Jochen Guck
AI-driven projection tomography with multicore fibre-optic cell rotation
Dec 12, 2023Abstract:Optical tomography has emerged as a non-invasive imaging method, providing three-dimensional insights into subcellular structures and thereby enabling a deeper understanding of cellular functions, interactions, and processes. Conventional optical tomography methods are constrained by a limited illumination scanning range, leading to anisotropic resolution and incomplete imaging of cellular structures. To overcome this problem, we employ a compact multi-core fibre-optic cell rotator system that facilitates precise optical manipulation of cells within a microfluidic chip, achieving full-angle projection tomography with isotropic resolution. Moreover, we demonstrate an AI-driven tomographic reconstruction workflow, which can be a paradigm shift from conventional computational methods, often demanding manual processing, to a fully autonomous process. The performance of the proposed cell rotation tomography approach is validated through the three-dimensional reconstruction of cell phantoms and HL60 human cancer cells. The versatility of this learning-based tomographic reconstruction workflow paves the way for its broad application across diverse tomographic imaging modalities, including but not limited to flow cytometry tomography and acoustic rotation tomography. Therefore, this AI-driven approach can propel advancements in cell biology, aiding in the inception of pioneering therapeutics, and augmenting early-stage cancer diagnostics.
Quantitative phase imaging through an ultra-thin lensless fiber endoscope
Dec 25, 2021
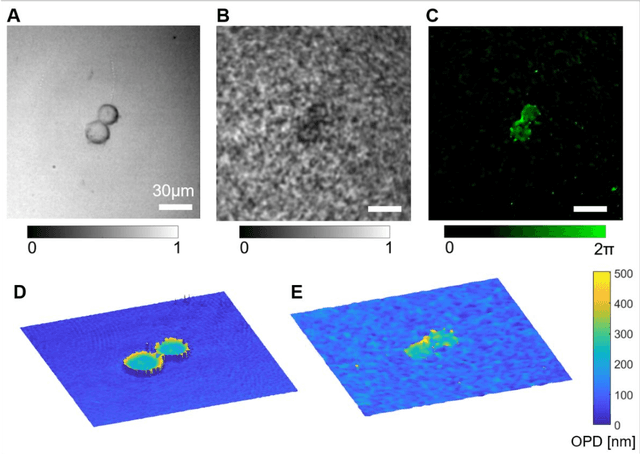
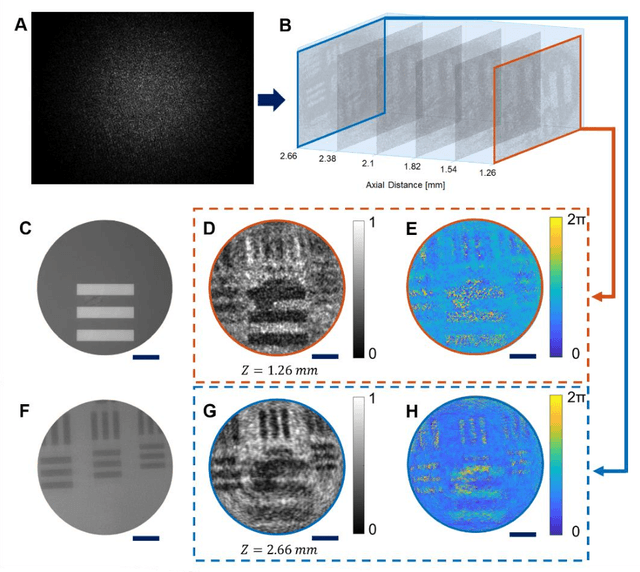
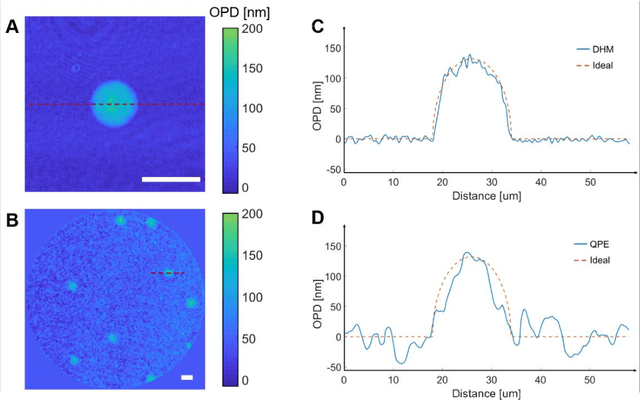

Abstract:Quantitative phase imaging (QPI) is a label-free technique providing both morphology and quantitative biophysical information in biomedicine. However, applying such a powerful technique to in vivo pathological diagnosis remains challenging. Multi-core fiber bundles (MCFs) enable ultra-thin probes for in vivo imaging, but current MCF imaging techniques are limited to amplitude imaging modalities. We demonstrate a computational lensless microendoscope that uses an ultra-thin bare MCF to perform quantitative phase imaging of biomedical samples with up to 1 {\mu}m lateral resolution and nanoscale axial resolution. The incident complex light field at the measurement side is precisely reconstructed from a single-shot far-field speckle pattern at the detection side, enabling digital focusing and 3D volumetric reconstruction without any mechanical movement. The accuracy of the quantitative phase reconstruction is validated by imaging the phase target and hydrogel beads through the MCF. With the proposed imaging modality, 3D imaging of human cancer cells is achieved through the ultra-thin fiber endoscope, promising widespread clinical applications.
Cell Mechanics Based Computational Classification of Red Blood Cells Via Machine Intelligence Applied to Morpho-Rheological Markers
Mar 02, 2020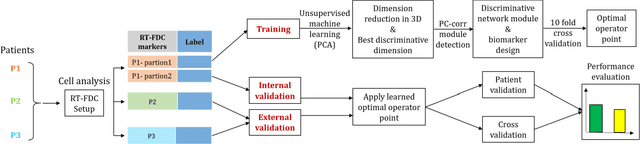
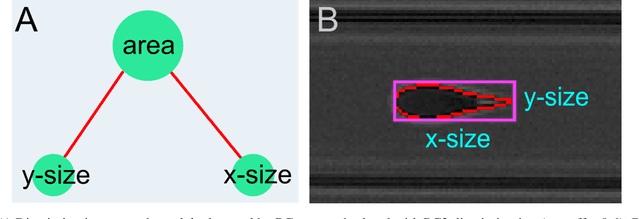
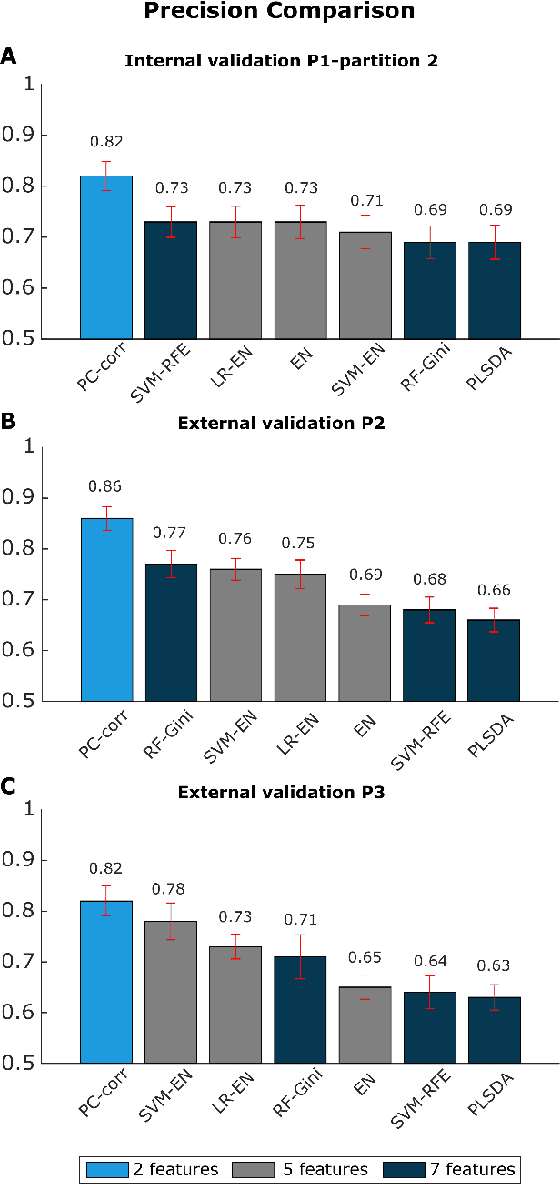
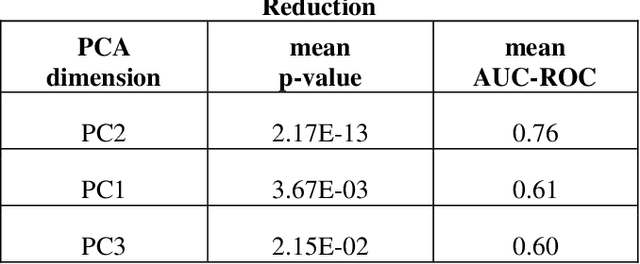
Abstract:Despite fluorescent cell-labelling being widely employed in biomedical studies, some of its drawbacks are inevitable, with unsuitable fluorescent probes or probes inducing a functional change being the main limitations. Consequently, the demand for and development of label-free methodologies to classify cells is strong and its impact on precision medicine is relevant. Towards this end, high-throughput techniques for cell mechanical phenotyping have been proposed to get a multidimensional biophysical characterization of single cells. With this motivation, our goal here is to investigate the extent to which an unsupervised machine learning methodology, which is applied exclusively on morpho-rheological markers obtained by real-time deformability and fluorescence cytometry (RT-FDC), can address the difficult task of providing label-free discrimination of reticulocytes from mature red blood cells. We focused on this problem, since the characterization of reticulocytes (their percentage and cellular features) in the blood is vital in multiple human disease conditions, especially bone-marrow disorders such as anemia and leukemia. Our approach reports promising label-free results in the classification of reticulocytes from mature red blood cells, and it represents a step forward in the development of high-throughput morpho-rheological-based methodologies for the computational categorization of single cells. Besides, our methodology can be an alternative but also a complementary method to integrate with existing cell-labelling techniques.
* 13 pages, 3 figures, 4 tables
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge