Jianming Liang
Project Imaging-X: A Survey of 1000+ Open-Access Medical Imaging Datasets for Foundation Model Development
Mar 29, 2026Abstract:Foundation models have demonstrated remarkable success across diverse domains and tasks, primarily due to the thrive of large-scale, diverse, and high-quality datasets. However, in the field of medical imaging, the curation and assembling of such medical datasets are highly challenging due to the reliance on clinical expertise and strict ethical and privacy constraints, resulting in a scarcity of large-scale unified medical datasets and hindering the development of powerful medical foundation models. In this work, we present the largest survey to date of medical image datasets, covering over 1,000 open-access datasets with a systematic catalog of their modalities, tasks, anatomies, annotations, limitations, and potential for integration. Our analysis exposes a landscape that is modest in scale, fragmented across narrowly scoped tasks, and unevenly distributed across organs and modalities, which in turn limits the utility of existing medical image datasets for developing versatile and robust medical foundation models. To turn fragmentation into scale, we propose a metadata-driven fusion paradigm (MDFP) that integrates public datasets with shared modalities or tasks, thereby transforming multiple small data silos into larger, more coherent resources. Building on MDFP, we release an interactive discovery portal that enables end-to-end, automated medical image dataset integration, and compile all surveyed datasets into a unified, structured table that clearly summarizes their key characteristics and provides reference links, offering the community an accessible and comprehensive repository. By charting the current terrain and offering a principled path to dataset consolidation, our survey provides a practical roadmap for scaling medical imaging corpora, supporting faster data discovery, more principled dataset creation, and more capable medical foundation models.
Lamps: Learning Anatomy from Multiple Perspectives via Self-supervision in Chest Radiographs
Jan 02, 2026Abstract:Foundation models have been successful in natural language processing and computer vision because they are capable of capturing the underlying structures (foundation) of natural languages. However, in medical imaging, the key foundation lies in human anatomy, as these images directly represent the internal structures of the body, reflecting the consistency, coherence, and hierarchy of human anatomy. Yet, existing self-supervised learning (SSL) methods often overlook these perspectives, limiting their ability to effectively learn anatomical features. To overcome the limitation, we built Lamps (learning anatomy from multiple perspectives via self-supervision) pre-trained on large-scale chest radiographs by harmoniously utilizing the consistency, coherence, and hierarchy of human anatomy as the supervision signal. Extensive experiments across 10 datasets evaluated through fine-tuning and emergent property analysis demonstrate Lamps' superior robustness, transferability, and clinical potential when compared to 10 baseline models. By learning from multiple perspectives, Lamps presents a unique opportunity for foundation models to develop meaningful, robust representations that are aligned with the structure of human anatomy.
Learning Anatomy from Multiple Perspectives via Self-supervision in Chest Radiographs
Dec 28, 2025Abstract:Foundation models have been successful in natural language processing and computer vision because they are capable of capturing the underlying structures (foundation) of natural languages. However, in medical imaging, the key foundation lies in human anatomy, as these images directly represent the internal structures of the body, reflecting the consistency, coherence, and hierarchy of human anatomy. Yet, existing self-supervised learning (SSL) methods often overlook these perspectives, limiting their ability to effectively learn anatomical features. To overcome the limitation, we built Lamps (learning anatomy from multiple perspectives via self-supervision) pre-trained on large-scale chest radiographs by harmoniously utilizing the consistency, coherence, and hierarchy of human anatomy as the supervision signal. Extensive experiments across 10 datasets evaluated through fine-tuning and emergent property analysis demonstrate Lamps' superior robustness, transferability, and clinical potential when compared to 10 baseline models. By learning from multiple perspectives, Lamps presents a unique opportunity for foundation models to develop meaningful, robust representations that are aligned with the structure of human anatomy.
Foundation X: Integrating Classification, Localization, and Segmentation through Lock-Release Pretraining Strategy for Chest X-ray Analysis
Mar 12, 2025Abstract:Developing robust and versatile deep-learning models is essential for enhancing diagnostic accuracy and guiding clinical interventions in medical imaging, but it requires a large amount of annotated data. The advancement of deep learning has facilitated the creation of numerous medical datasets with diverse expert-level annotations. Aggregating these datasets can maximize data utilization and address the inadequacy of labeled data. However, the heterogeneity of expert-level annotations across tasks such as classification, localization, and segmentation presents a significant challenge for learning from these datasets. To this end, we introduce nFoundation X, an end-to-end framework that utilizes diverse expert-level annotations from numerous public datasets to train a foundation model capable of multiple tasks including classification, localization, and segmentation. To address the challenges of annotation and task heterogeneity, we propose a Lock-Release pretraining strategy to enhance the cyclic learning from multiple datasets, combined with the student-teacher learning paradigm, ensuring the model retains general knowledge for all tasks while preventing overfitting to any single task. To demonstrate the effectiveness of Foundation X, we trained a model using 11 chest X-ray datasets, covering annotations for classification, localization, and segmentation tasks. Our experimental results show that Foundation X achieves notable performance gains through extensive annotation utilization, excels in cross-dataset and cross-task learning, and further enhances performance in organ localization and segmentation tasks. All code and pretrained models are publicly accessible at https://github.com/jlianglab/Foundation_X.
ACE: Anatomically Consistent Embeddings in Composition and Decomposition
Jan 17, 2025Abstract:Medical images acquired from standardized protocols show consistent macroscopic or microscopic anatomical structures, and these structures consist of composable/decomposable organs and tissues, but existing self-supervised learning (SSL) methods do not appreciate such composable/decomposable structure attributes inherent to medical images. To overcome this limitation, this paper introduces a novel SSL approach called ACE to learn anatomically consistent embedding via composition and decomposition with two key branches: (1) global consistency, capturing discriminative macro-structures via extracting global features; (2) local consistency, learning fine-grained anatomical details from composable/decomposable patch features via corresponding matrix matching. Experimental results across 6 datasets 2 backbones, evaluated in few-shot learning, fine-tuning, and property analysis, show ACE's superior robustness, transferability, and clinical potential. The innovations of our ACE lie in grid-wise image cropping, leveraging the intrinsic properties of compositionality and decompositionality of medical images, bridging the semantic gap from high-level pathologies to low-level tissue anomalies, and providing a new SSL method for medical imaging.
Representing Part-Whole Hierarchies in Foundation Models by Learning Localizability, Composability, and Decomposability from Anatomy via Self-Supervision
Apr 24, 2024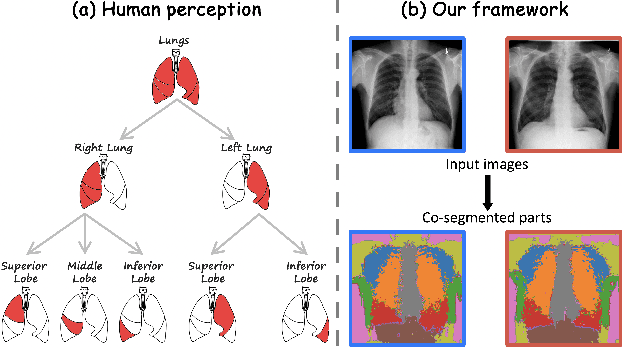
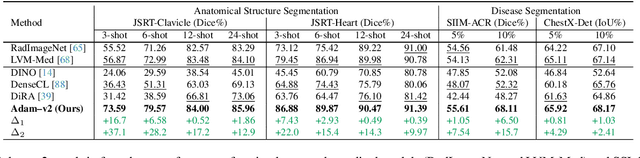
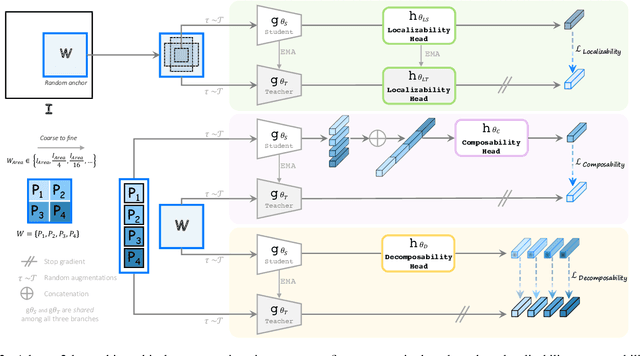
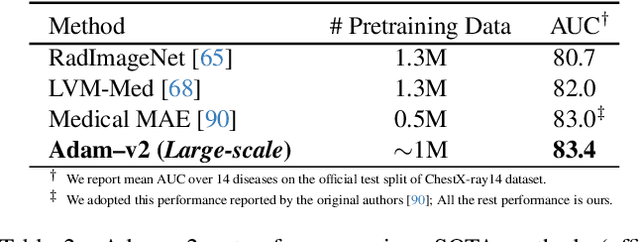
Abstract:Humans effortlessly interpret images by parsing them into part-whole hierarchies; deep learning excels in learning multi-level feature spaces, but they often lack explicit coding of part-whole relations, a prominent property of medical imaging. To overcome this limitation, we introduce Adam-v2, a new self-supervised learning framework extending Adam [79] by explicitly incorporating part-whole hierarchies into its learning objectives through three key branches: (1) Localizability, acquiring discriminative representations to distinguish different anatomical patterns; (2) Composability, learning each anatomical structure in a parts-to-whole manner; and (3) Decomposability, comprehending each anatomical structure in a whole-to-parts manner. Experimental results across 10 tasks, compared to 11 baselines in zero-shot, few-shot transfer, and full fine-tuning settings, showcase Adam-v2's superior performance over large-scale medical models and existing SSL methods across diverse downstream tasks. The higher generality and robustness of Adam-v2's representations originate from its explicit construction of hierarchies for distinct anatomical structures from unlabeled medical images. Adam-v2 preserves a semantic balance of anatomical diversity and harmony in its embedding, yielding representations that are both generic and semantically meaningful, yet overlooked in existing SSL methods. All code and pretrained models are available at https://github.com/JLiangLab/Eden.
Learning Anatomically Consistent Embedding for Chest Radiography
Dec 01, 2023Abstract:Self-supervised learning (SSL) approaches have recently shown substantial success in learning visual representations from unannotated images. Compared with photographic images, medical images acquired with the same imaging protocol exhibit high consistency in anatomy. To exploit this anatomical consistency, this paper introduces a novel SSL approach, called PEAC (patch embedding of anatomical consistency), for medical image analysis. Specifically, in this paper, we propose to learn global and local consistencies via stable grid-based matching, transfer pre-trained PEAC models to diverse downstream tasks, and extensively demonstrate that (1) PEAC achieves significantly better performance than the existing state-of-the-art fully/self-supervised methods, and (2) PEAC captures the anatomical structure consistency across views of the same patient and across patients of different genders, weights, and healthy statuses, which enhances the interpretability of our method for medical image analysis.
Foundation Ark: Accruing and Reusing Knowledge for Superior and Robust Performance
Oct 14, 2023Abstract:Deep learning nowadays offers expert-level and sometimes even super-expert-level performance, but achieving such performance demands massive annotated data for training (e.g., Google's proprietary CXR Foundation Model (CXR-FM) was trained on 821,544 labeled and mostly private chest X-rays (CXRs)). Numerous datasets are publicly available in medical imaging but individually small and heterogeneous in expert labels. We envision a powerful and robust foundation model that can be trained by aggregating numerous small public datasets. To realize this vision, we have developed Ark, a framework that accrues and reuses knowledge from heterogeneous expert annotations in various datasets. As a proof of concept, we have trained two Ark models on 335,484 and 704,363 CXRs, respectively, by merging several datasets including ChestX-ray14, CheXpert, MIMIC-II, and VinDr-CXR, evaluated them on a wide range of imaging tasks covering both classification and segmentation via fine-tuning, linear-probing, and gender-bias analysis, and demonstrated our Ark's superior and robust performance over the SOTA fully/self-supervised baselines and Google's proprietary CXR-FM. This enhanced performance is attributed to our simple yet powerful observation that aggregating numerous public datasets diversifies patient populations and accrues knowledge from diverse experts, yielding unprecedented performance yet saving annotation cost. With all codes and pretrained models released at GitHub.com/JLiangLab/Ark, we hope that Ark exerts an important impact on open science, as accruing and reusing knowledge from expert annotations in public datasets can potentially surpass the performance of proprietary models trained on unusually large data, inspiring many more researchers worldwide to share codes and datasets to build open foundation models, accelerate open science, and democratize deep learning for medical imaging.
Towards Foundation Models Learned from Anatomy in Medical Imaging via Self-Supervision
Sep 27, 2023Abstract:Human anatomy is the foundation of medical imaging and boasts one striking characteristic: its hierarchy in nature, exhibiting two intrinsic properties: (1) locality: each anatomical structure is morphologically distinct from the others; and (2) compositionality: each anatomical structure is an integrated part of a larger whole. We envision a foundation model for medical imaging that is consciously and purposefully developed upon this foundation to gain the capability of "understanding" human anatomy and to possess the fundamental properties of medical imaging. As our first step in realizing this vision towards foundation models in medical imaging, we devise a novel self-supervised learning (SSL) strategy that exploits the hierarchical nature of human anatomy. Our extensive experiments demonstrate that the SSL pretrained model, derived from our training strategy, not only outperforms state-of-the-art (SOTA) fully/self-supervised baselines but also enhances annotation efficiency, offering potential few-shot segmentation capabilities with performance improvements ranging from 9% to 30% for segmentation tasks compared to SSL baselines. This performance is attributed to the significance of anatomy comprehension via our learning strategy, which encapsulates the intrinsic attributes of anatomical structures-locality and compositionality-within the embedding space, yet overlooked in existing SSL methods. All code and pretrained models are available at https://github.com/JLiangLab/Eden.
Unifying Visual Perception by Dispersible Points Learning
Aug 18, 2022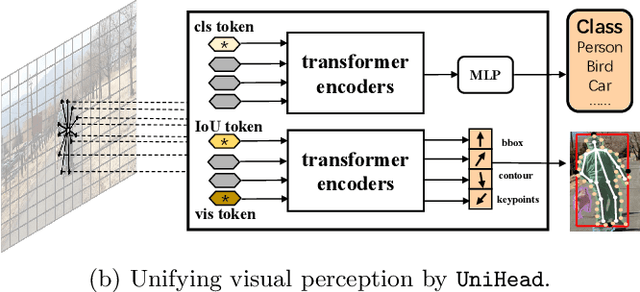
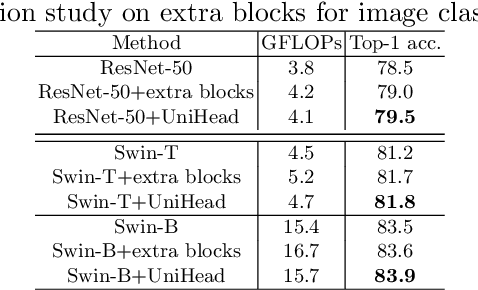
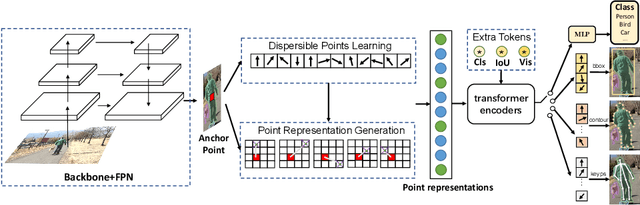
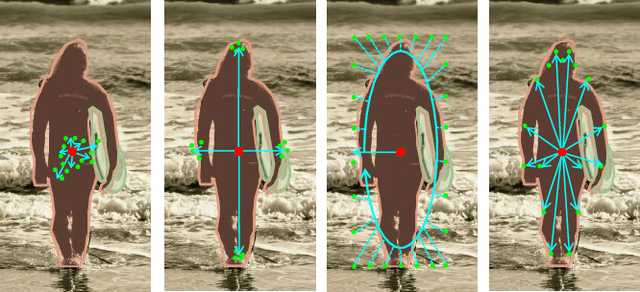
Abstract:We present a conceptually simple, flexible, and universal visual perception head for variant visual tasks, e.g., classification, object detection, instance segmentation and pose estimation, and different frameworks, such as one-stage or two-stage pipelines. Our approach effectively identifies an object in an image while simultaneously generating a high-quality bounding box or contour-based segmentation mask or set of keypoints. The method, called UniHead, views different visual perception tasks as the dispersible points learning via the transformer encoder architecture. Given a fixed spatial coordinate, UniHead adaptively scatters it to different spatial points and reasons about their relations by transformer encoder. It directly outputs the final set of predictions in the form of multiple points, allowing us to perform different visual tasks in different frameworks with the same head design. We show extensive evaluations on ImageNet classification and all three tracks of the COCO suite of challenges, including object detection, instance segmentation and pose estimation. Without bells and whistles, UniHead can unify these visual tasks via a single visual head design and achieve comparable performance compared to expert models developed for each task.We hope our simple and universal UniHead will serve as a solid baseline and help promote universal visual perception research. Code and models are available at https://github.com/Sense-X/UniHead.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge