Da-Woon Heo
FIESTA: Fourier-Based Semantic Augmentation with Uncertainty Guidance for Enhanced Domain Generalizability in Medical Image Segmentation
Jun 20, 2024
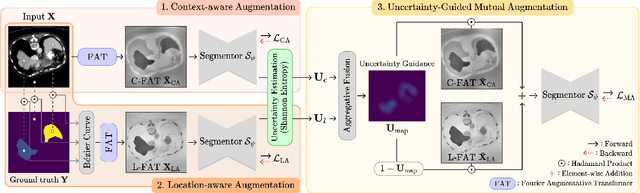
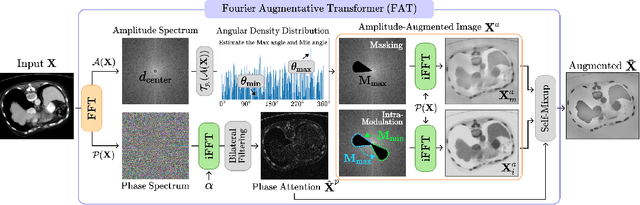
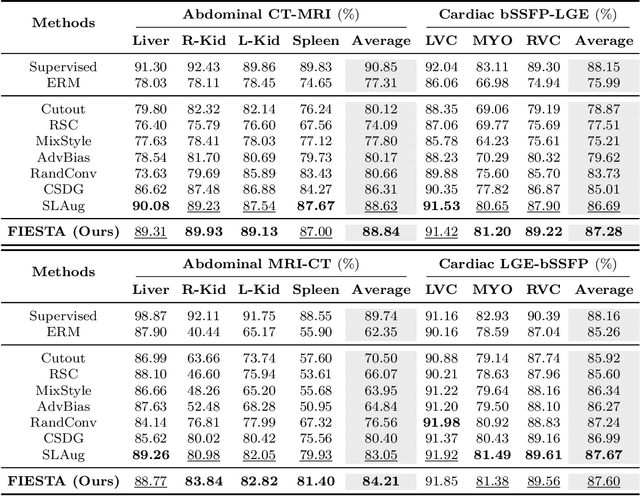
Abstract:Single-source domain generalization (SDG) in medical image segmentation (MIS) aims to generalize a model using data from only one source domain to segment data from an unseen target domain. Despite substantial advances in SDG with data augmentation, existing methods often fail to fully consider the details and uncertain areas prevalent in MIS, leading to mis-segmentation. This paper proposes a Fourier-based semantic augmentation method called FIESTA using uncertainty guidance to enhance the fundamental goals of MIS in an SDG context by manipulating the amplitude and phase components in the frequency domain. The proposed Fourier augmentative transformer addresses semantic amplitude modulation based on meaningful angular points to induce pertinent variations and harnesses the phase spectrum to ensure structural coherence. Moreover, FIESTA employs epistemic uncertainty to fine-tune the augmentation process, improving the ability of the model to adapt to diverse augmented data and concentrate on areas with higher ambiguity. Extensive experiments across three cross-domain scenarios demonstrate that FIESTA surpasses recent state-of-the-art SDG approaches in segmentation performance and significantly contributes to boosting the applicability of the model in medical imaging modalities.
Transferring Ultrahigh-Field Representations for Intensity-Guided Brain Segmentation of Low-Field Magnetic Resonance Imaging
Feb 13, 2024



Abstract:Ultrahigh-field (UHF) magnetic resonance imaging (MRI), i.e., 7T MRI, provides superior anatomical details of internal brain structures owing to its enhanced signal-to-noise ratio and susceptibility-induced contrast. However, the widespread use of 7T MRI is limited by its high cost and lower accessibility compared to low-field (LF) MRI. This study proposes a deep-learning framework that systematically fuses the input LF magnetic resonance feature representations with the inferred 7T-like feature representations for brain image segmentation tasks in a 7T-absent environment. Specifically, our adaptive fusion module aggregates 7T-like features derived from the LF image by a pre-trained network and then refines them to be effectively assimilable UHF guidance into LF image features. Using intensity-guided features obtained from such aggregation and assimilation, segmentation models can recognize subtle structural representations that are usually difficult to recognize when relying only on LF features. Beyond such advantages, this strategy can seamlessly be utilized by modulating the contrast of LF features in alignment with UHF guidance, even when employing arbitrary segmentation models. Exhaustive experiments demonstrated that the proposed method significantly outperformed all baseline models on both brain tissue and whole-brain segmentation tasks; further, it exhibited remarkable adaptability and scalability by successfully integrating diverse segmentation models and tasks. These improvements were not only quantifiable but also visible in the superlative visual quality of segmentation masks.
A Quantitatively Interpretable Model for Alzheimer's Disease Prediction Using Deep Counterfactuals
Oct 05, 2023



Abstract:Deep learning (DL) for predicting Alzheimer's disease (AD) has provided timely intervention in disease progression yet still demands attentive interpretability to explain how their DL models make definitive decisions. Recently, counterfactual reasoning has gained increasing attention in medical research because of its ability to provide a refined visual explanatory map. However, such visual explanatory maps based on visual inspection alone are insufficient unless we intuitively demonstrate their medical or neuroscientific validity via quantitative features. In this study, we synthesize the counterfactual-labeled structural MRIs using our proposed framework and transform it into a gray matter density map to measure its volumetric changes over the parcellated region of interest (ROI). We also devised a lightweight linear classifier to boost the effectiveness of constructed ROIs, promoted quantitative interpretation, and achieved comparable predictive performance to DL methods. Throughout this, our framework produces an ``AD-relatedness index'' for each ROI and offers an intuitive understanding of brain status for an individual patient and across patient groups with respect to AD progression.
Medical Transformer: Universal Brain Encoder for 3D MRI Analysis
Apr 28, 2021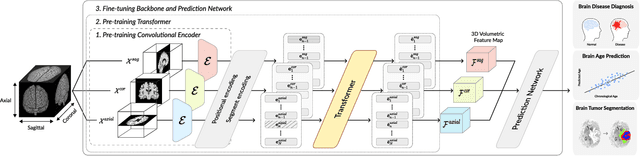
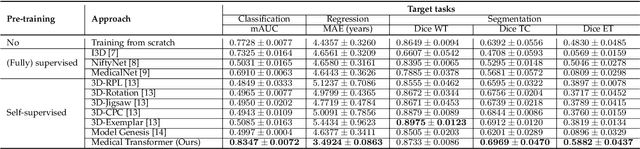
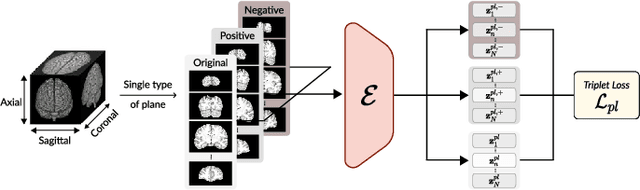
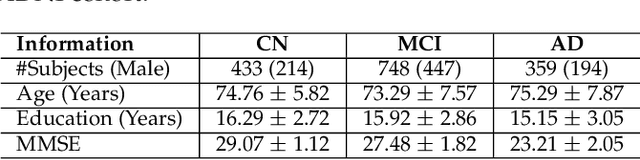
Abstract:Transfer learning has gained attention in medical image analysis due to limited annotated 3D medical datasets for training data-driven deep learning models in the real world. Existing 3D-based methods have transferred the pre-trained models to downstream tasks, which achieved promising results with only a small number of training samples. However, they demand a massive amount of parameters to train the model for 3D medical imaging. In this work, we propose a novel transfer learning framework, called Medical Transformer, that effectively models 3D volumetric images in the form of a sequence of 2D image slices. To make a high-level representation in 3D-form empowering spatial relations better, we take a multi-view approach that leverages plenty of information from the three planes of 3D volume, while providing parameter-efficient training. For building a source model generally applicable to various tasks, we pre-train the model in a self-supervised learning manner for masked encoding vector prediction as a proxy task, using a large-scale normal, healthy brain magnetic resonance imaging (MRI) dataset. Our pre-trained model is evaluated on three downstream tasks: (i) brain disease diagnosis, (ii) brain age prediction, and (iii) brain tumor segmentation, which are actively studied in brain MRI research. The experimental results show that our Medical Transformer outperforms the state-of-the-art transfer learning methods, efficiently reducing the number of parameters up to about 92% for classification and
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge