Ding Xue
DrugOOD: Out-of-Distribution (OOD) Dataset Curator and Benchmark for AI-aided Drug Discovery -- A Focus on Affinity Prediction Problems with Noise Annotations
Jan 24, 2022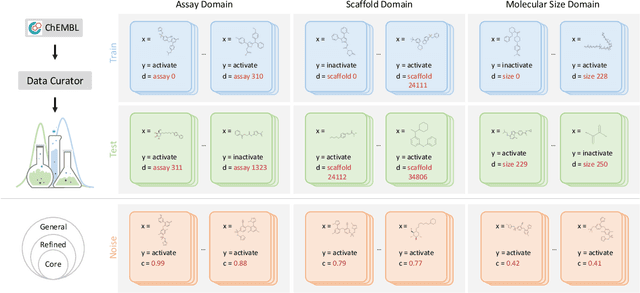
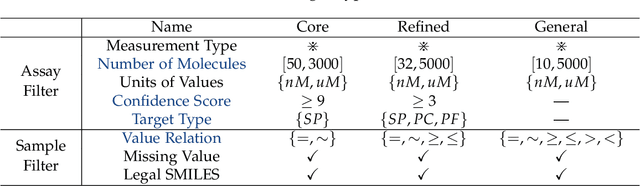

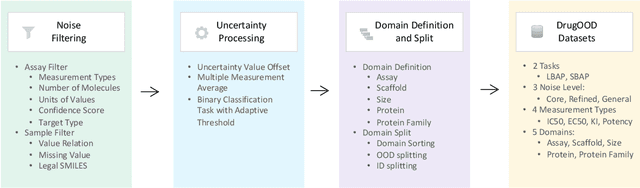
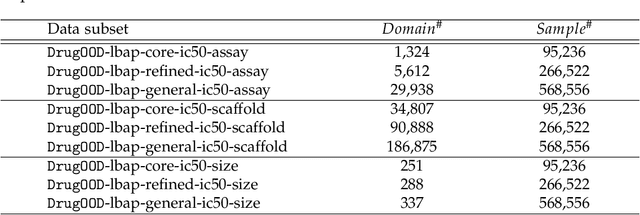
Abstract:AI-aided drug discovery (AIDD) is gaining increasing popularity due to its promise of making the search for new pharmaceuticals quicker, cheaper and more efficient. In spite of its extensive use in many fields, such as ADMET prediction, virtual screening, protein folding and generative chemistry, little has been explored in terms of the out-of-distribution (OOD) learning problem with \emph{noise}, which is inevitable in real world AIDD applications. In this work, we present DrugOOD, a systematic OOD dataset curator and benchmark for AI-aided drug discovery, which comes with an open-source Python package that fully automates the data curation and OOD benchmarking processes. We focus on one of the most crucial problems in AIDD: drug target binding affinity prediction, which involves both macromolecule (protein target) and small-molecule (drug compound). In contrast to only providing fixed datasets, DrugOOD offers automated dataset curator with user-friendly customization scripts, rich domain annotations aligned with biochemistry knowledge, realistic noise annotations and rigorous benchmarking of state-of-the-art OOD algorithms. Since the molecular data is often modeled as irregular graphs using graph neural network (GNN) backbones, DrugOOD also serves as a valuable testbed for \emph{graph OOD learning} problems. Extensive empirical studies have shown a significant performance gap between in-distribution and out-of-distribution experiments, which highlights the need to develop better schemes that can allow for OOD generalization under noise for AIDD.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge