Jochen Kruecker
Ultrasound Image Synthesis Using Generative AI for Lung Ultrasound Detection
Jan 10, 2025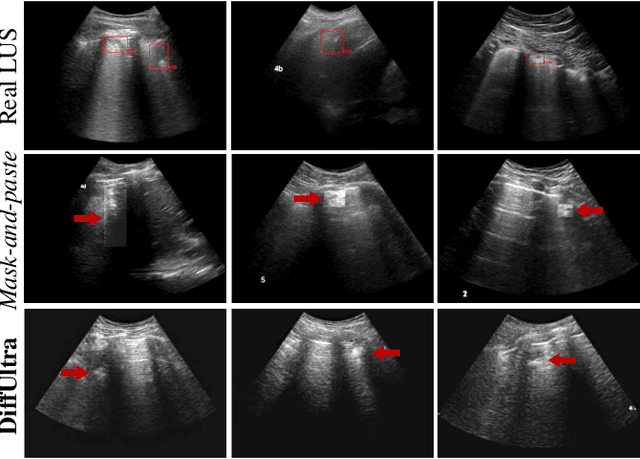
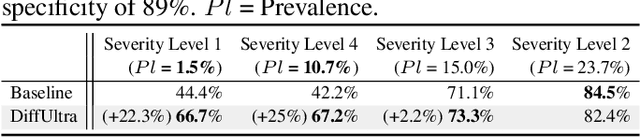

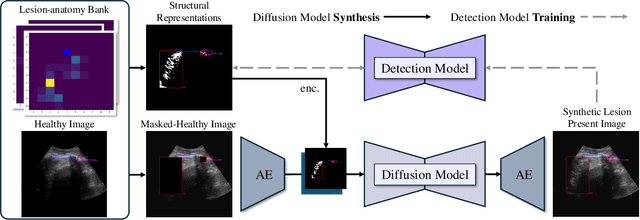

Abstract:Developing reliable healthcare AI models requires training with representative and diverse data. In imbalanced datasets, model performance tends to plateau on the more prevalent classes while remaining low on less common cases. To overcome this limitation, we propose DiffUltra, the first generative AI technique capable of synthesizing realistic Lung Ultrasound (LUS) images with extensive lesion variability. Specifically, we condition the generative AI by the introduced Lesion-anatomy Bank, which captures the lesion's structural and positional properties from real patient data to guide the image synthesis.We demonstrate that DiffUltra improves consolidation detection by 5.6% in AP compared to the models trained solely on real patient data. More importantly, DiffUltra increases data diversity and prevalence of rare cases, leading to a 25% AP improvement in detecting rare instances such as large lung consolidations, which make up only 10% of the dataset.
Contrastive Self-Supervised Learning for Spatio-Temporal Analysis of Lung Ultrasound Videos
Oct 14, 2023Abstract:Self-supervised learning (SSL) methods have shown promise for medical imaging applications by learning meaningful visual representations, even when the amount of labeled data is limited. Here, we extend state-of-the-art contrastive learning SSL methods to 2D+time medical ultrasound video data by introducing a modified encoder and augmentation method capable of learning meaningful spatio-temporal representations, without requiring constraints on the input data. We evaluate our method on the challenging clinical task of identifying lung consolidations (an important pathological feature) in ultrasound videos. Using a multi-center dataset of over 27k lung ultrasound videos acquired from over 500 patients, we show that our method can significantly improve performance on downstream localization and classification of lung consolidation. Comparisons against baseline models trained without SSL show that the proposed methods are particularly advantageous when the size of labeled training data is limited (e.g., as little as 5% of the training set).
Weakly Semi-Supervised Detection in Lung Ultrasound Videos
Aug 08, 2023Abstract:Frame-by-frame annotation of bounding boxes by clinical experts is often required to train fully supervised object detection models on medical video data. We propose a method for improving object detection in medical videos through weak supervision from video-level labels. More concretely, we aggregate individual detection predictions into video-level predictions and extend a teacher-student training strategy to provide additional supervision via a video-level loss. We also introduce improvements to the underlying teacher-student framework, including methods to improve the quality of pseudo-labels based on weak supervision and adaptive schemes to optimize knowledge transfer between the student and teacher networks. We apply this approach to the clinically important task of detecting lung consolidations (seen in respiratory infections such as COVID-19 pneumonia) in medical ultrasound videos. Experiments reveal that our framework improves detection accuracy and robustness compared to baseline semi-supervised models, and improves efficiency in data and annotation usage.
Learning Deep Similarity Metric for 3D MR-TRUS Registration
Oct 15, 2018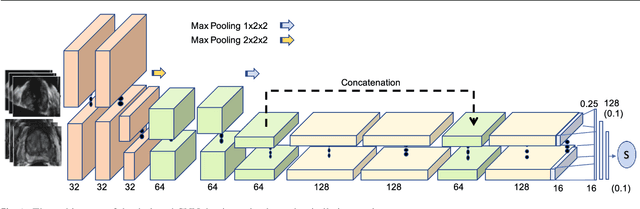
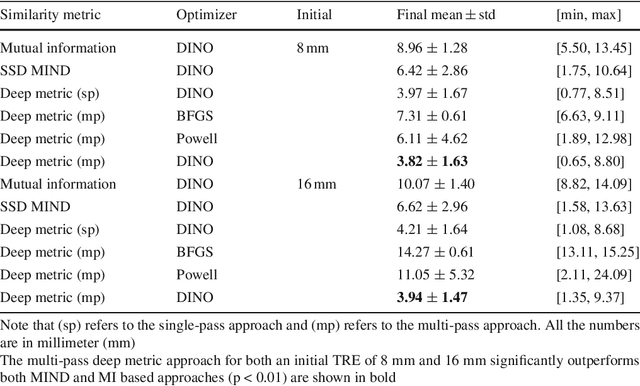
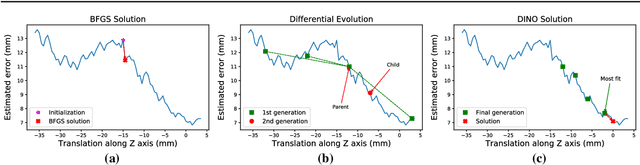
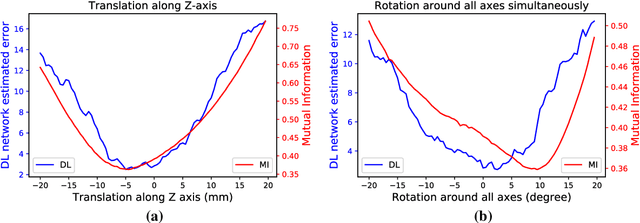
Abstract:Purpose: The fusion of transrectal ultrasound (TRUS) and magnetic resonance (MR) images for guiding targeted prostate biopsy has significantly improved the biopsy yield of aggressive cancers. A key component of MR-TRUS fusion is image registration. However, it is very challenging to obtain a robust automatic MR-TRUS registration due to the large appearance difference between the two imaging modalities. The work presented in this paper aims to tackle this problem by addressing two challenges: (i) the definition of a suitable similarity metric and (ii) the determination of a suitable optimization strategy. Methods: This work proposes the use of a deep convolutional neural network to learn a similarity metric for MR-TRUS registration. We also use a composite optimization strategy that explores the solution space in order to search for a suitable initialization for the second-order optimization of the learned metric. Further, a multi-pass approach is used in order to smooth the metric for optimization. Results: The learned similarity metric outperforms the classical mutual information and also the state-of-the-art MIND feature based methods. The results indicate that the overall registration framework has a large capture range. The proposed deep similarity metric based approach obtained a mean TRE of 3.86mm (with an initial TRE of 16mm) for this challenging problem. Conclusion: A similarity metric that is learned using a deep neural network can be used to assess the quality of any given image registration and can be used in conjunction with the aforementioned optimization framework to perform automatic registration that is robust to poor initialization.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge