Cesar César Quilodrán-Casas
T3D: Towards 3D Medical Image Understanding through Vision-Language Pre-training
Dec 05, 2023
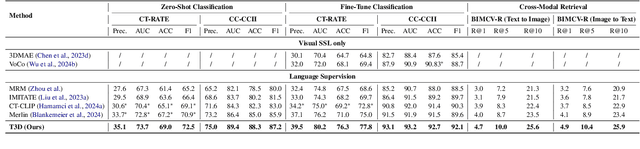
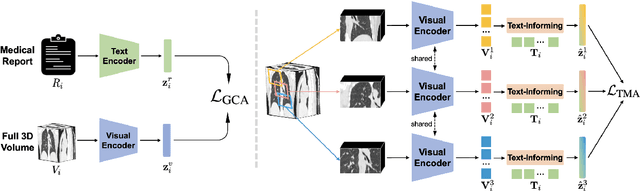
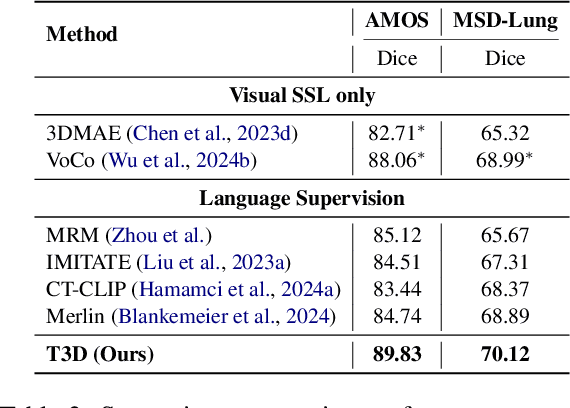
Abstract:Expert annotation of 3D medical image for downstream analysis is resource-intensive, posing challenges in clinical applications. Visual self-supervised learning (vSSL), though effective for learning visual invariance, neglects the incorporation of domain knowledge from medicine. To incorporate medical knowledge into visual representation learning, vision-language pre-training (VLP) has shown promising results in 2D image. However, existing VLP approaches become generally impractical when applied to high-resolution 3D medical images due to GPU hardware constraints and the potential loss of critical details caused by downsampling, which is the intuitive solution to hardware constraints. To address the above limitations, we introduce T3D, the first VLP framework designed for high-resolution 3D medical images. T3D incorporates two text-informed pretext tasks: (\lowerromannumeral{1}) text-informed contrastive learning; (\lowerromannumeral{2}) text-informed image restoration. These tasks focus on learning 3D visual representations from high-resolution 3D medical images and integrating clinical knowledge from radiology reports, without distorting information through forced alignment of downsampled volumes with detailed anatomical text. Trained on a newly curated large-scale dataset of 3D medical images and radiology reports, T3D significantly outperforms current vSSL methods in tasks like organ and tumor segmentation, as well as disease classification. This underlines T3D's potential in representation learning for 3D medical image analysis. All data and code will be available upon acceptance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge