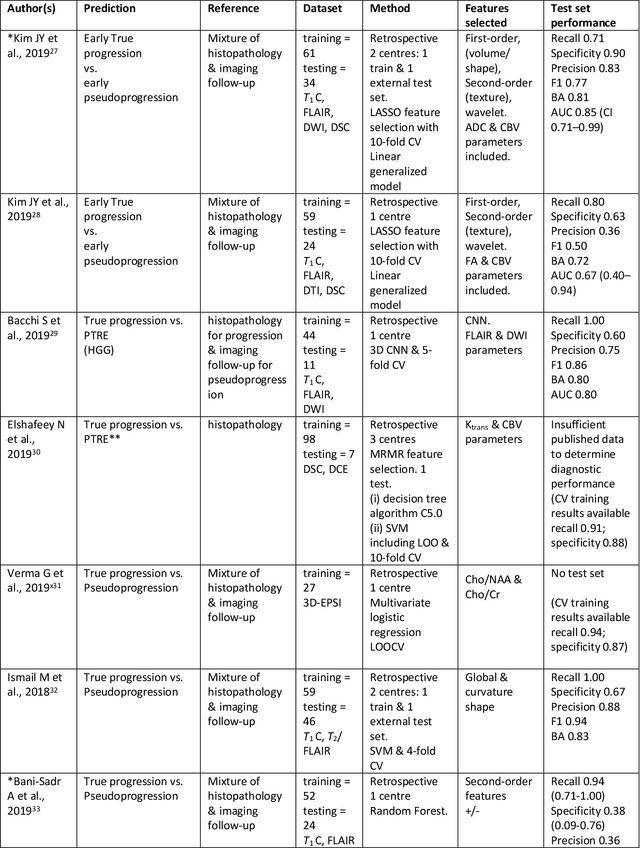
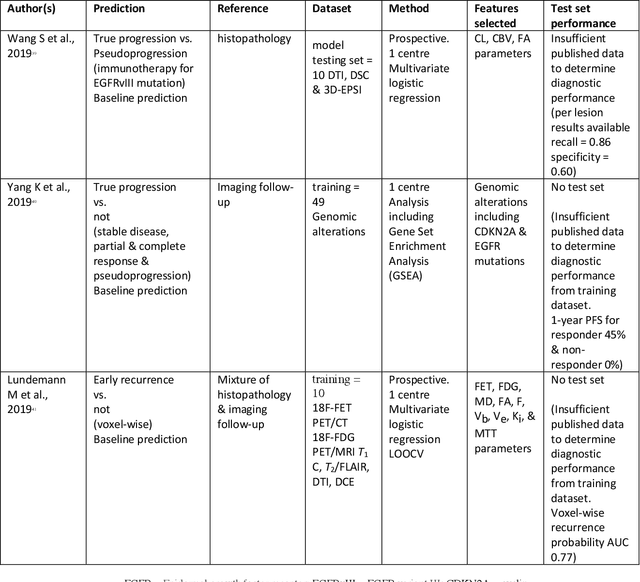
Abstract:The aim of the systematic review was to assess recently published studies on diagnostic test accuracy of glioblastoma treatment response monitoring biomarkers in adults, developed through machine learning (ML). Articles were searched for using MEDLINE, EMBASE, and the Cochrane Register. Included study participants were adult patients with high grade glioma who had undergone standard treatment (maximal resection, radiotherapy with concomitant and adjuvant temozolomide) and subsequently underwent follow-up imaging to determine treatment response status. Risk of bias and applicability was assessed with QUADAS 2 methodology. Contingency tables were created for hold-out test sets and recall, specificity, precision, F1-score, balanced accuracy calculated. Fifteen studies were included with 1038 patients in training sets and 233 in test sets. To determine whether there was progression or a mimic, the reference standard combination of follow-up imaging and histopathology at re-operation was applied in 67% of studies. The small numbers of patient included in studies, the high risk of bias and concerns of applicability in the study designs (particularly in relation to the reference standard and patient selection due to confounding), and the low level of evidence, suggest that limited conclusions can be drawn from the data. There is likely good diagnostic performance of machine learning models that use MRI features to distinguish between progression and mimics. The diagnostic performance of ML using implicit features did not appear to be superior to ML using explicit features. There are a range of ML-based solutions poised to become treatment response monitoring biomarkers for glioblastoma. To achieve this, the development and validation of ML models require large, well-annotated datasets where the potential for confounding in the study design has been carefully considered.
Using convolution neural networks to learn enhanced fiber orientation distribution models from commercially available diffusion magnetic resonance imaging
Aug 12, 2020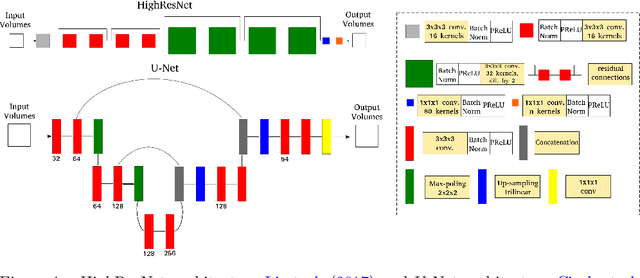
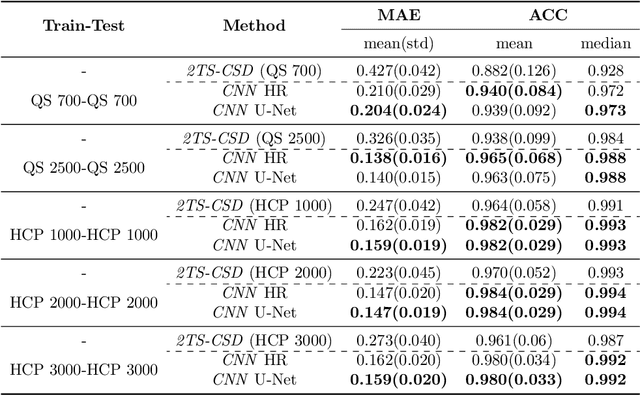
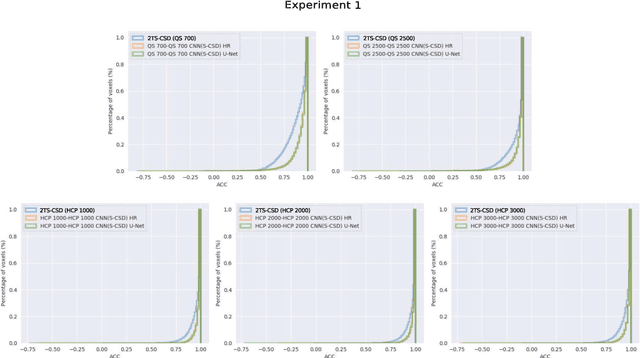
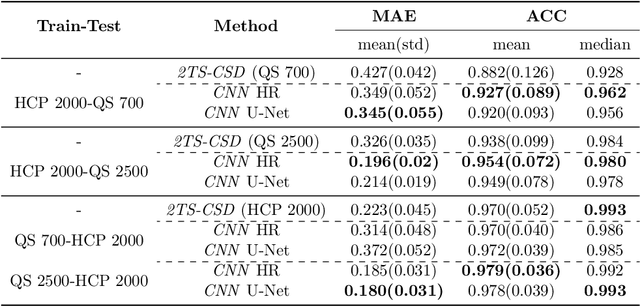
Abstract:Accurate local fiber orientation distribution (FOD) modeling based on diffusion magnetic resonance imaging (dMRI) capable of resolving complex fiber configurations benefit from specific acquisition protocols that impose a high number of gradient directions (b-vecs), a high maximum b-value (b-vals) and multiple b-values (multi-shell). However, acquisition time is limited in a clinical setting and commercial scanners may not provide robust state-of-the-art dMRI sequences. Therefore, dMRI is often acquired as single-shell (SS) (single b-value). Here, we learn improved FODs for commercially acquired dMRI. We evaluate the use of 3D convolutional neural networks (CNNs) to regress multi-shell FOS representations from single-shell representations, using the spherical harmonics basis obtained from constrained spherical deconvolution (CSD) to model FODs. We use U-Net and HighResNet 3D CNN architectures and data from the publicly available Human Connectome Dataset and a dataset acquired at National Hospital For Neurology and Neurosurgery Queen Square. We evaluate how well the CNN models can resolve local fiber orientation 1) when training and testing on datasets with same dMRI acquisition protocol; 2) when testing on dataset with a different dMRI acquisition protocol than used training the CNN models; and 3) when testing on datasets with a fewer number dMRI gradient directions than used training the CNN models. Our approach may enable robust CSD model estimation on dMRI acquisition protocols which are single shell and with a few gradient directions, reducing acquisition times, and thus, facilitating translation to time-limited clinical environments.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge