David A. Bluemke
Deep Learning-based Automated Aortic Area and Distensibility Assessment: The Multi-Ethnic Study of Atherosclerosis
Mar 03, 2021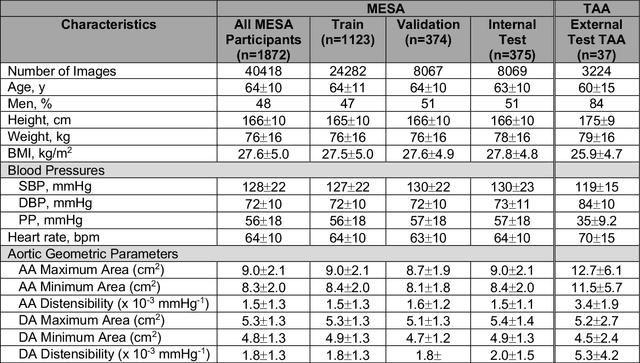
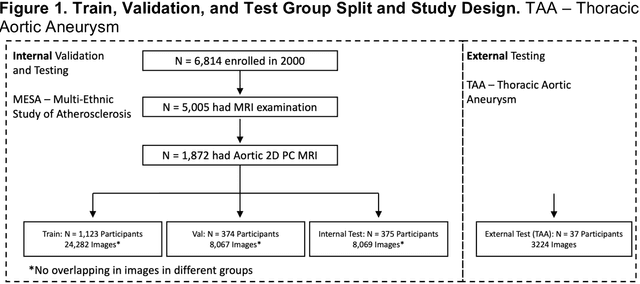


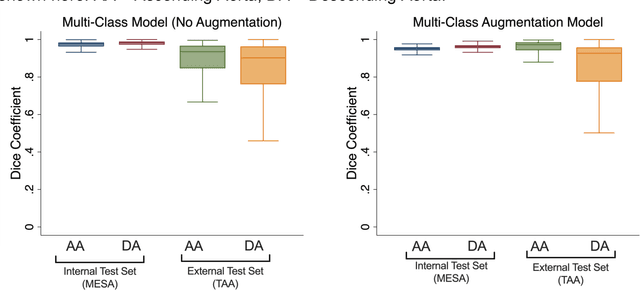
Abstract:This study applies convolutional neural network (CNN)-based automatic segmentation and distensibility measurement of the ascending and descending aorta from 2D phase-contrast cine magnetic resonance imaging (PC-cine MRI) within the large MESA cohort with subsequent assessment on an external cohort of thoracic aortic aneurysm (TAA) patients. 2D PC-cine MRI images of the ascending and descending aorta at the pulmonary artery bifurcation from the MESA study were included. Train, validation, and internal test sets consisted of 1123 studies (24282 images), 374 studies (8067 images), and 375 studies (8069 images), respectively. An external test set of TAAs consisted of 37 studies (3224 images). A U-Net based CNN was constructed, and performance was evaluated utilizing dice coefficient (for segmentation) and concordance correlation coefficients (CCC) of aortic geometric parameters by comparing to manual segmentation and parameter estimation. Dice coefficients for aorta segmentation were 97.6% (CI: 97.5%-97.6%) and 93.6% (84.6%-96.7%) on the internal and external test of TAAs, respectively. CCC for comparison of manual and CNN maximum and minimum ascending aortic areas were 0.97 and 0.95, respectively, on the internal test set and 0.997 and 0.995, respectively, for the external test. CCCs for maximum and minimum descending aortic areas were 0.96 and 0. 98, respectively, on the internal test set and 0.93 and 0.93, respectively, on the external test set. We successfully developed and validated a U-Net based ascending and descending aortic segmentation and distensibility quantification model in a large multi-ethnic database and in an external cohort of TAA patients.
Ω-Net : Fully Automatic, Multi-View Cardiac MR Detection, Orientation, and Segmentation with Deep Neural Networks
Mar 20, 2018
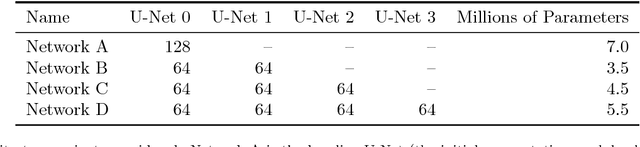

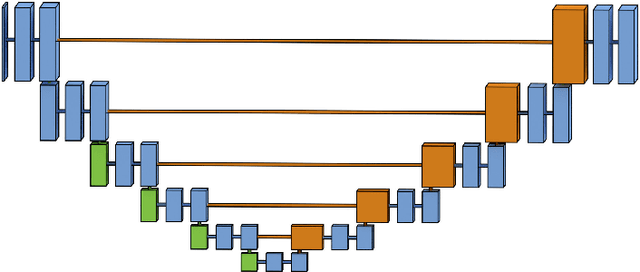
Abstract:Pixelwise segmentation of the left ventricular (LV) myocardium and the four cardiac chambers in 2-D steady state free precession (SSFP) cine sequences is an essential preprocessing step for a wide range of analyses. Variability in contrast, appearance, orientation, and placement of the heart between patients, clinical views, scanners, and protocols makes fully automatic semantic segmentation a notoriously difficult problem. Here, we present ${\Omega}$-Net (Omega-Net): a novel convolutional neural network (CNN) architecture for simultaneous localization, transformation into a canonical orientation, and semantic segmentation. First, an initial segmentation is performed on the input image, second, the features learned during this initial segmentation are used to predict the parameters needed to transform the input image into a canonical orientation, and third, a final segmentation is performed on the transformed image. In this work, ${\Omega}$-Nets of varying depths were trained to detect five foreground classes in any of three clinical views (short axis, SA, four-chamber, 4C, two-chamber, 2C), without prior knowledge of the view being segmented. The architecture was trained on a cohort of patients with hypertrophic cardiomyopathy and healthy control subjects. Network performance as measured by weighted foreground intersection-over-union (IoU) was substantially improved in the best-performing ${\Omega}$- Net compared with U-Net segmentation without localization or orientation. In addition, {\Omega}-Net was retrained from scratch on the 2017 MICCAI ACDC dataset, and achieves state-of-the-art results on the LV and RV bloodpools, and performed slightly worse in segmentation of the LV myocardium. We conclude this architecture represents a substantive advancement over prior approaches, with implications for biomedical image segmentation more generally.
Feature Tracking Cardiac Magnetic Resonance via Deep Learning and Spline Optimization
Apr 12, 2017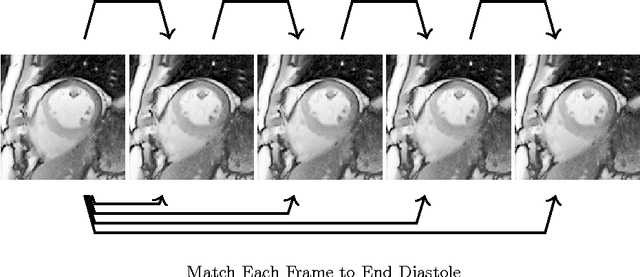

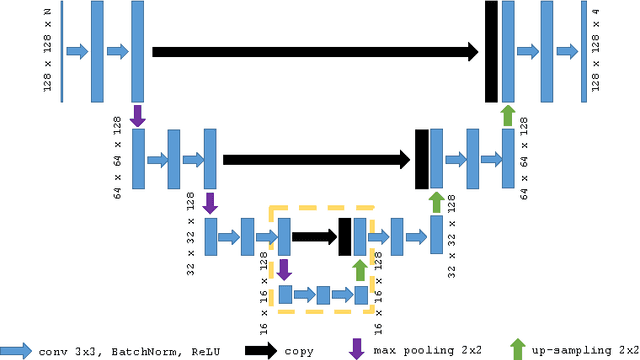

Abstract:Feature tracking Cardiac Magnetic Resonance (CMR) has recently emerged as an area of interest for quantification of regional cardiac function from balanced, steady state free precession (SSFP) cine sequences. However, currently available techniques lack full automation, limiting reproducibility. We propose a fully automated technique whereby a CMR image sequence is first segmented with a deep, fully convolutional neural network (CNN) architecture, and quadratic basis splines are fitted simultaneously across all cardiac frames using least squares optimization. Experiments are performed using data from 42 patients with hypertrophic cardiomyopathy (HCM) and 21 healthy control subjects. In terms of segmentation, we compared state-of-the-art CNN frameworks, U-Net and dilated convolution architectures, with and without temporal context, using cross validation with three folds. Performance relative to expert manual segmentation was similar across all networks: pixel accuracy was ~97%, intersection-over-union (IoU) across all classes was ~87%, and IoU across foreground classes only was ~85%. Endocardial left ventricular circumferential strain calculated from the proposed pipeline was significantly different in control and disease subjects (-25.3% vs -29.1%, p = 0.006), in agreement with the current clinical literature.
Explaining Radiological Emphysema Subtypes with Unsupervised Texture Prototypes: MESA COPD Study
Dec 05, 2016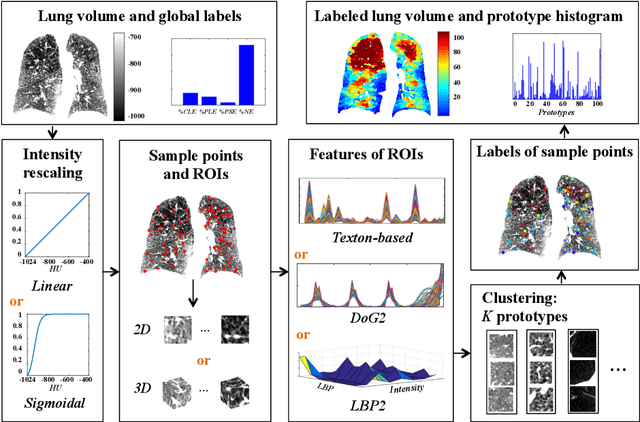
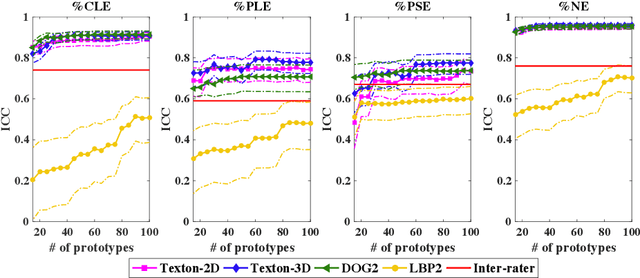
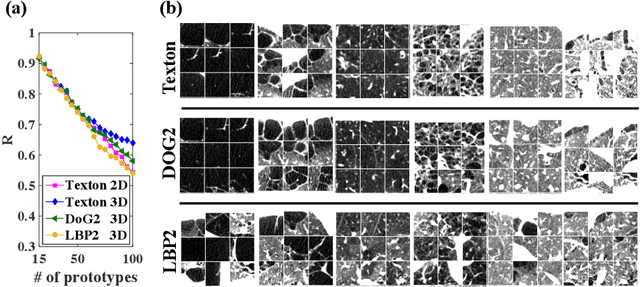

Abstract:Pulmonary emphysema is traditionally subcategorized into three subtypes, which have distinct radiological appearances on computed tomography (CT) and can help with the diagnosis of chronic obstructive pulmonary disease (COPD). Automated texture-based quantification of emphysema subtypes has been successfully implemented via supervised learning of these three emphysema subtypes. In this work, we demonstrate that unsupervised learning on a large heterogeneous database of CT scans can generate texture prototypes that are visually homogeneous and distinct, reproducible across subjects, and capable of predicting accurately the three standard radiological subtypes. These texture prototypes enable automated labeling of lung volumes, and open the way to new interpretations of lung CT scans with finer subtyping of emphysema.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge