Jeanne Shen
Pathology-CoT: Learning Visual Chain-of-Thought Agent from Expert Whole Slide Image Diagnosis Behavior
Oct 06, 2025Abstract:Diagnosing a whole-slide image is an interactive, multi-stage process involving changes in magnification and movement between fields. Although recent pathology foundation models are strong, practical agentic systems that decide what field to examine next, adjust magnification, and deliver explainable diagnoses are still lacking. The blocker is data: scalable, clinically aligned supervision of expert viewing behavior that is tacit and experience-based, not written in textbooks or online, and therefore absent from large language model training. We introduce the AI Session Recorder, which works with standard WSI viewers to unobtrusively record routine navigation and convert the viewer logs into standardized behavioral commands (inspect or peek at discrete magnifications) and bounding boxes. A lightweight human-in-the-loop review turns AI-drafted rationales into the Pathology-CoT dataset, a form of paired "where to look" and "why it matters" supervision produced at roughly six times lower labeling time. Using this behavioral data, we build Pathologist-o3, a two-stage agent that first proposes regions of interest and then performs behavior-guided reasoning. On gastrointestinal lymph-node metastasis detection, it achieved 84.5% precision, 100.0% recall, and 75.4% accuracy, exceeding the state-of-the-art OpenAI o3 model and generalizing across backbones. To our knowledge, this constitutes one of the first behavior-grounded agentic systems in pathology. Turning everyday viewer logs into scalable, expert-validated supervision, our framework makes agentic pathology practical and establishes a path to human-aligned, upgradeable clinical AI.
Deep Learning-Based Sparse Whole-Slide Image Analysis for the Diagnosis of Gastric Intestinal Metaplasia
Jan 05, 2022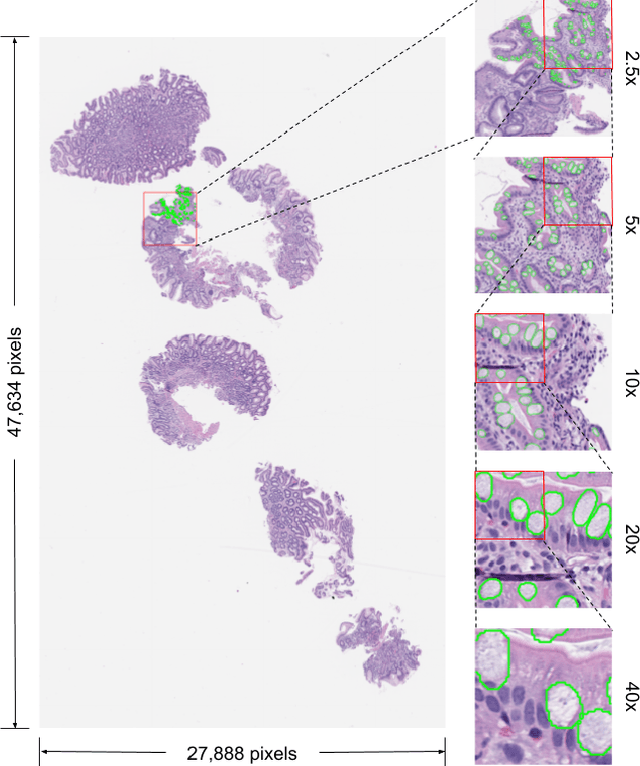
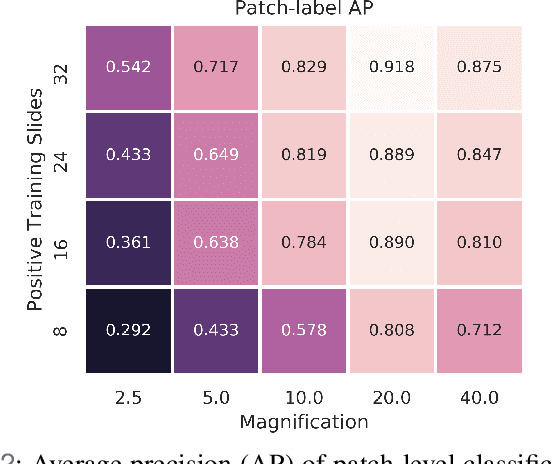

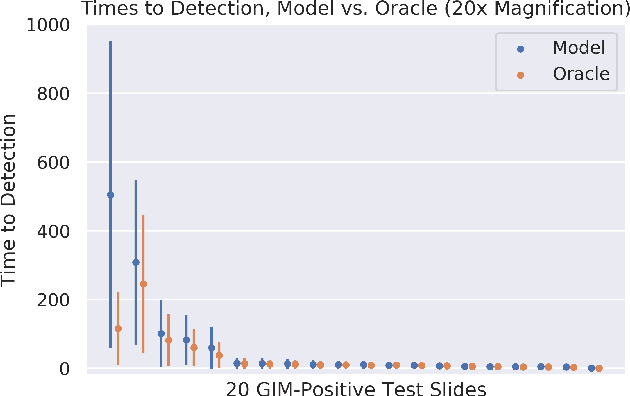
Abstract:In recent years, deep learning has successfully been applied to automate a wide variety of tasks in diagnostic histopathology. However, fast and reliable localization of small-scale regions-of-interest (ROI) has remained a key challenge, as discriminative morphologic features often occupy only a small fraction of a gigapixel-scale whole-slide image (WSI). In this paper, we propose a sparse WSI analysis method for the rapid identification of high-power ROI for WSI-level classification. We develop an evaluation framework inspired by the early classification literature, in order to quantify the tradeoff between diagnostic performance and inference time for sparse analytic approaches. We test our method on a common but time-consuming task in pathology - that of diagnosing gastric intestinal metaplasia (GIM) on hematoxylin and eosin (H&E)-stained slides from endoscopic biopsy specimens. GIM is a well-known precursor lesion along the pathway to development of gastric cancer. We performed a thorough evaluation of the performance and inference time of our approach on a test set of GIM-positive and GIM-negative WSI, finding that our method successfully detects GIM in all positive WSI, with a WSI-level classification area under the receiver operating characteristic curve (AUC) of 0.98 and an average precision (AP) of 0.95. Furthermore, we show that our method can attain these metrics in under one minute on a standard CPU. Our results are applicable toward the goal of developing neural networks that can easily be deployed in clinical settings to support pathologists in quickly localizing and diagnosing small-scale morphologic features in WSI.
Learning domain-agnostic visual representation for computational pathology using medically-irrelevant style transfer augmentation
Feb 02, 2021
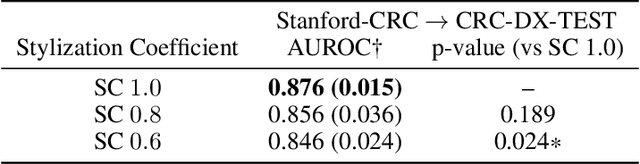
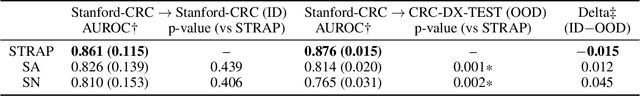


Abstract:Suboptimal generalization of machine learning models on unseen data is a key challenge which hampers the clinical applicability of such models to medical imaging. Although various methods such as domain adaptation and domain generalization have evolved to combat this challenge, learning robust and generalizable representations is core to medical image understanding, and continues to be a problem. Here, we propose STRAP (Style TRansfer Augmentation for histoPathology), a form of data augmentation based on random style transfer from artistic paintings, for learning domain-agnostic visual representations in computational pathology. Style transfer replaces the low-level texture content of images with the uninformative style of randomly selected artistic paintings, while preserving high-level semantic content. This improves robustness to domain shift and can be used as a simple yet powerful tool for learning domain-agnostic representations. We demonstrate that STRAP leads to state-of-the-art performance, particularly in the presence of domain shifts, on a particular classification task of predicting microsatellite status in colorectal cancer using digitized histopathology images.
Analysis Of Multi Field Of View Cnn And Attention Cnn On H&E Stained Whole-slide Images On Hepatocellular Carcinoma
Feb 18, 2020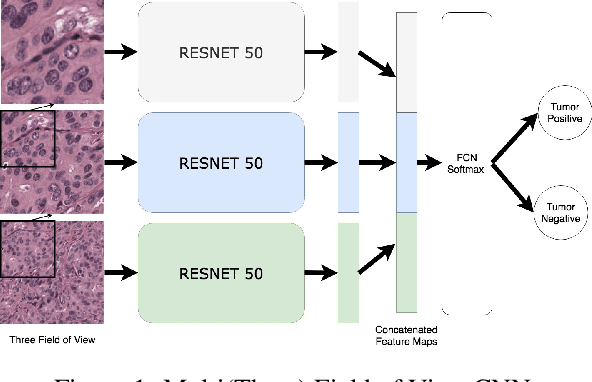
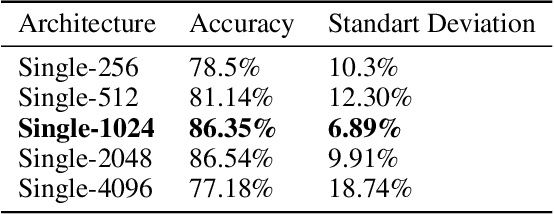

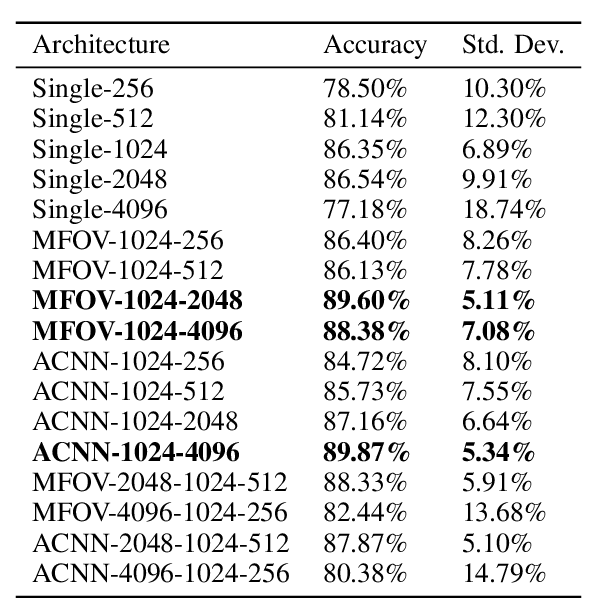
Abstract:Hepatocellular carcinoma (HCC) is a leading cause of cancer-related death worldwide. Whole-slide imaging which is a method of scanning glass slides have been employed for diagnosis of HCC. Using high resolution Whole-slide images is infeasible for Convolutional Neural Network applications. Hence tiling the Whole-slide images is a common methodology for assigning Convolutional Neural Networks for classification and segmentation. Determination of the tile size affects the performance of the algorithms since small field of view can not capture the information on a larger scale and large field of view can not capture the information on a cellular scale. In this work, the effect of tile size on performance for classification problem is analysed. In addition, Multi Field of View CNN is assigned for taking advantage of the information provided by different tile sizes and Attention CNN is assigned for giving the capability of voting most contributing tile size. It is found that employing more than one tile size significantly increases the performance of the classification by 3.97% and both algorithms are found successful over the algorithm which uses only one tile size.
Plexus Convolutional Neural Network (PlexusNet): A novel neural network architecture for histologic image analysis
Aug 24, 2019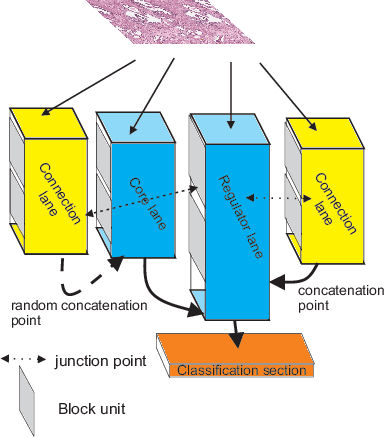
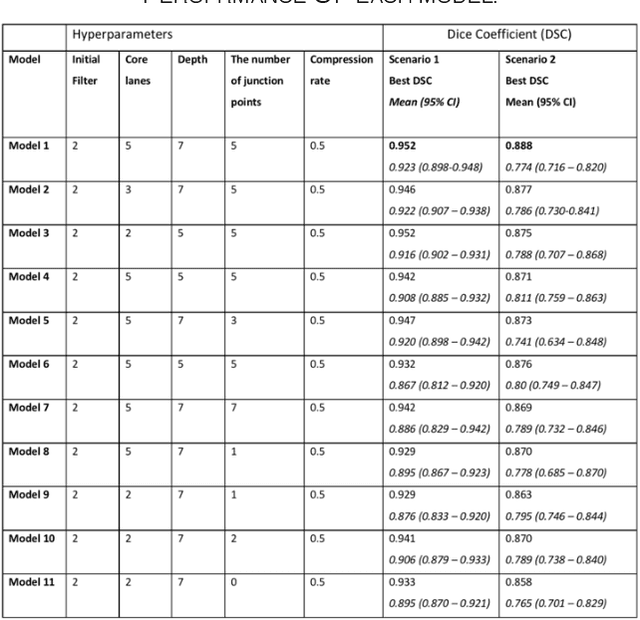
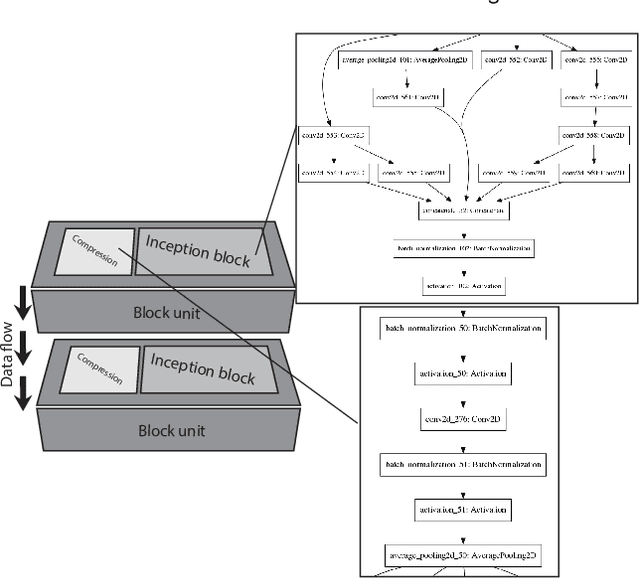
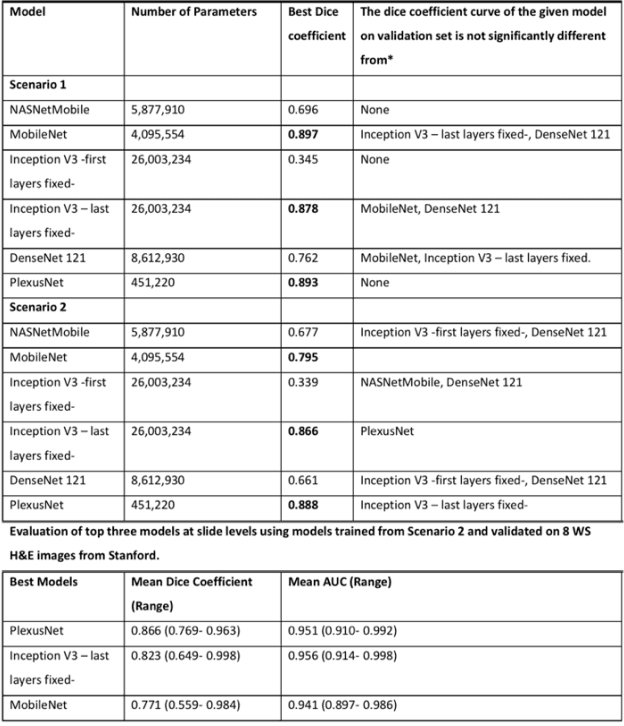
Abstract:Different convolutional neural network (CNN) models have been tested for their application in histologic imaging analyses. However, these models are prone to overfitting due to their large parameter capacity, requiring more data and expensive computational resources for model training. Given these limitations, we developed and tested PlexusNet for histologic evaluation using a single GPU by a batch dimension of 16x512x512x3. We utilized 62 Hematoxylin and eosin stain (H&E) annotated histological images of radical prostatectomy cases from TCGA-PRAD and Stanford University, and 24 H&E whole-slide images with hepatocellular carcinoma from TCGA-LIHC diagnostic histology images. Base models were DenseNet, Inception V3, and MobileNet and compared with PlexusNet. The dice coefficient (DSC) was evaluated for each model. PlexusNet delivered comparable classification performance (DSC at patch level: 0.89) for H&E whole-slice images in distinguishing prostate cancer from normal tissues. The parameter capacity of PlexusNet is 9 times smaller than MobileNet or 58 times smaller than Inception V3, respectively. Similar findings were observed in distinguishing hepatocellular carcinoma from non-cancerous liver histologies (DSC at patch level: 0.85). As conclusion, PlexusNet represents a novel model architecture for histological image analysis that achieves classification performance comparable to the base models while providing orders-of-magnitude memory savings.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge