Chaowen Hu
From $\boldsymbol{\logπ}$ to $\boldsymbolπ$: Taming Divergence in Soft Clipping via Bilateral Decoupled Decay of Probability Gradient Weight
Mar 15, 2026Abstract:Reinforcement Learning with Verifiable Rewards (RLVR) has catalyzed a leap in Large Language Model (LLM) reasoning, yet its optimization dynamics remain fragile. Standard algorithms like GRPO enforce stability via ``hard clipping'', which inadvertently stifles exploration by discarding gradients of tokens outside the trust region. While recent ``soft clipping'' methods attempt to recover these gradients, they suffer from a critical challenge: relying on log-probability gradient ($\nabla_θ\log π_θ$) yields divergent weights as probabilities vanish, destabilizing LLM training. We rethink this convention by establishing probability gradient ($\nabla_θπ_θ$) as the superior optimization primitive. Accordingly, we propose Decoupled Gradient Policy Optimization (DGPO), which employs a decoupled decay mechanism based on importance sampling ratios. By applying asymmetric, continuous decay to boundary tokens, DGPO resolves the conflict between stability and sustained exploration. Extensive experiments across DeepSeek-R1-Distill-Qwen series models (1.5B/7B/14B) demonstrate that DGPO consistently outperforms strong baselines on various mathematical benchmarks, offering a robust and scalable solution for RLVR. Our code and implementation are available at: https://github.com/VenomRose-Juri/DGPO-RL.
How to Allocate, How to Learn? Dynamic Rollout Allocation and Advantage Modulation for Policy Optimization
Feb 22, 2026Abstract:Reinforcement Learning with Verifiable Rewards (RLVR) has proven effective for Large Language Model (LLM) reasoning, yet current methods face key challenges in resource allocation and policy optimization dynamics: (i) uniform rollout allocation ignores gradient variance heterogeneity across problems, and (ii) the softmax policy structure causes gradient attenuation for high-confidence correct actions, while excessive gradient updates may destabilize training. Therefore, we propose DynaMO, a theoretically-grounded dual-pronged optimization framework. At the sequence level, we prove that uniform allocation is suboptimal and derive variance-minimizing allocation from the first principle, establishing Bernoulli variance as a computable proxy for gradient informativeness. At the token level, we develop gradient-aware advantage modulation grounded in theoretical analysis of gradient magnitude bounds. Our framework compensates for gradient attenuation of high-confidence correct actions while utilizing entropy changes as computable indicators to stabilize excessive update magnitudes. Extensive experiments conducted on a diverse range of mathematical reasoning benchmarks demonstrate consistent improvements over strong RLVR baselines. Our implementation is available at: \href{https://anonymous.4open.science/r/dynamo-680E/README.md}{https://anonymous.4open.science/r/dynamo}.
MASPO: Unifying Gradient Utilization, Probability Mass, and Signal Reliability for Robust and Sample-Efficient LLM Reasoning
Feb 19, 2026Abstract:Existing Reinforcement Learning with Verifiable Rewards (RLVR) algorithms, such as GRPO, rely on rigid, uniform, and symmetric trust region mechanisms that are fundamentally misaligned with the complex optimization dynamics of Large Language Models (LLMs). In this paper, we identify three critical challenges in these methods: (1) inefficient gradient utilization caused by the binary cutoff of hard clipping, (2) insensitive probability mass arising from uniform ratio constraints that ignore the token distribution, and (3) asymmetric signal reliability stemming from the disparate credit assignment ambiguity between positive and negative samples. To bridge these gaps, we propose Mass-Adaptive Soft Policy Optimization (MASPO), a unified framework designed to harmonize these three dimensions. MASPO integrates a differentiable soft Gaussian gating to maximize gradient utility, a mass-adaptive limiter to balance exploration across the probability spectrum, and an asymmetric risk controller to align update magnitudes with signal confidence. Extensive evaluations demonstrate that MASPO serves as a robust, all-in-one RLVR solution, significantly outperforming strong baselines. Our code is available at: https://anonymous.4open.science/r/ma1/README.md.
SERVAL: Synergy Learning between Vertical Models and LLMs towards Oracle-Level Zero-shot Medical Prediction
Mar 03, 2024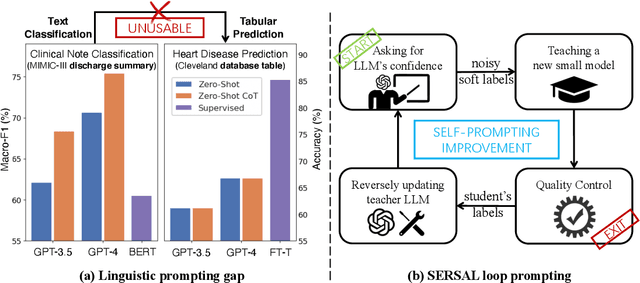

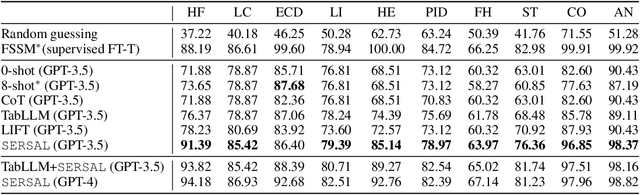
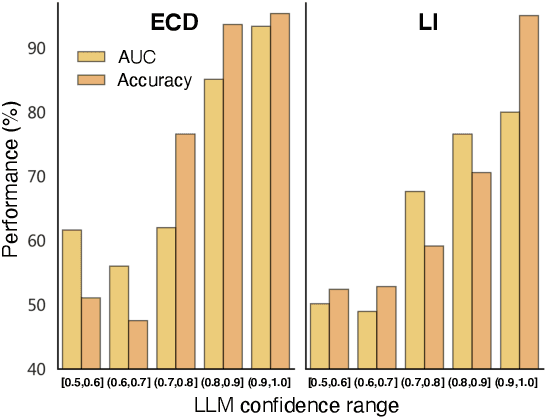
Abstract:Recent development of large language models (LLMs) has exhibited impressive zero-shot proficiency on generic and common sense questions. However, LLMs' application on domain-specific vertical questions still lags behind, primarily due to the humiliation problems and deficiencies in vertical knowledge. Furthermore, the vertical data annotation process often requires labor-intensive expert involvement, thereby presenting an additional challenge in enhancing the model's vertical capabilities. In this paper, we propose SERVAL, a synergy learning pipeline designed for unsupervised development of vertical capabilities in both LLMs and small models by mutual enhancement. Specifically, SERVAL utilizes the LLM's zero-shot outputs as annotations, leveraging its confidence to teach a robust vertical model from scratch. Reversely, the trained vertical model guides the LLM fine-tuning to enhance its zero-shot capability, progressively improving both models through an iterative process. In medical domain, known for complex vertical knowledge and costly annotations, comprehensive experiments show that, without access to any gold labels, SERVAL with the synergy learning of OpenAI GPT-3.5 and a simple model attains fully-supervised competitive performance across ten widely used medical datasets. These datasets represent vertically specialized medical diagnostic scenarios (e.g., diabetes, heart diseases, COVID-19), highlighting the potential of SERVAL in refining the vertical capabilities of LLMs and training vertical models from scratch, all achieved without the need for annotations.
Sample-efficient Multi-objective Molecular Optimization with GFlowNets
Feb 08, 2023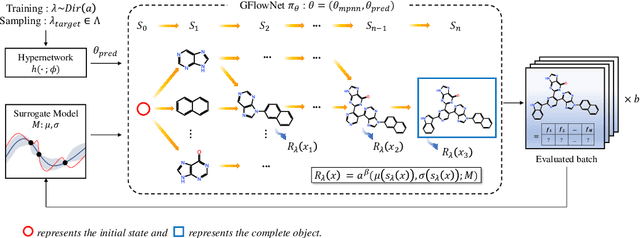
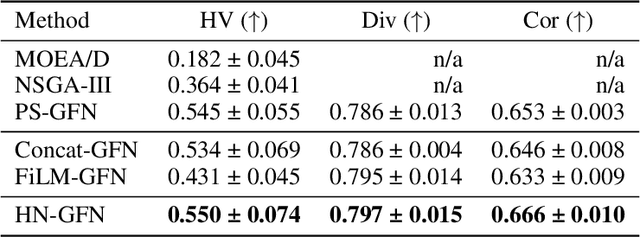
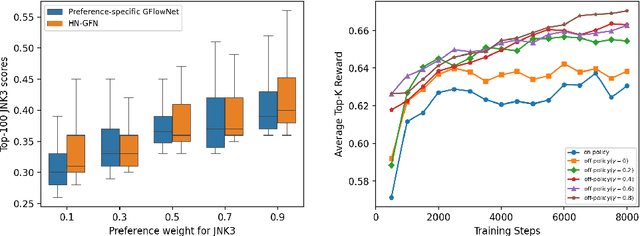
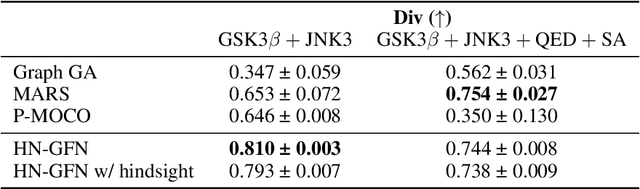
Abstract:Many crucial scientific problems involve designing novel molecules with desired properties, which can be formulated as an expensive black-box optimization problem over the discrete chemical space. Computational methods have achieved initial success but still struggle with simultaneously optimizing multiple competing properties in a sample-efficient manner. In this work, we propose a multi-objective Bayesian optimization (MOBO) algorithm leveraging the hypernetwork-based GFlowNets (HN-GFN) as an acquisition function optimizer, with the purpose of sampling a diverse batch of candidate molecular graphs from an approximate Pareto front. Using a single preference-conditioned hypernetwork, HN-GFN learns to explore various trade-offs between objectives. Inspired by reinforcement learning, we further propose a hindsight-like off-policy strategy to share high-performing molecules among different preferences in order to speed up learning for HN-GFN. Through synthetic experiments, we illustrate that HN-GFN has adequate capacity to generalize over preferences. Extensive experiments show that our framework outperforms the best baselines by a large margin in terms of hypervolume in various real-world MOBO settings.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge