Deep Learning-Based Sparse Whole-Slide Image Analysis for the Diagnosis of Gastric Intestinal Metaplasia
Paper and Code
Jan 05, 2022
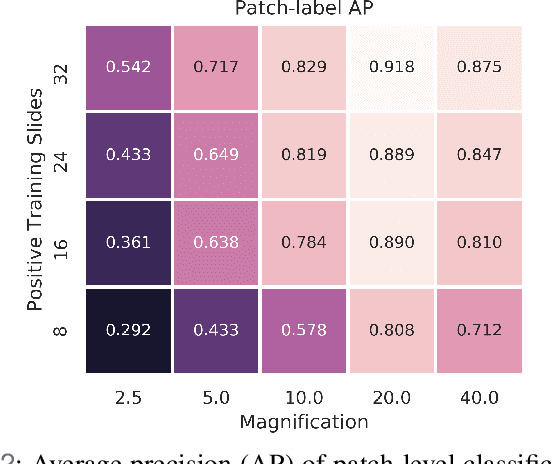

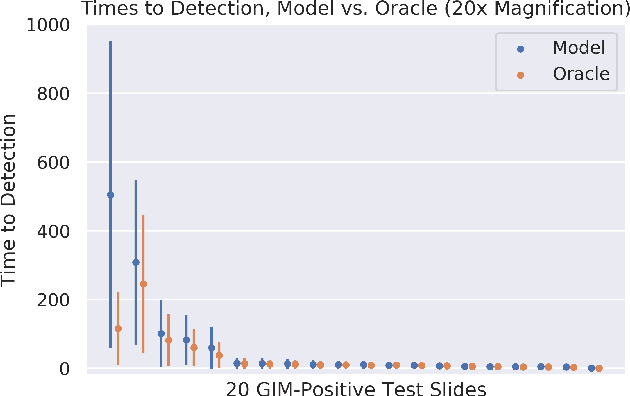
In recent years, deep learning has successfully been applied to automate a wide variety of tasks in diagnostic histopathology. However, fast and reliable localization of small-scale regions-of-interest (ROI) has remained a key challenge, as discriminative morphologic features often occupy only a small fraction of a gigapixel-scale whole-slide image (WSI). In this paper, we propose a sparse WSI analysis method for the rapid identification of high-power ROI for WSI-level classification. We develop an evaluation framework inspired by the early classification literature, in order to quantify the tradeoff between diagnostic performance and inference time for sparse analytic approaches. We test our method on a common but time-consuming task in pathology - that of diagnosing gastric intestinal metaplasia (GIM) on hematoxylin and eosin (H&E)-stained slides from endoscopic biopsy specimens. GIM is a well-known precursor lesion along the pathway to development of gastric cancer. We performed a thorough evaluation of the performance and inference time of our approach on a test set of GIM-positive and GIM-negative WSI, finding that our method successfully detects GIM in all positive WSI, with a WSI-level classification area under the receiver operating characteristic curve (AUC) of 0.98 and an average precision (AP) of 0.95. Furthermore, we show that our method can attain these metrics in under one minute on a standard CPU. Our results are applicable toward the goal of developing neural networks that can easily be deployed in clinical settings to support pathologists in quickly localizing and diagnosing small-scale morphologic features in WSI.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge