Antonio Foncubierta-Rodriguez
MaxCorrMGNN: A Multi-Graph Neural Network Framework for Generalized Multimodal Fusion of Medical Data for Outcome Prediction
Jul 13, 2023
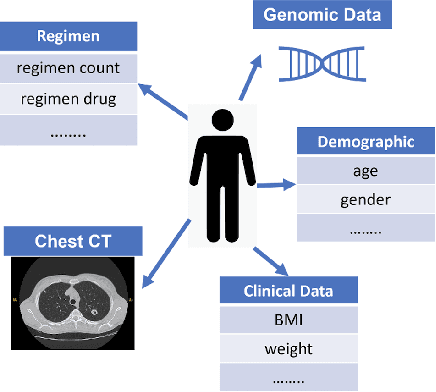
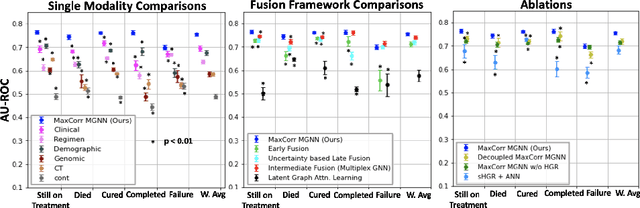
Abstract:With the emergence of multimodal electronic health records, the evidence for an outcome may be captured across multiple modalities ranging from clinical to imaging and genomic data. Predicting outcomes effectively requires fusion frameworks capable of modeling fine-grained and multi-faceted complex interactions between modality features within and across patients. We develop an innovative fusion approach called MaxCorr MGNN that models non-linear modality correlations within and across patients through Hirschfeld-Gebelein-Renyi maximal correlation (MaxCorr) embeddings, resulting in a multi-layered graph that preserves the identities of the modalities and patients. We then design, for the first time, a generalized multi-layered graph neural network (MGNN) for task-informed reasoning in multi-layered graphs, that learns the parameters defining patient-modality graph connectivity and message passing in an end-to-end fashion. We evaluate our model an outcome prediction task on a Tuberculosis (TB) dataset consistently outperforming several state-of-the-art neural, graph-based and traditional fusion techniques.
Fusing Modalities by Multiplexed Graph Neural Networks for Outcome Prediction in Tuberculosis
Oct 25, 2022Abstract:In a complex disease such as tuberculosis, the evidence for the disease and its evolution may be present in multiple modalities such as clinical, genomic, or imaging data. Effective patient-tailored outcome prediction and therapeutic guidance will require fusing evidence from these modalities. Such multimodal fusion is difficult since the evidence for the disease may not be uniform across all modalities, not all modality features may be relevant, or not all modalities may be present for all patients. All these nuances make simple methods of early, late, or intermediate fusion of features inadequate for outcome prediction. In this paper, we present a novel fusion framework using multiplexed graphs and derive a new graph neural network for learning from such graphs. Specifically, the framework allows modalities to be represented through their targeted encodings, and models their relationship explicitly via multiplexed graphs derived from salient features in a combined latent space. We present results that show that our proposed method outperforms state-of-the-art methods of fusing modalities for multi-outcome prediction on a large Tuberculosis (TB) dataset.
Learning-based Defect Recognition for Quasi-Periodic Microscope Images
Jul 02, 2020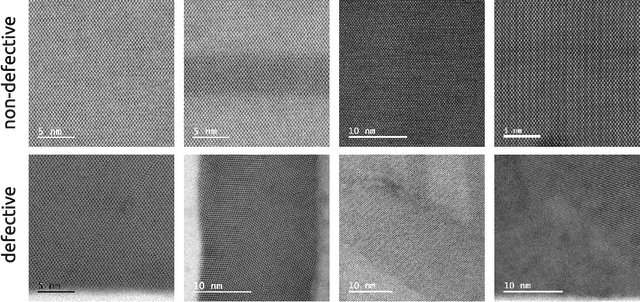

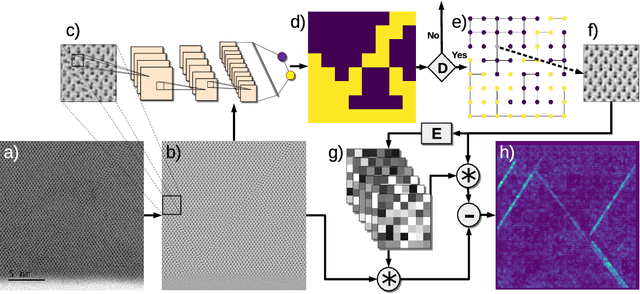
Abstract:The detailed control of crystalline material defects is a crucial process, as they affect properties of the material that may be detrimental or beneficial for the final performance of a device. Defect analysis on the sub-nanometer scale is enabled by high-resolution transmission electron microscopy (HRTEM), where the identification of defects is currently carried out based on human expertise. However, the process is tedious, highly time consuming and, in some cases, can yield to ambiguous results. Here we propose a semi-supervised machine learning method that assists in the detection of lattice defects from atomic resolution microscope images. It involves a convolutional neural network that classifies image patches as defective or non-defective, a graph-based heuristic that chooses one non-defective patch as a model, and finally an automatically generated convolutional filter bank, which highlights symmetry breaking such as stacking faults, twin defects and grain boundaries. Additionally, a variance filter is suggested to segment amorphous regions and beam defects. The algorithm is tested on III-V/Si crystalline materials and successfully evaluated against different metrics, showing promising results even for extremely small data sets. By combining the data-driven classification generality, robustness and speed of deep learning with the effectiveness of image filters in segmenting faulty symmetry arrangements, we provide a valuable open-source tool to the microscopist community that can streamline future HRTEM analyses of crystalline materials.
Mitosis Detection Under Limited Annotation: A Joint Learning Approach
Jul 02, 2020



Abstract:Mitotic counting is a vital prognostic marker of tumor proliferation in breast cancer. Deep learning-based mitotic detection is on par with pathologists, but it requires large labeled data for training. We propose a deep classification framework for enhancing mitosis detection by leveraging class label information, via softmax loss, and spatial distribution information among samples, via distance metric learning. We also investigate strategies towards steadily providing informative samples to boost the learning. The efficacy of the proposed framework is established through evaluation on ICPR 2012 and AMIDA 2013 mitotic data. Our framework significantly improves the detection with small training data and achieves on par or superior performance compared to state-of-the-art methods for using the entire training data.
Towards Explainable Graph Representations in Digital Pathology
Jul 01, 2020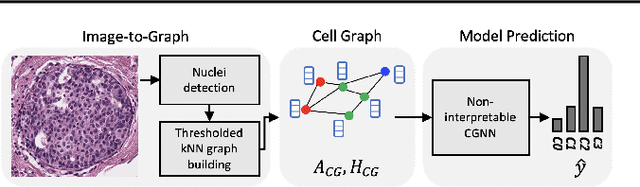
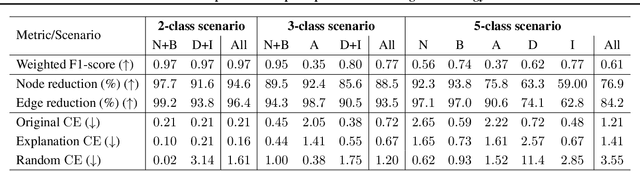
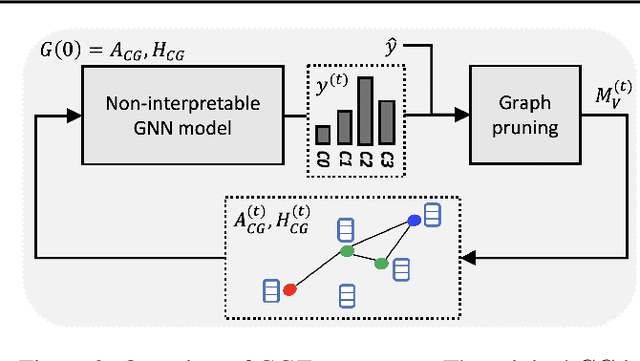
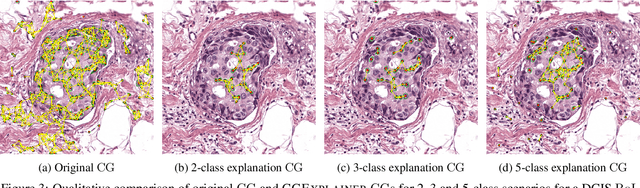
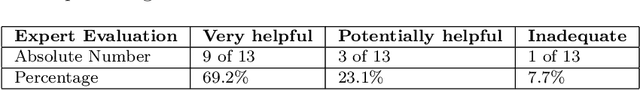
Abstract:Explainability of machine learning (ML) techniques in digital pathology (DP) is of great significance to facilitate their wide adoption in clinics. Recently, graph techniques encoding relevant biological entities have been employed to represent and assess DP images. Such paradigm shift from pixel-wise to entity-wise analysis provides more control over concept representation. In this paper, we introduce a post-hoc explainer to derive compact per-instance explanations emphasizing diagnostically important entities in the graph. Although we focus our analyses to cells and cellular interactions in breast cancer subtyping, the proposed explainer is generic enough to be extended to other topological representations in DP. Qualitative and quantitative analyses demonstrate the efficacy of the explainer in generating comprehensive and compact explanations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge