Ayagoz Mussabayeva
Image Registration and Predictive Modeling: Learning the Metric on the Space of Diffeomorphisms
Aug 10, 2018

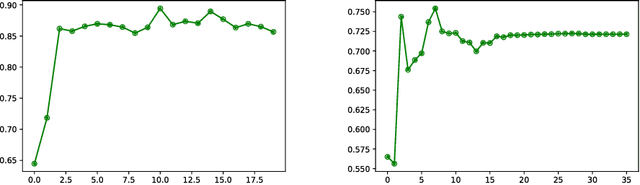
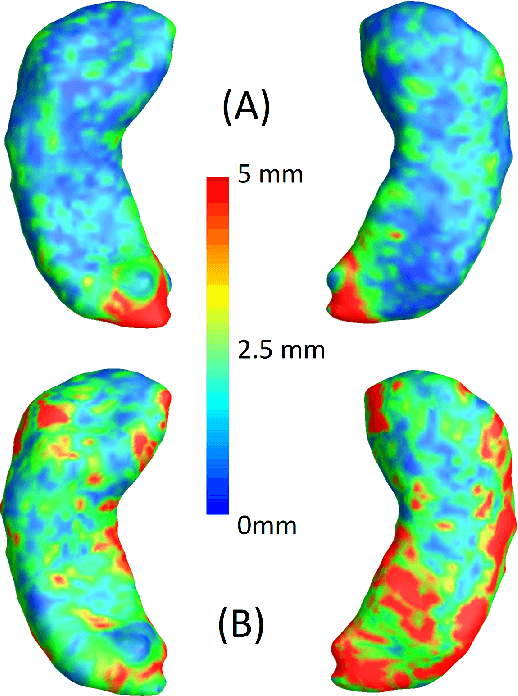
Abstract:We present a method for metric optimization in the Large Deformation Diffeomorphic Metric Mapping (LDDMM) framework, by treating the induced Riemannian metric on the space of diffeomorphisms as a kernel in a machine learning context. For simplicity, we choose the kernel Fischer Linear Discriminant Analysis (KLDA) as the framework. Optimizing the kernel parameters in an Expectation-Maximization framework, we define model fidelity via the hinge loss of the decision function. The resulting algorithm optimizes the parameters of the LDDMM norm-inducing differential operator as a solution to a group-wise registration and classification problem. In practice, this may lead to a biology-aware registration, focusing its attention on the predictive task at hand such as identifying the effects of disease. We first tested our algorithm on a synthetic dataset, showing that our parameter selection improves registration quality and classification accuracy. We then tested the algorithm on 3D subcortical shapes from the Schizophrenia cohort Schizconnect. Our Schizpohrenia-Control predictive model showed significant improvement in ROC AUC compared to baseline parameters.
Connectivity-Driven Brain Parcellation via Consensus Clustering
Aug 10, 2018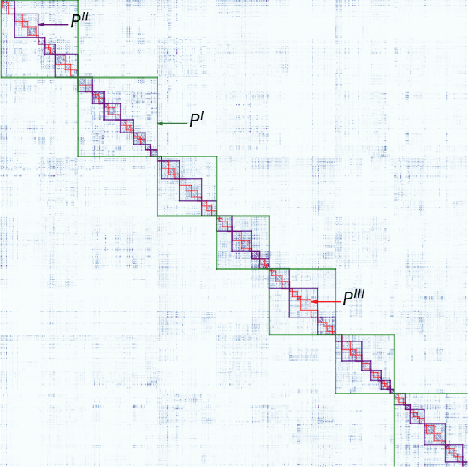
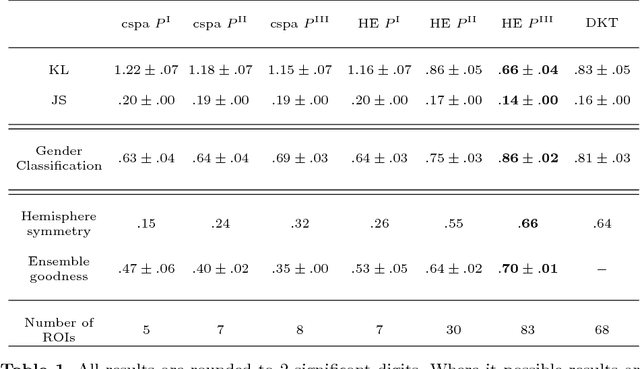
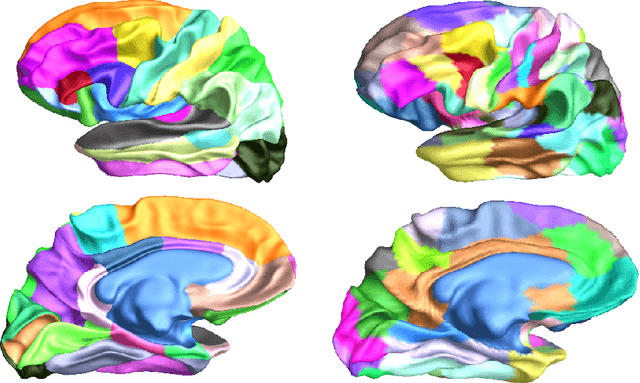
Abstract:We present two related methods for deriving connectivity-based brain atlases from individual connectomes. The proposed methods exploit a previously proposed dense connectivity representation, termed continuous connectivity, by first performing graph-based hierarchical clustering of individual brains, and subsequently aggregating the individual parcellations into a consensus parcellation. The search for consensus minimizes the sum of cluster membership distances, effectively estimating a pseudo-Karcher mean of individual parcellations. We assess the quality of our parcellations using (1) Kullback-Liebler and Jensen-Shannon divergence with respect to the dense connectome representation, (2) inter-hemispheric symmetry, and (3) performance of the simplified connectome in a biological sex classification task. We find that the parcellation based-atlas computed using a greedy search at a hierarchical depth 3 outperforms all other parcellation-based atlases as well as the standard Dessikan-Killiany anatomical atlas in all three assessments.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge