Jiadi Luo
SMILE: a Scale-aware Multiple Instance Learning Method for Multicenter STAS Lung Cancer Histopathology Diagnosis
Mar 18, 2025Abstract:Spread through air spaces (STAS) represents a newly identified aggressive pattern in lung cancer, which is known to be associated with adverse prognostic factors and complex pathological features. Pathologists currently rely on time consuming manual assessments, which are highly subjective and prone to variation. This highlights the urgent need for automated and precise diag nostic solutions. 2,970 lung cancer tissue slides are comprised from multiple centers, re-diagnosed them, and constructed and publicly released three lung cancer STAS datasets: STAS CSU (hospital), STAS TCGA, and STAS CPTAC. All STAS datasets provide corresponding pathological feature diagnoses and related clinical data. To address the bias, sparse and heterogeneous nature of STAS, we propose an scale-aware multiple instance learning(SMILE) method for STAS diagnosis of lung cancer. By introducing a scale-adaptive attention mechanism, the SMILE can adaptively adjust high attention instances, reducing over-reliance on local regions and promoting consistent detection of STAS lesions. Extensive experiments show that SMILE achieved competitive diagnostic results on STAS CSU, diagnosing 251 and 319 STAS samples in CPTAC andTCGA,respectively, surpassing clinical average AUC. The 11 open baseline results are the first to be established for STAS research, laying the foundation for the future expansion, interpretability, and clinical integration of computational pathology technologies. The datasets and code are available at https://anonymous.4open.science/r/IJCAI25-1DA1.
Feature-interactive Siamese graph encoder-based image analysis to predict STAS from histopathology images in lung cancer
Nov 22, 2024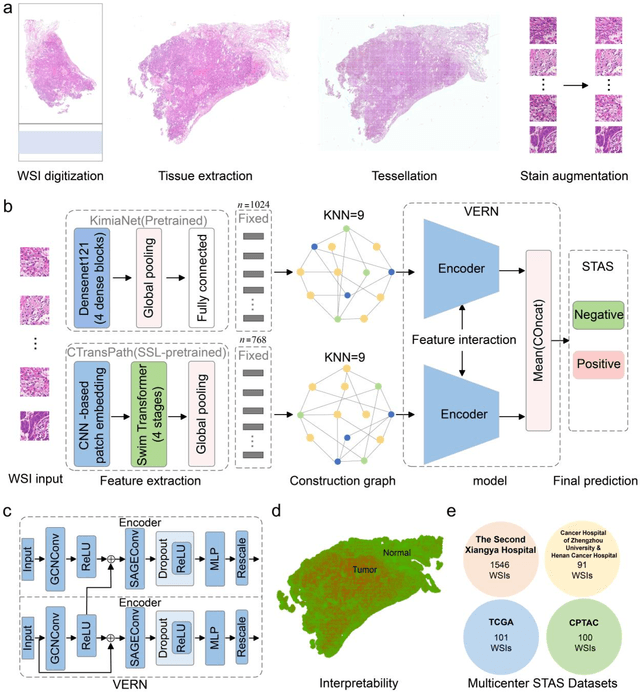
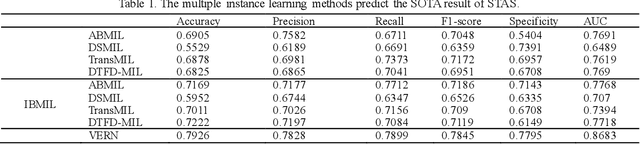
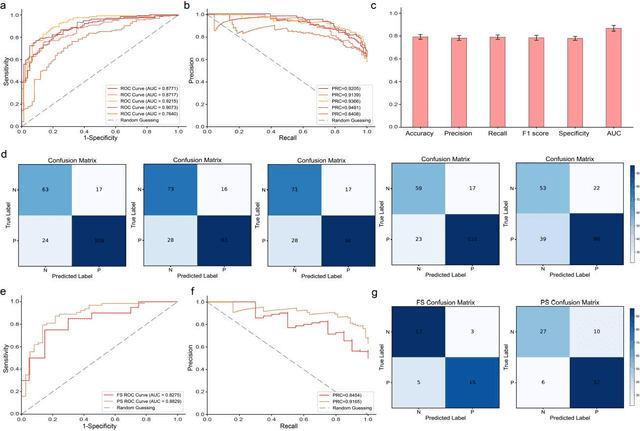
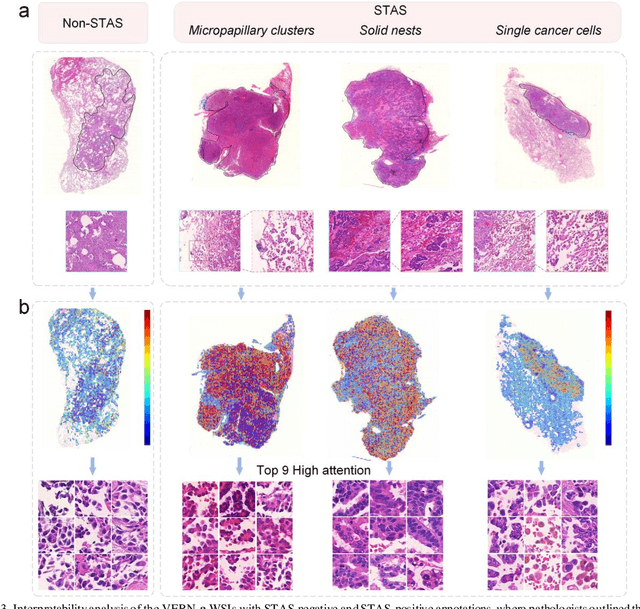
Abstract:Spread through air spaces (STAS) is a distinct invasion pattern in lung cancer, crucial for prognosis assessment and guiding surgical decisions. Histopathology is the gold standard for STAS detection, yet traditional methods are subjective, time-consuming, and prone to misdiagnosis, limiting large-scale applications. We present VERN, an image analysis model utilizing a feature-interactive Siamese graph encoder to predict STAS from lung cancer histopathological images. VERN captures spatial topological features with feature sharing and skip connections to enhance model training. Using 1,546 histopathology slides, we built a large single-cohort STAS lung cancer dataset. VERN achieved an AUC of 0.9215 in internal validation and AUCs of 0.8275 and 0.8829 in frozen and paraffin-embedded test sections, respectively, demonstrating clinical-grade performance. Validated on a single-cohort and three external datasets, VERN showed robust predictive performance and generalizability, providing an open platform (http://plr.20210706.xyz:5000/) to enhance STAS diagnosis efficiency and accuracy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge