Benjamin M. Smith
Multi-View Transformers for Airway-To-Lung Ratio Inference on Cardiac CT Scans: The C4R Study
Jan 15, 2025
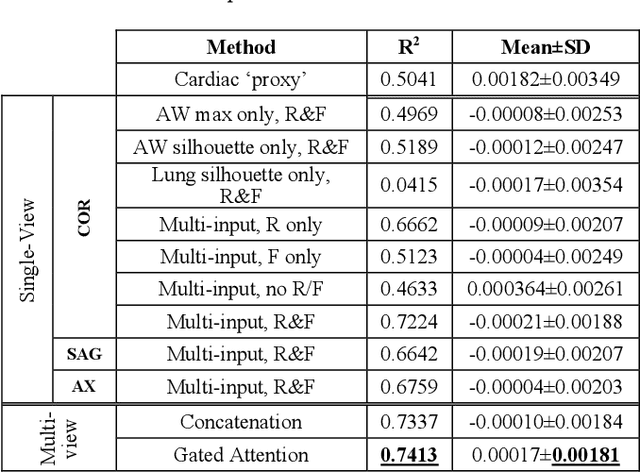
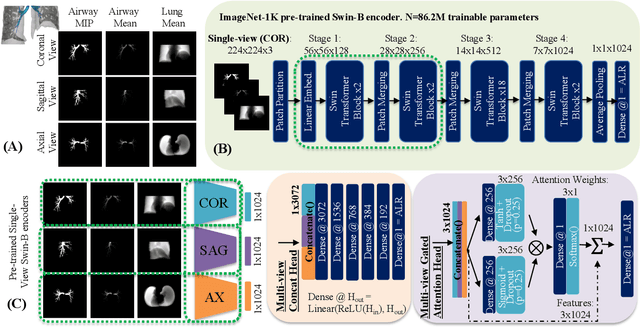
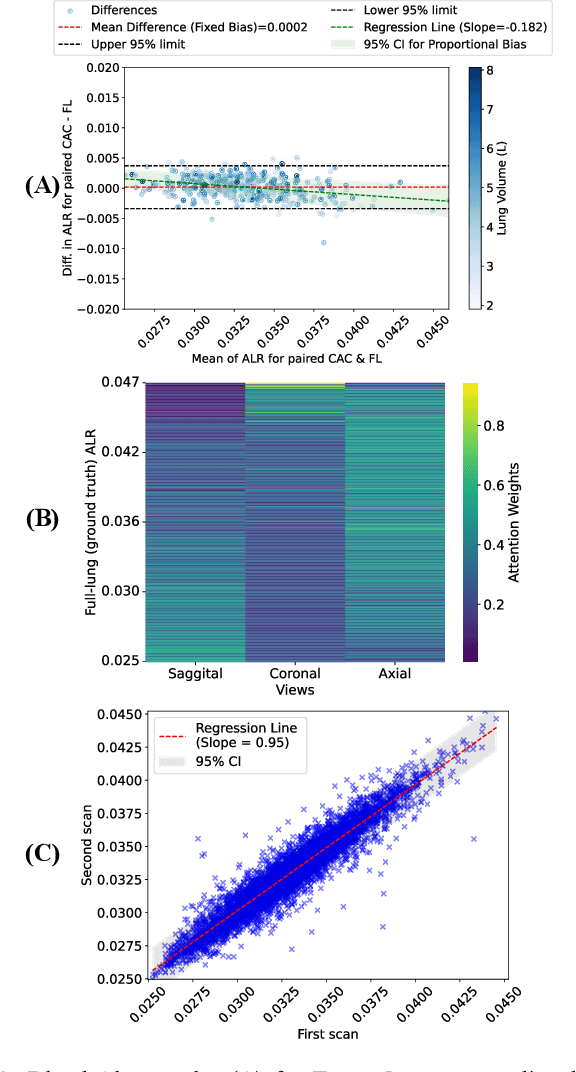
Abstract:The ratio of airway tree lumen to lung size (ALR), assessed at full inspiration on high resolution full-lung computed tomography (CT), is a major risk factor for chronic obstructive pulmonary disease (COPD). There is growing interest to infer ALR from cardiac CT images, which are widely available in epidemiological cohorts, to investigate the relationship of ALR to severe COVID-19 and post-acute sequelae of SARS-CoV-2 infection (PASC). Previously, cardiac scans included approximately 2/3 of the total lung volume with 5-6x greater slice thickness than high-resolution (HR) full-lung (FL) CT. In this study, we present a novel attention-based Multi-view Swin Transformer to infer FL ALR values from segmented cardiac CT scans. For the supervised training we exploit paired full-lung and cardiac CTs acquired in the Multi-Ethnic Study of Atherosclerosis (MESA). Our network significantly outperforms a proxy direct ALR inference on segmented cardiac CT scans and achieves accuracy and reproducibility comparable with a scan-rescan reproducibility of the FL ALR ground-truth.
Robust Quantification of Percent Emphysema on CT via Domain Attention: the Multi-Ethnic Study of Atherosclerosis (MESA) Lung Study
Mar 06, 2024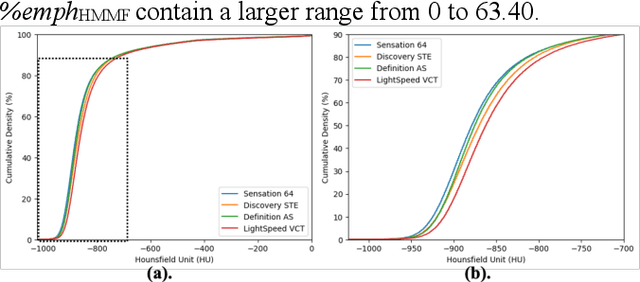
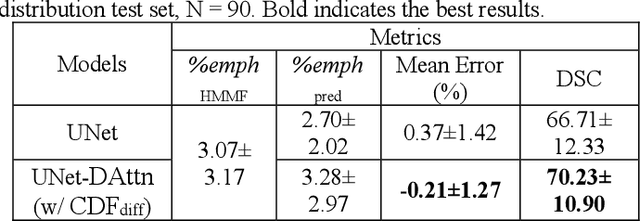
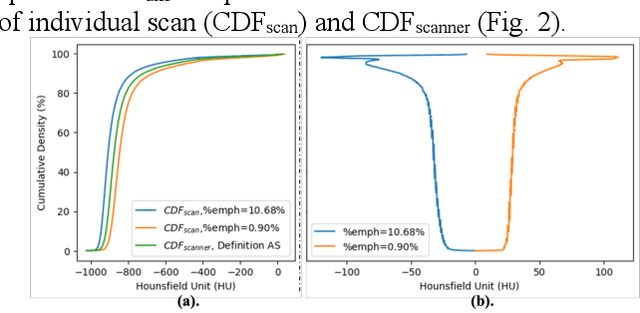
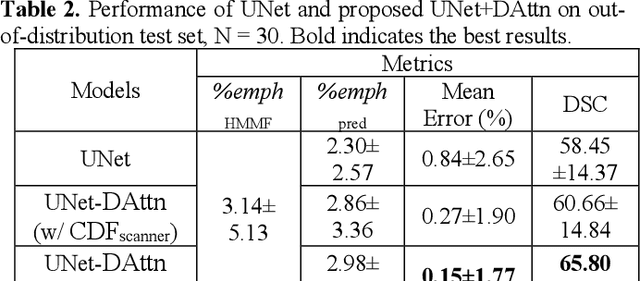
Abstract:Robust quantification of pulmonary emphysema on computed tomography (CT) remains challenging for large-scale research studies that involve scans from different scanner types and for translation to clinical scans. Existing studies have explored several directions to tackle this challenge, including density correction, noise filtering, regression, hidden Markov measure field (HMMF) model-based segmentation, and volume-adjusted lung density. Despite some promising results, previous studies either required a tedious workflow or limited opportunities for downstream emphysema subtyping, limiting efficient adaptation on a large-scale study. To alleviate this dilemma, we developed an end-to-end deep learning framework based on an existing HMMF segmentation framework. We first demonstrate that a regular UNet cannot replicate the existing HMMF results because of the lack of scanner priors. We then design a novel domain attention block to fuse image feature with quantitative scanner priors which significantly improves the results.
Unsupervised Airway Tree Clustering with Deep Learning: The Multi-Ethnic Study of Atherosclerosis (MESA) Lung Study
Feb 28, 2024
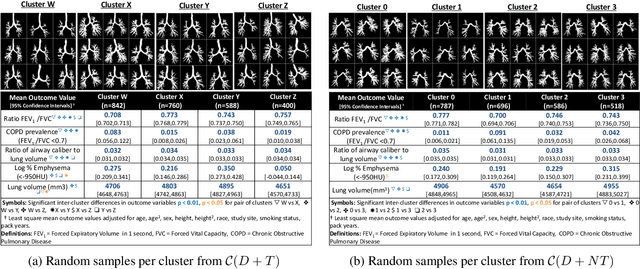



Abstract:High-resolution full lung CT scans now enable the detailed segmentation of airway trees up to the 6th branching generation. The airway binary masks display very complex tree structures that may encode biological information relevant to disease risk and yet remain challenging to exploit via traditional methods such as meshing or skeletonization. Recent clinical studies suggest that some variations in shape patterns and caliber of the human airway tree are highly associated with adverse health outcomes, including all-cause mortality and incident COPD. However, quantitative characterization of variations observed on CT segmented airway tree remain incomplete, as does our understanding of the clinical and developmental implications of such. In this work, we present an unsupervised deep-learning pipeline for feature extraction and clustering of human airway trees, learned directly from projections of 3D airway segmentations. We identify four reproducible and clinically distinct airway sub-types in the MESA Lung CT cohort.
Novel Subtypes of Pulmonary Emphysema Based on Spatially-Informed Lung Texture Learning
Jul 09, 2020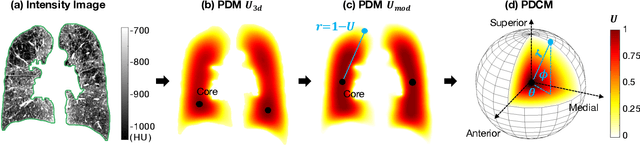
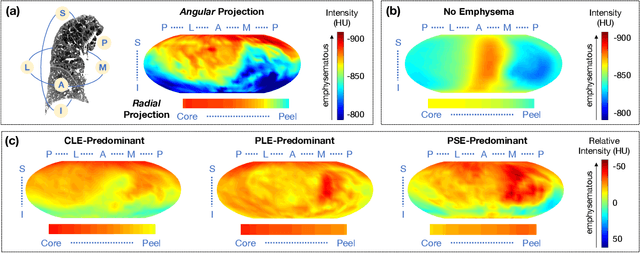
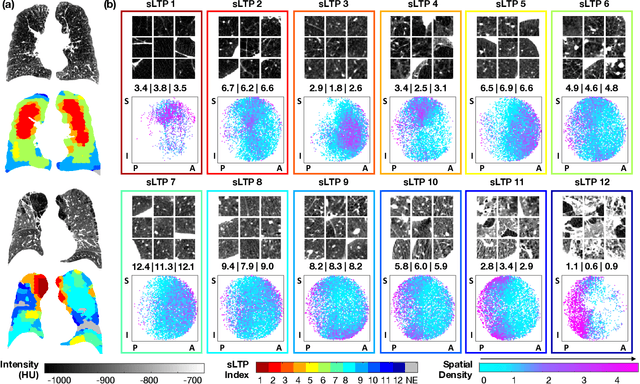
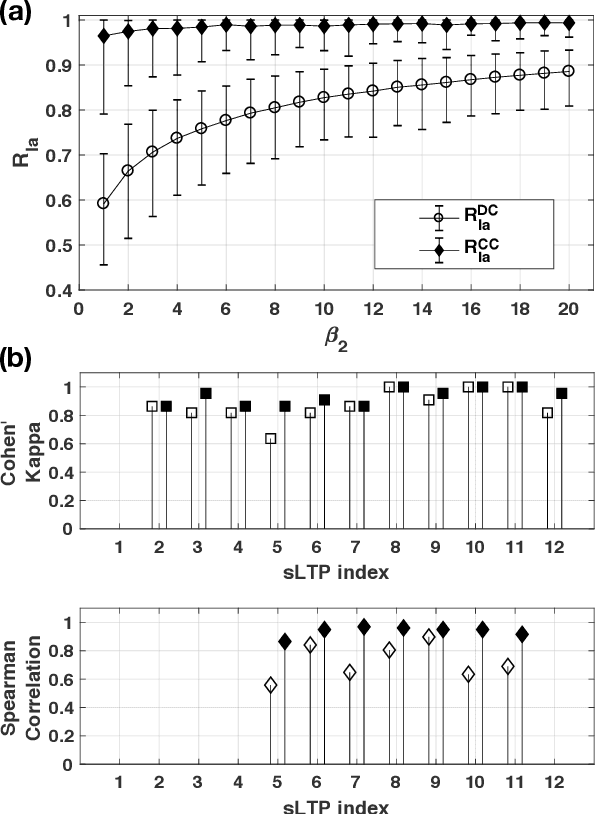
Abstract:Pulmonary emphysema overlaps considerably with chronic obstructive pulmonary disease (COPD), and is traditionally subcategorized into three subtypes previously identified on autopsy. Unsupervised learning of emphysema subtypes on computed tomography (CT) opens the way to new definitions of emphysema subtypes and eliminates the need of thorough manual labeling. However, CT-based emphysema subtypes have been limited to texture-based patterns without considering spatial location. In this work, we introduce a standardized spatial mapping of the lung for quantitative study of lung texture location, and propose a novel framework for combining spatial and texture information to discover spatially-informed lung texture patterns (sLTPs) that represent novel emphysema subtypes. Exploiting two cohorts of full-lung CT scans from the MESA COPD and EMCAP studies, we first show that our spatial mapping enables population-wide study of emphysema spatial location. We then evaluate the characteristics of the sLTPs discovered on MESA COPD, and show that they are reproducible, able to encode standard emphysema subtypes, and associated with physiological symptoms.
Explaining Radiological Emphysema Subtypes with Unsupervised Texture Prototypes: MESA COPD Study
Dec 05, 2016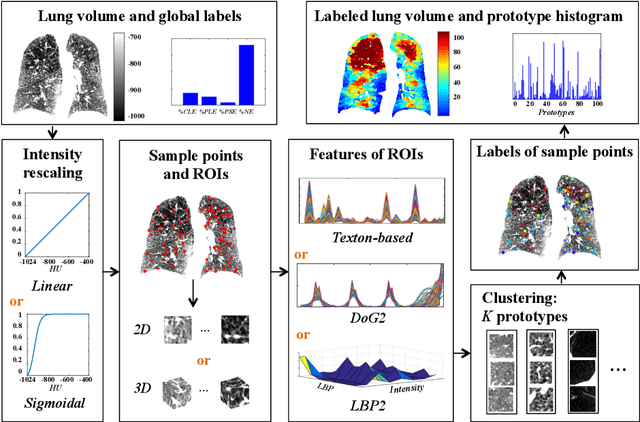
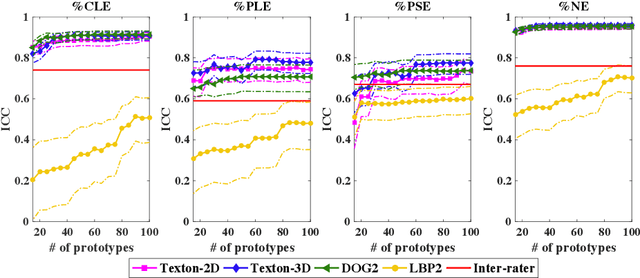
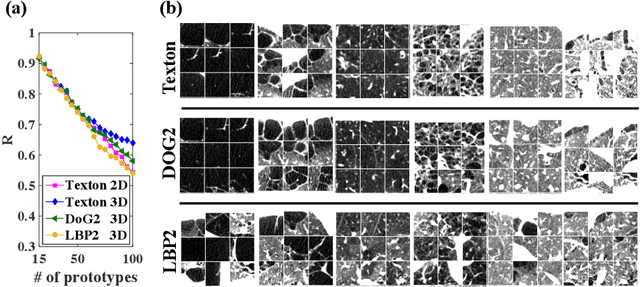

Abstract:Pulmonary emphysema is traditionally subcategorized into three subtypes, which have distinct radiological appearances on computed tomography (CT) and can help with the diagnosis of chronic obstructive pulmonary disease (COPD). Automated texture-based quantification of emphysema subtypes has been successfully implemented via supervised learning of these three emphysema subtypes. In this work, we demonstrate that unsupervised learning on a large heterogeneous database of CT scans can generate texture prototypes that are visually homogeneous and distinct, reproducible across subjects, and capable of predicting accurately the three standard radiological subtypes. These texture prototypes enable automated labeling of lung volumes, and open the way to new interpretations of lung CT scans with finer subtyping of emphysema.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge