Jason Z. Kim
University of Pennsylvania
$Γ$-VAE: Curvature regularized variational autoencoders for uncovering emergent low dimensional geometric structure in high dimensional data
Mar 02, 2024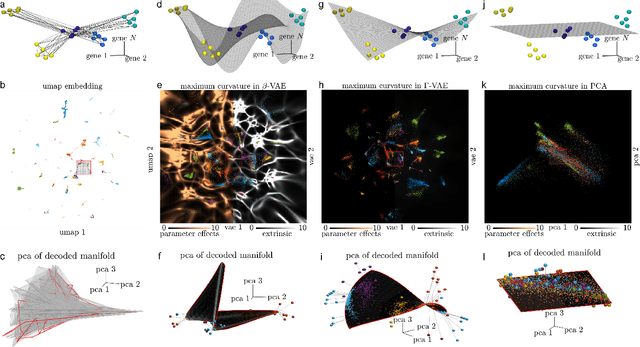
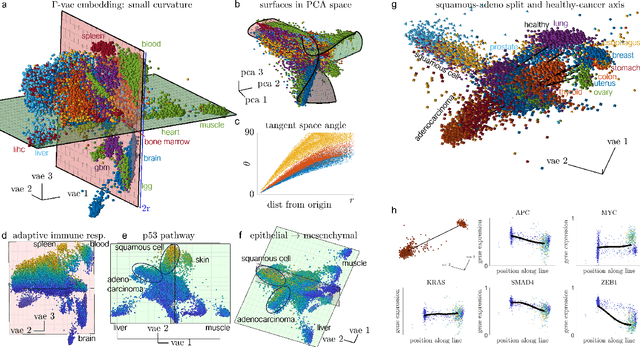
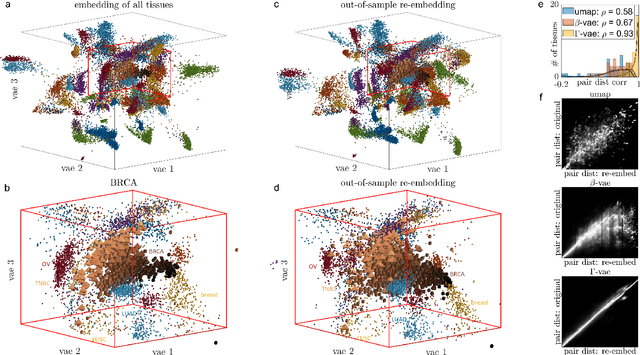

Abstract:Natural systems with emergent behaviors often organize along low-dimensional subsets of high-dimensional spaces. For example, despite the tens of thousands of genes in the human genome, the principled study of genomics is fruitful because biological processes rely on coordinated organization that results in lower dimensional phenotypes. To uncover this organization, many nonlinear dimensionality reduction techniques have successfully embedded high-dimensional data into low-dimensional spaces by preserving local similarities between data points. However, the nonlinearities in these methods allow for too much curvature to preserve general trends across multiple non-neighboring data clusters, thereby limiting their interpretability and generalizability to out-of-distribution data. Here, we address both of these limitations by regularizing the curvature of manifolds generated by variational autoencoders, a process we coin ``$\Gamma$-VAE''. We demonstrate its utility using two example data sets: bulk RNA-seq from the The Cancer Genome Atlas (TCGA) and the Genotype Tissue Expression (GTEx); and single cell RNA-seq from a lineage tracing experiment in hematopoietic stem cell differentiation. We find that the resulting regularized manifolds identify mesoscale structure associated with different cancer cell types, and accurately re-embed tissues from completely unseen, out-of distribution cancers as if they were originally trained on them. Finally, we show that preserving long-range relationships to differentiated cells separates undifferentiated cells -- which have not yet specialized -- according to their eventual fate. Broadly, we anticipate that regularizing the curvature of generative models will enable more consistent, predictive, and generalizable models in any high-dimensional system with emergent low-dimensional behavior.
Learning Continuous Chaotic Attractors with a Reservoir Computer
Oct 16, 2021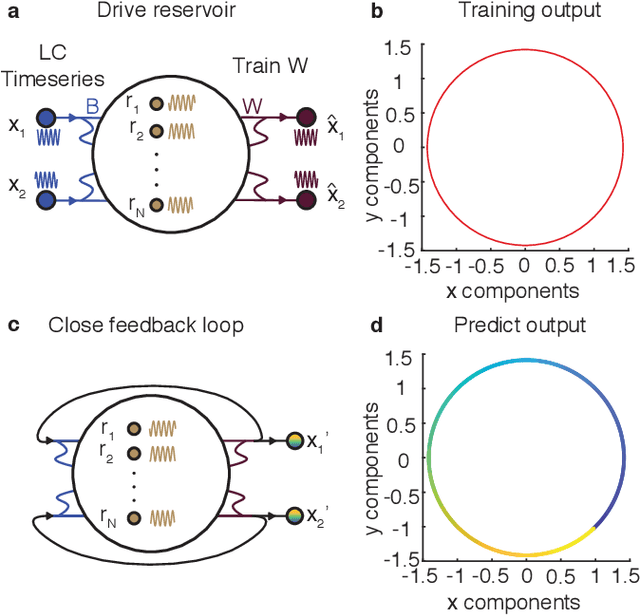
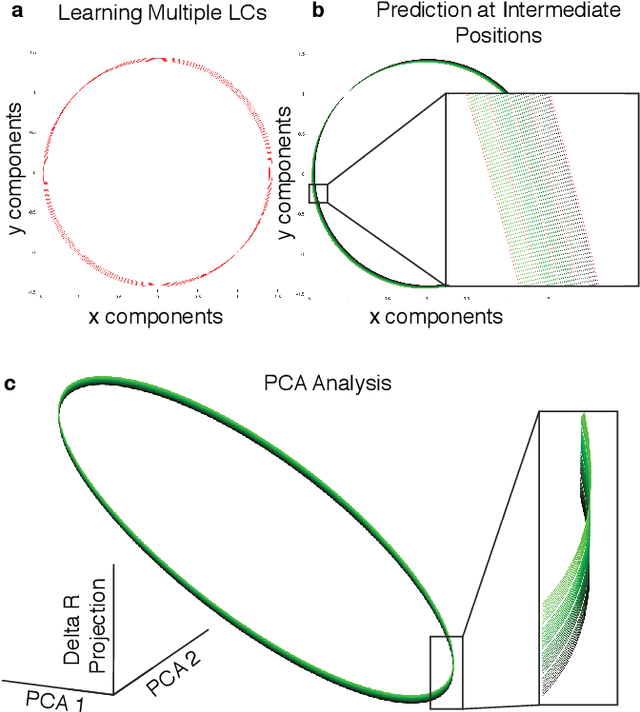
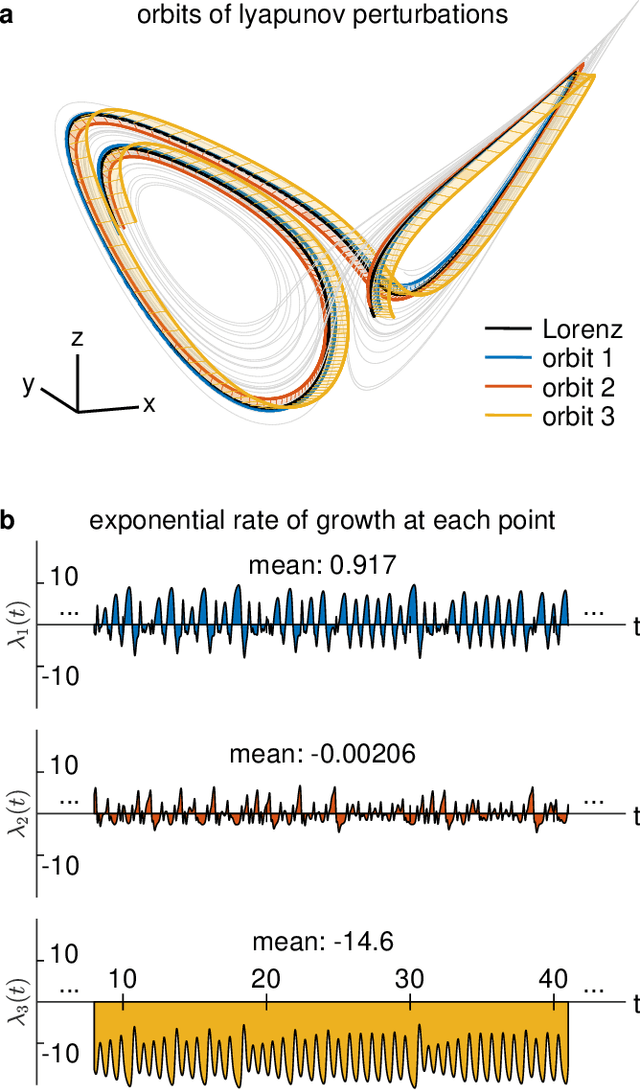
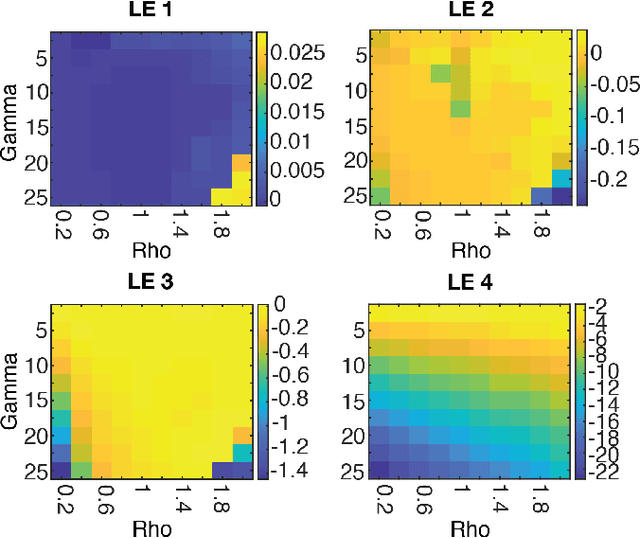
Abstract:Neural systems are well known for their ability to learn and store information as memories. Even more impressive is their ability to abstract these memories to create complex internal representations, enabling advanced functions such as the spatial manipulation of mental representations. While recurrent neural networks (RNNs) are capable of representing complex information, the exact mechanisms of how dynamical neural systems perform abstraction are still not well-understood, thereby hindering the development of more advanced functions. Here, we train a 1000-neuron RNN -- a reservoir computer (RC) -- to abstract a continuous dynamical attractor memory from isolated examples of dynamical attractor memories. Further, we explain the abstraction mechanism with new theory. By training the RC on isolated and shifted examples of either stable limit cycles or chaotic Lorenz attractors, the RC learns a continuum of attractors, as quantified by an extra Lyapunov exponent equal to zero. We propose a theoretical mechanism of this abstraction by combining ideas from differentiable generalized synchronization and feedback dynamics. Our results quantify abstraction in simple neural systems, enabling us to design artificial RNNs for abstraction, and leading us towards a neural basis of abstraction.
Teaching Recurrent Neural Networks to Modify Chaotic Memories by Example
May 03, 2020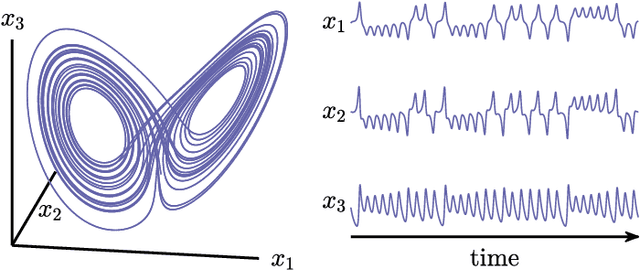
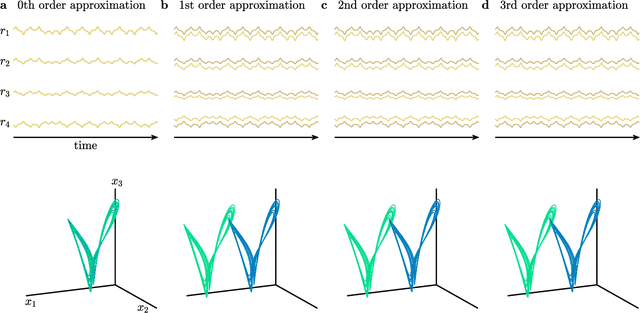
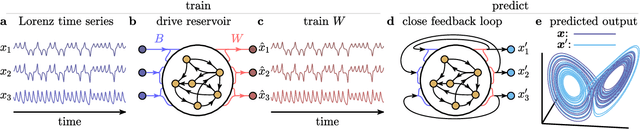
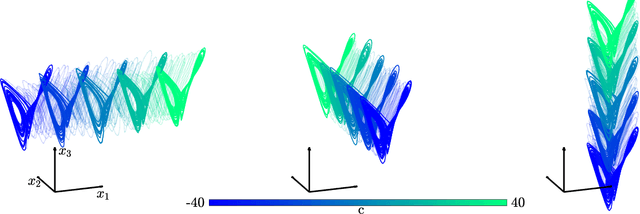
Abstract:The ability to store and manipulate information is a hallmark of computational systems. Whereas computers are carefully engineered to represent and perform mathematical operations on structured data, neurobiological systems perform analogous functions despite flexible organization and unstructured sensory input. Recent efforts have made progress in modeling the representation and recall of information in neural systems. However, precisely how neural systems learn to modify these representations remains far from understood. Here we demonstrate that a recurrent neural network (RNN) can learn to modify its representation of complex information using only examples, and we explain the associated learning mechanism with new theory. Specifically, we drive an RNN with examples of translated, linearly transformed, or pre-bifurcated time series from a chaotic Lorenz system, alongside an additional control signal that changes value for each example. By training the network to replicate the Lorenz inputs, it learns to autonomously evolve about a Lorenz-shaped manifold. Additionally, it learns to continuously interpolate and extrapolate the translation, transformation, and bifurcation of this representation far beyond the training data by changing the control signal. Finally, we provide a mechanism for how these computations are learned, and demonstrate that a single network can simultaneously learn multiple computations. Together, our results provide a simple but powerful mechanism by which an RNN can learn to manipulate internal representations of complex information, allowing for the principled study and precise design of RNNs.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge