Federico Raimondo
for the Alzheimer's Disease Neuroimaging Initiative
Region-wise stacking ensembles for estimating brain-age using MRI
Jan 17, 2025



Abstract:Predictive modeling using structural magnetic resonance imaging (MRI) data is a prominent approach to study brain-aging. Machine learning algorithms and feature extraction methods have been employed to improve predictions and explore healthy and accelerated aging e.g. neurodegenerative and psychiatric disorders. The high-dimensional MRI data pose challenges to building generalizable and interpretable models as well as for data privacy. Common practices are resampling or averaging voxels within predefined parcels, which reduces anatomical specificity and biological interpretability as voxels within a region may differently relate to aging. Effectively, naive fusion by averaging can result in information loss and reduced accuracy. We present a conceptually novel two-level stacking ensemble (SE) approach. The first level comprises regional models for predicting individuals' age based on voxel-wise information, fused by a second-level model yielding final predictions. Eight data fusion scenarios were explored using as input Gray matter volume (GMV) estimates from four datasets covering the adult lifespan. Performance, measured using mean absolute error (MAE), R2, correlation and prediction bias, showed that SE outperformed the region-wise averages. The best performance was obtained when first-level regional predictions were obtained as out-of-sample predictions on the application site with second-level models trained on independent and site-specific data (MAE=4.75 vs baseline regional mean GMV MAE=5.68). Performance improved as more datasets were used for training. First-level predictions showed improved and more robust aging signal providing new biological insights and enhanced data privacy. Overall, the SE improves accuracy compared to the baseline while preserving or enhancing data privacy.
Impact of Leakage on Data Harmonization in Machine Learning Pipelines in Class Imbalance Across Sites
Oct 25, 2024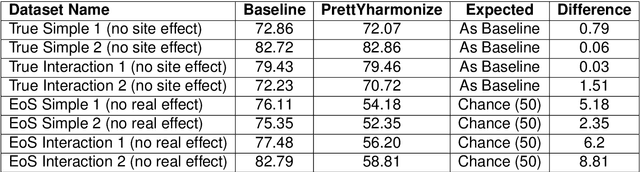
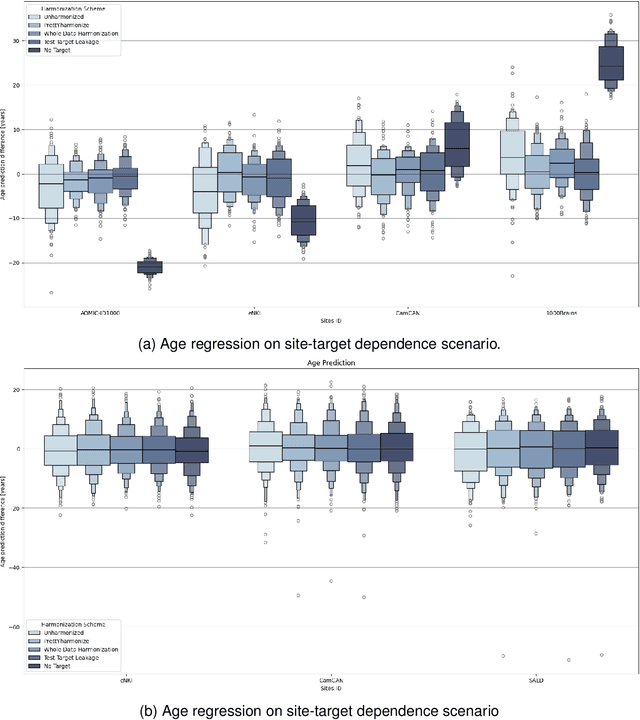
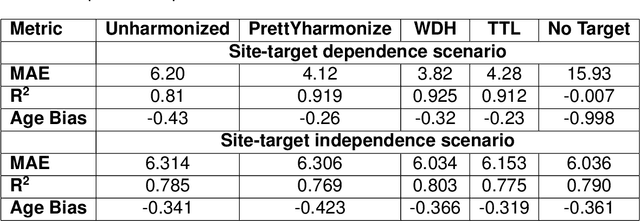
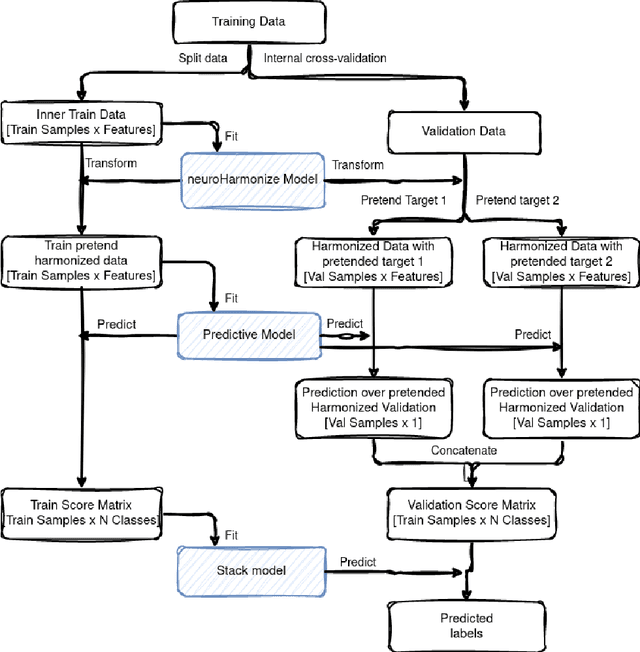
Abstract:Machine learning (ML) models benefit from large datasets. Collecting data in biomedical domains is costly and challenging, hence, combining datasets has become a common practice. However, datasets obtained under different conditions could present undesired site-specific variability. Data harmonization methods aim to remove site-specific variance while retaining biologically relevant information. This study evaluates the effectiveness of popularly used ComBat-based methods for harmonizing data in scenarios where the class balance is not equal across sites. We find that these methods struggle with data leakage issues. To overcome this problem, we propose a novel approach PrettYharmonize, designed to harmonize data by pretending the target labels. We validate our approach using controlled datasets designed to benchmark the utility of harmonization. Finally, using real-world MRI and clinical data, we compare leakage-prone methods with PrettYharmonize and show that it achieves comparable performance while avoiding data leakage, particularly in site-target-dependence scenarios.
On Leakage in Machine Learning Pipelines
Nov 07, 2023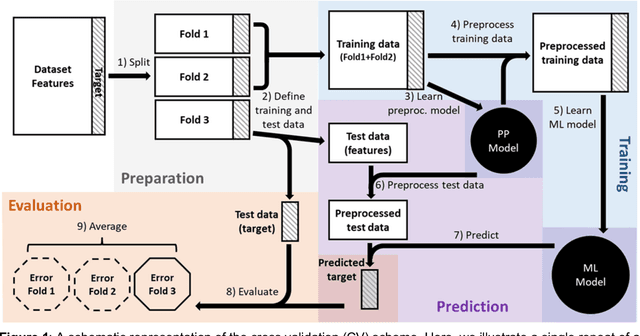
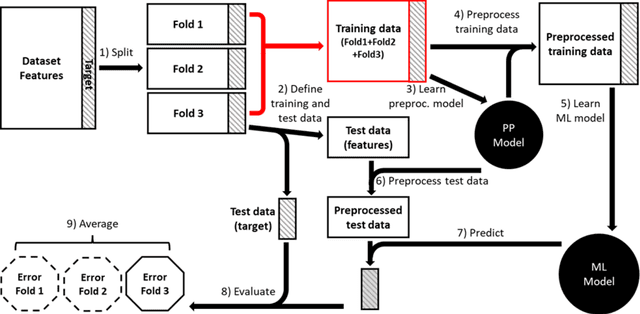
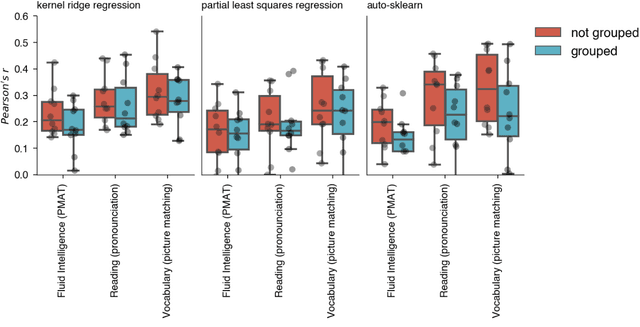
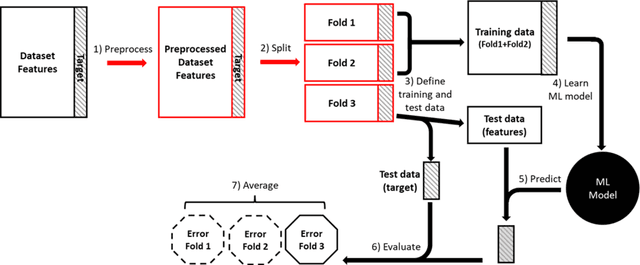
Abstract:Machine learning (ML) provides powerful tools for predictive modeling. ML's popularity stems from the promise of sample-level prediction with applications across a variety of fields from physics and marketing to healthcare. However, if not properly implemented and evaluated, ML pipelines may contain leakage typically resulting in overoptimistic performance estimates and failure to generalize to new data. This can have severe negative financial and societal implications. Our aim is to expand understanding associated with causes leading to leakage when designing, implementing, and evaluating ML pipelines. Illustrated by concrete examples, we provide a comprehensive overview and discussion of various types of leakage that may arise in ML pipelines.
Julearn: an easy-to-use library for leakage-free evaluation and inspection of ML models
Oct 19, 2023Abstract:The fast-paced development of machine learning (ML) methods coupled with its increasing adoption in research poses challenges for researchers without extensive training in ML. In neuroscience, for example, ML can help understand brain-behavior relationships, diagnose diseases, and develop biomarkers using various data sources like magnetic resonance imaging and electroencephalography. The primary objective of ML is to build models that can make accurate predictions on unseen data. Researchers aim to prove the existence of such generalizable models by evaluating performance using techniques such as cross-validation (CV), which uses systematic subsampling to estimate the generalization performance. Choosing a CV scheme and evaluating an ML pipeline can be challenging and, if used improperly, can lead to overestimated results and incorrect interpretations. We created julearn, an open-source Python library, that allow researchers to design and evaluate complex ML pipelines without encountering in common pitfalls. In this manuscript, we present the rationale behind julearn's design, its core features, and showcase three examples of previously-published research projects that can be easily implemented using this novel library. Julearn aims to simplify the entry into the ML world by providing an easy-to-use environment with built in guards against some of the most common ML pitfalls. With its design, unique features and simple interface, it poses as a useful Python-based library for research projects.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge