Kevin Wells
Fatigue-Aware Learning to Defer via Constrained Optimisation
Apr 01, 2026Abstract:Learning to defer (L2D) enables human-AI cooperation by deciding when an AI system should act autonomously or defer to a human expert. Existing L2D methods, however, assume static human performance, contradicting well-established findings on fatigue-induced degradation. We propose Fatigue-Aware Learning to Defer via Constrained Optimisation (FALCON), which explicitly models workload-varying human performance using psychologically grounded fatigue curves. FALCON formulates L2D as a Constrained Markov Decision Process (CMDP) whose state includes both task features and cumulative human workload, and optimises accuracy under human-AI cooperation budgets via PPO-Lagrangian training. We further introduce FA-L2D, a benchmark that systematically varies fatigue dynamics from near-static to rapidly degrading regimes. Experiments across multiple datasets show that FALCON consistently outperforms state-of-the-art L2D methods across coverage levels, generalises zero-shot to unseen experts with different fatigue patterns, and demonstrates the advantage of adaptive human-AI collaboration over AI-only or human-only decision-making when coverage lies strictly between 0 and 1.
Fair Distillation: Teaching Fairness from Biased Teachers in Medical Imaging
Nov 18, 2024



Abstract:Deep learning has achieved remarkable success in image classification and segmentation tasks. However, fairness concerns persist, as models often exhibit biases that disproportionately affect demographic groups defined by sensitive attributes such as race, gender, or age. Existing bias-mitigation techniques, including Subgroup Re-balancing, Adversarial Training, and Domain Generalization, aim to balance accuracy across demographic groups, but often fail to simultaneously improve overall accuracy, group-specific accuracy, and fairness due to conflicts among these interdependent objectives. We propose the Fair Distillation (FairDi) method, a novel fairness approach that decomposes these objectives by leveraging biased ``teacher'' models, each optimized for a specific demographic group. These teacher models then guide the training of a unified ``student'' model, which distills their knowledge to maximize overall and group-specific accuracies, while minimizing inter-group disparities. Experiments on medical imaging datasets show that FairDi achieves significant gains in both overall and group-specific accuracy, along with improved fairness, compared to existing methods. FairDi is adaptable to various medical tasks, such as classification and segmentation, and provides an effective solution for equitable model performance.
Coverage-Constrained Human-AI Cooperation with Multiple Experts
Nov 18, 2024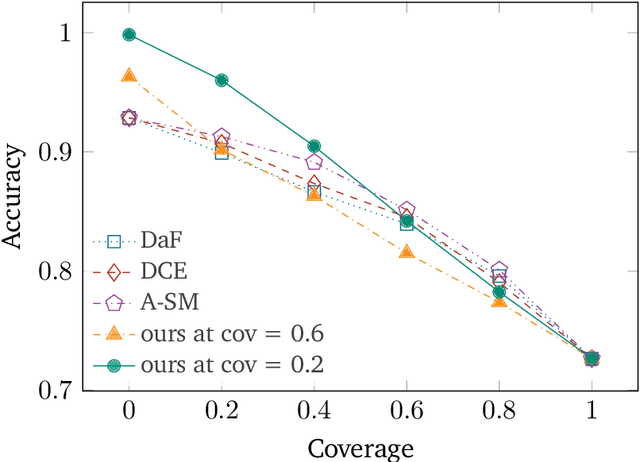
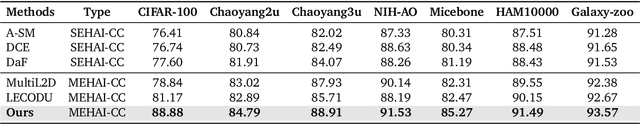
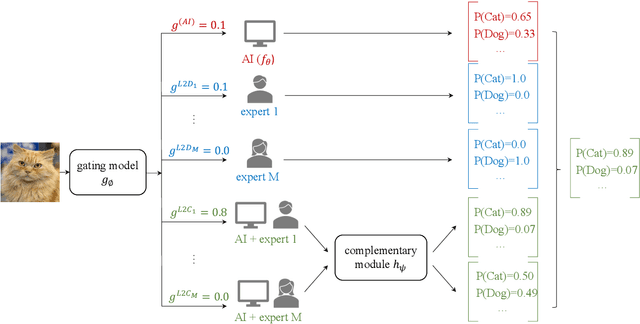
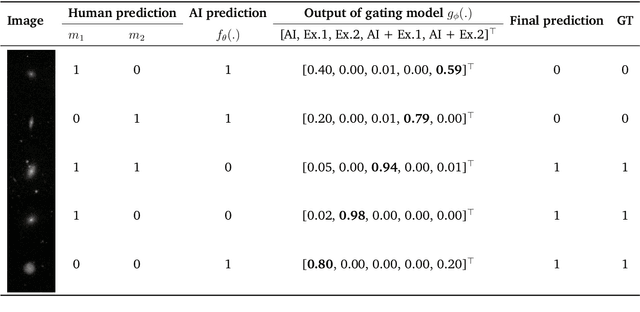
Abstract:Human-AI cooperative classification (HAI-CC) approaches aim to develop hybrid intelligent systems that enhance decision-making in various high-stakes real-world scenarios by leveraging both human expertise and AI capabilities. Current HAI-CC methods primarily focus on learning-to-defer (L2D), where decisions are deferred to human experts, and learning-to-complement (L2C), where AI and human experts make predictions cooperatively. However, a notable research gap remains in effectively exploring both L2D and L2C under diverse expert knowledge to improve decision-making, particularly when constrained by the cooperation cost required to achieve a target probability for AI-only selection (i.e., coverage). In this paper, we address this research gap by proposing the Coverage-constrained Learning to Defer and Complement with Specific Experts (CL2DC) method. CL2DC makes final decisions through either AI prediction alone or by deferring to or complementing a specific expert, depending on the input data. Furthermore, we propose a coverage-constrained optimisation to control the cooperation cost, ensuring it approximates a target probability for AI-only selection. This approach enables an effective assessment of system performance within a specified budget. Also, CL2DC is designed to address scenarios where training sets contain multiple noisy-label annotations without any clean-label references. Comprehensive evaluations on both synthetic and real-world datasets demonstrate that CL2DC achieves superior performance compared to state-of-the-art HAI-CC methods.
Learning to Complement and to Defer to Multiple Users
Jul 09, 2024



Abstract:With the development of Human-AI Collaboration in Classification (HAI-CC), integrating users and AI predictions becomes challenging due to the complex decision-making process. This process has three options: 1) AI autonomously classifies, 2) learning to complement, where AI collaborates with users, and 3) learning to defer, where AI defers to users. Despite their interconnected nature, these options have been studied in isolation rather than as components of a unified system. In this paper, we address this weakness with the novel HAI-CC methodology, called Learning to Complement and to Defer to Multiple Users (LECODU). LECODU not only combines learning to complement and learning to defer strategies, but it also incorporates an estimation of the optimal number of users to engage in the decision process. The training of LECODU maximises classification accuracy and minimises collaboration costs associated with user involvement. Comprehensive evaluations across real-world and synthesized datasets demonstrate LECODU's superior performance compared to state-of-the-art HAI-CC methods. Remarkably, even when relying on unreliable users with high rates of label noise, LECODU exhibits significant improvement over both human decision-makers alone and AI alone.
Learning to Complement with Multiple Humans (LECOMH): Integrating Multi-rater and Noisy-Label Learning into Human-AI Collaboration
Nov 22, 2023



Abstract:The advent of learning with noisy labels (LNL), multi-rater learning, and human-AI collaboration has revolutionised the development of robust classifiers, enabling them to address the challenges posed by different types of data imperfections and complex decision processes commonly encountered in real-world applications. While each of these methodologies has individually made significant strides in addressing their unique challenges, the development of techniques that can simultaneously tackle these three problems remains underexplored. This paper addresses this research gap by integrating noisy-label learning, multi-rater learning, and human-AI collaboration with new benchmarks and the innovative Learning to Complement with Multiple Humans (LECOMH) approach. LECOMH optimises the level of human collaboration during testing, aiming to optimise classification accuracy while minimising collaboration costs that vary from 0 to M, where M is the maximum number of human collaborators. We quantitatively compare LECOMH with leading human-AI collaboration methods using our proposed benchmarks. LECOMH consistently outperforms the competition, with accuracy improving as collaboration costs increase. Notably, LECOMH is the only method enhancing human labeller performance across all benchmarks.
Automatic Delineation of Kidney Region in DCE-MRI
May 26, 2019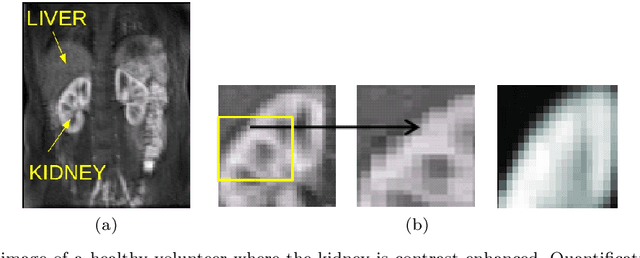
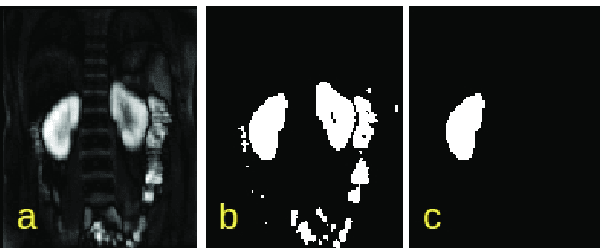
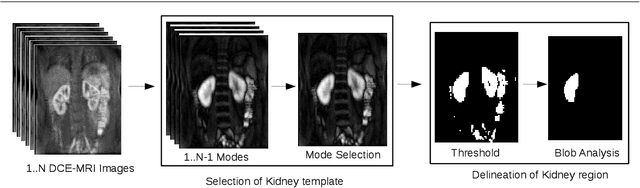
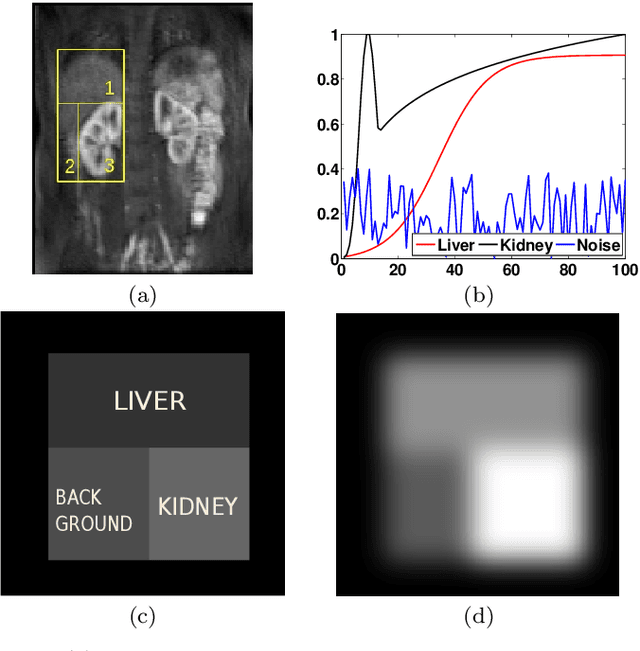
Abstract:Delineation of the kidney region in dynamic contrast-enhanced magnetic resonance Imaging (DCE-MRI) is required during post-acquisition analysis in order to quantify various aspects of renal function, such as filtration and perfusion or blood flow. However, this can be obfuscated by the Partial Volume Effect (PVE), caused due to the mixing of any single voxel with two or more signal intensities from adjacent regions such as liver region and other tissues. To avoid this problem, firstly, a kidney region of interest (ROI) needs to be defined for the analysis. A clinician may choose to select a region avoiding edges where PV mixing is likely to be significant. However, this approach is time-consuming and labour intensive. To address this issue, we present Dynamic Mode Decomposition (DMD) coupled with thresholding and blob analysis as a framework for automatic delineation of the kidney region. This method is first validated on synthetically generated data with ground-truth available and then applied to ten healthy volunteers' kidney DCE-MRI datasets. We found that the result obtained from our proposed framework is comparable to that of a human expert. For example, while our result gives an average Root Mean Square Error (RMSE) of 0.0097, the baseline achieves an average RMSE of 0.1196 across the 10 datasets. As a result, we conclude automatic modelling via DMD framework is a promising approach.
Functional Segmentation through Dynamic Mode Decomposition: Automatic Quantification of Kidney Function in DCE-MRI Images
May 24, 2019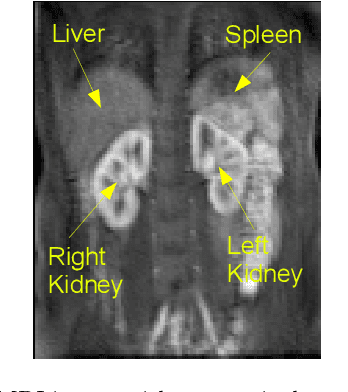
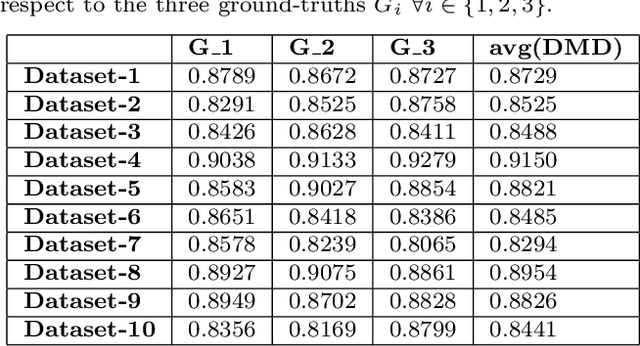
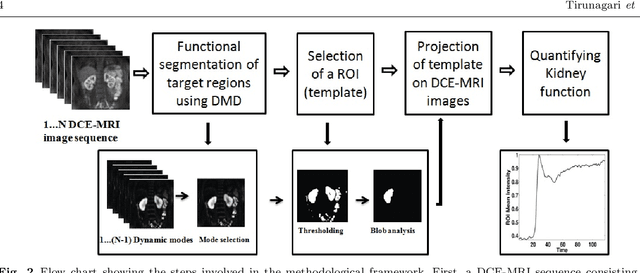
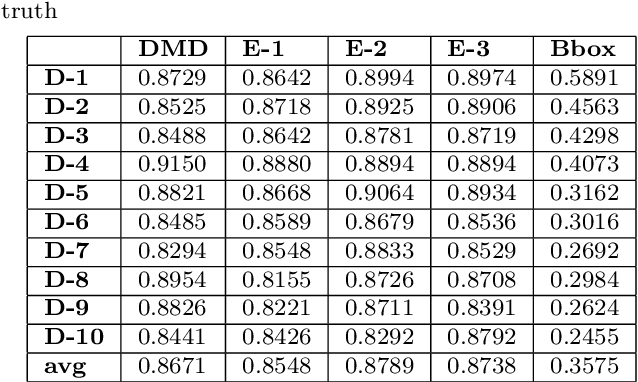
Abstract:Quantification of kidney function in Dynamic Contrast-Enhanced Magnetic Resonance Imaging (DCE-MRI) requires careful segmentation of the renal region of interest (ROI). Traditionally, human experts are required to manually delineate the kidney ROI across multiple images in the dynamic sequence. This approach is costly, time-consuming and labour intensive, and therefore acts to limit patient throughout and acts as one of the factors limiting the wider adoption of DCR-MRI in clinical practice. Therefore, to address this issue, we present the first use of Dynamic Mode Decomposition (DMD) as a basis for automatic segmentation of a dynamic sequence, in this case, kidney ROIs in DCE-MRI. Using DMD coupled combined with thresholding and connected component analysis is first validated on synthetically generated data with known ground-truth, and then applied to ten healthy volunteers' DCE-MRI datasets. We find that the segmentation result obtained from our proposed DMD framework is comparable to that of expert observers and very significantly better than that of an a-priori bounding box segmentation. Our result gives a mean Jaccard coefficient of 0.87, compared to mean scores of 0.85, 0.88 and 0.87 produced from three independent manual annotations. This represents the first use of DMD as a robust automatic data-driven segmentation approach without requiring any human intervention. This is a viable, efficient alternative approach to current manual methods of isolation of kidney function in DCE-MRI.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge