Kaidi Fu
FoldMark: Protecting Protein Generative Models with Watermarking
Oct 27, 2024Abstract:Protein structure is key to understanding protein function and is essential for progress in bioengineering, drug discovery, and molecular biology. Recently, with the incorporation of generative AI, the power and accuracy of computational protein structure prediction/design have been improved significantly. However, ethical concerns such as copyright protection and harmful content generation (biosecurity) pose challenges to the wide implementation of protein generative models. Here, we investigate whether it is possible to embed watermarks into protein generative models and their outputs for copyright authentication and the tracking of generated structures. As a proof of concept, we propose a two-stage method FoldMark as a generalized watermarking strategy for protein generative models. FoldMark first pretrain watermark encoder and decoder, which can minorly adjust protein structures to embed user-specific information and faithfully recover the information from the encoded structure. In the second step, protein generative models are fine-tuned with watermark Low-Rank Adaptation (LoRA) modules to preserve generation quality while learning to generate watermarked structures with high recovery rates. Extensive experiments are conducted on open-source protein structure prediction models (e.g., ESMFold and MultiFlow) and de novo structure design models (e.g., FrameDiff and FoldFlow) and we demonstrate that our method is effective across all these generative models. Meanwhile, our watermarking framework only exerts a negligible impact on the original protein structure quality and is robust under potential post-processing and adaptive attacks.
Latent Diffusion Models for Controllable RNA Sequence Generation
Sep 15, 2024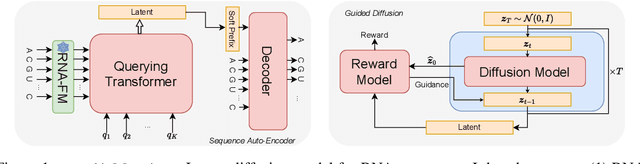
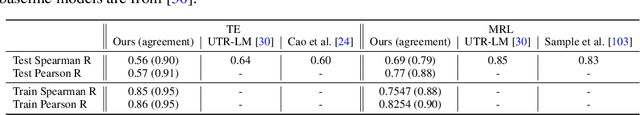
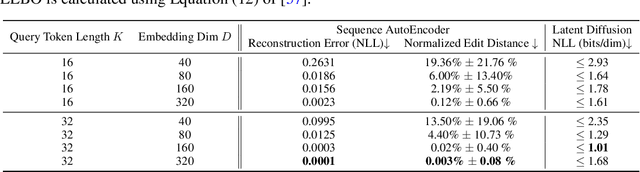

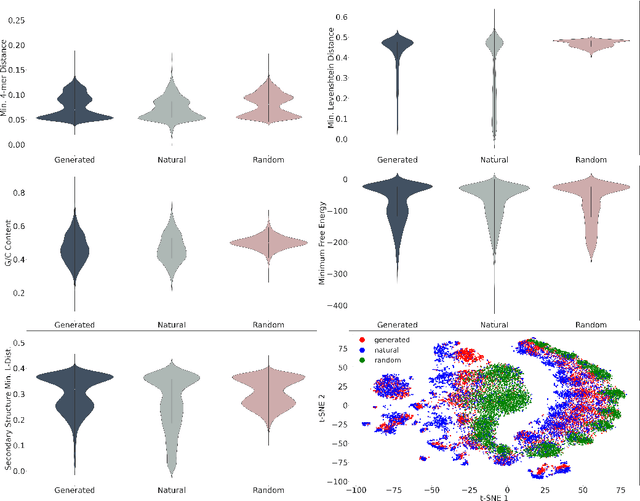
Abstract:This paper presents RNAdiffusion, a latent diffusion model for generating and optimizing discrete RNA sequences. RNA is a particularly dynamic and versatile molecule in biological processes. RNA sequences exhibit high variability and diversity, characterized by their variable lengths, flexible three-dimensional structures, and diverse functions. We utilize pretrained BERT-type models to encode raw RNAs into token-level biologically meaningful representations. A Q-Former is employed to compress these representations into a fixed-length set of latent vectors, with an autoregressive decoder trained to reconstruct RNA sequences from these latent variables. We then develop a continuous diffusion model within this latent space. To enable optimization, we train reward networks to estimate functional properties of RNA from the latent variables. We employ gradient-based guidance during the backward diffusion process, aiming to generate RNA sequences that are optimized for higher rewards. Empirical experiments confirm that RNAdiffusion generates non-coding RNAs that align with natural distributions across various biological indicators. We fine-tuned the diffusion model on untranslated regions (UTRs) of mRNA and optimize sample sequences for protein translation efficiencies. Our guided diffusion model effectively generates diverse UTR sequences with high Mean Ribosome Loading (MRL) and Translation Efficiency (TE), surpassing baselines. These results hold promise for studies on RNA sequence-function relationships, protein synthesis, and enhancing therapeutic RNA design.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge