Alicia Durrer
Towards MR-Based Trochleoplasty Planning
Aug 08, 2025Abstract:To treat Trochlear Dysplasia (TD), current approaches rely mainly on low-resolution clinical Magnetic Resonance (MR) scans and surgical intuition. The surgeries are planned based on surgeons experience, have limited adoption of minimally invasive techniques, and lead to inconsistent outcomes. We propose a pipeline that generates super-resolved, patient-specific 3D pseudo-healthy target morphologies from conventional clinical MR scans. First, we compute an isotropic super-resolved MR volume using an Implicit Neural Representation (INR). Next, we segment femur, tibia, patella, and fibula with a multi-label custom-trained network. Finally, we train a Wavelet Diffusion Model (WDM) to generate pseudo-healthy target morphologies of the trochlear region. In contrast to prior work producing pseudo-healthy low-resolution 3D MR images, our approach enables the generation of sub-millimeter resolved 3D shapes compatible for pre- and intraoperative use. These can serve as preoperative blueprints for reshaping the femoral groove while preserving the native patella articulation. Furthermore, and in contrast to other work, we do not require a CT for our pipeline - reducing the amount of radiation. We evaluated our approach on 25 TD patients and could show that our target morphologies significantly improve the sulcus angle (SA) and trochlear groove depth (TGD). The code and interactive visualization are available at https://wehrlimi.github.io/sr-3d-planning/.
fastWDM3D: Fast and Accurate 3D Healthy Tissue Inpainting
Jul 17, 2025


Abstract:Healthy tissue inpainting has significant applications, including the generation of pseudo-healthy baselines for tumor growth models and the facilitation of image registration. In previous editions of the BraTS Local Synthesis of Healthy Brain Tissue via Inpainting Challenge, denoising diffusion probabilistic models (DDPMs) demonstrated qualitatively convincing results but suffered from low sampling speed. To mitigate this limitation, we adapted a 2D image generation approach, combining DDPMs with generative adversarial networks (GANs) and employing a variance-preserving noise schedule, for the task of 3D inpainting. Our experiments showed that the variance-preserving noise schedule and the selected reconstruction losses can be effectively utilized for high-quality 3D inpainting in a few time steps without requiring adversarial training. We applied our findings to a different architecture, a 3D wavelet diffusion model (WDM3D) that does not include a GAN component. The resulting model, denoted as fastWDM3D, obtained a SSIM of 0.8571, a MSE of 0.0079, and a PSNR of 22.26 on the BraTS inpainting test set. Remarkably, it achieved these scores using only two time steps, completing the 3D inpainting process in 1.81 s per image. When compared to other DDPMs used for healthy brain tissue inpainting, our model is up to 800 x faster while still achieving superior performance metrics. Our proposed method, fastWDM3D, represents a promising approach for fast and accurate healthy tissue inpainting. Our code is available at https://github.com/AliciaDurrer/fastWDM3D.
Generating 3D Pseudo-Healthy Knee MR Images to Support Trochleoplasty Planning
Dec 13, 2024

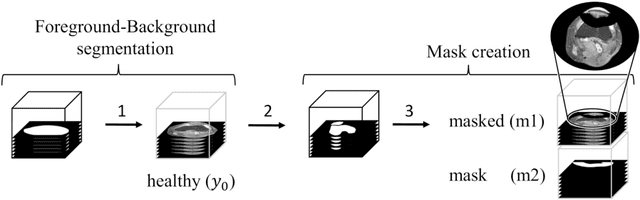
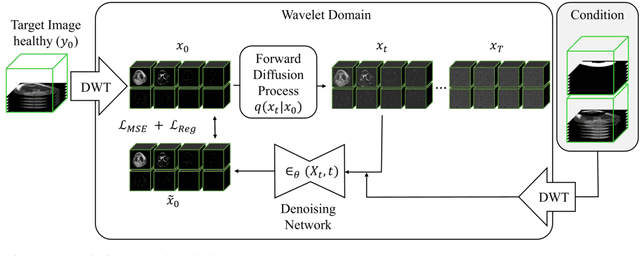
Abstract:Purpose: Trochlear Dysplasia (TD) is a common malformation in adolescents, leading to anterior knee pain and instability. Surgical interventions such as trochleoplasty require precise planning to correct the trochlear groove. However, no standardized preoperative plan exists to guide surgeons in reshaping the femur. This study aims to generate patient-specific, pseudo-healthy MR images of the trochlear region that should theoretically align with the respective patient's patella, potentially supporting the pre-operative planning of trochleoplasty. Methods: We employ a Wavelet Diffusion Model (WDM) to generate personalized pseudo-healthy, anatomically plausible MR scans of the trochlear region. We train our model using knee MR scans of healthy subjects. During inference, we mask out pathological regions around the patella in scans of patients affected by TD, and replace them with their pseudo-healthy counterpart. An orthopedic surgeon measured the sulcus angle (SA), trochlear groove depth (TGD) and D\'ejour classification in MR scans before and after inpainting. The code is available at https://github.com/wehrlimi/Generate-Pseudo-Healthy-Knee-MRI . Results: The inpainting by our model significantly improves the SA, TGD and D\'ejour classification in a study with 49 knee MR scans. Conclusion: This study demonstrates the potential of WDMs in providing surgeons with patient-specific guidance. By offering anatomically plausible MR scans, the method could potentially enhance the precision and preoperative planning of trochleoplasty, and pave the way to more minimally invasive surgeries.
cWDM: Conditional Wavelet Diffusion Models for Cross-Modality 3D Medical Image Synthesis
Nov 26, 2024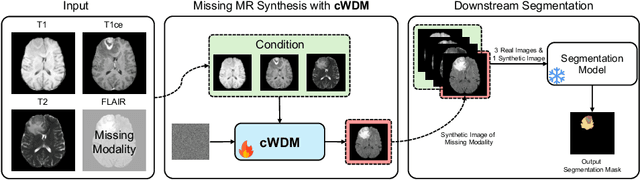
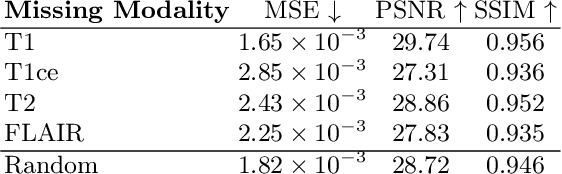

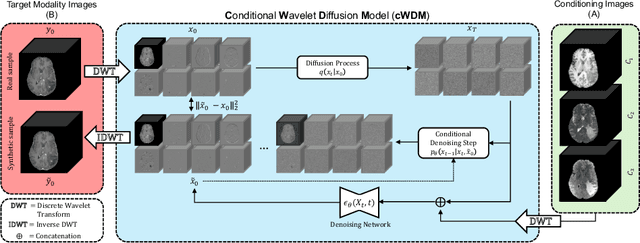

Abstract:This paper contributes to the "BraTS 2024 Brain MR Image Synthesis Challenge" and presents a conditional Wavelet Diffusion Model (cWDM) for directly solving a paired image-to-image translation task on high-resolution volumes. While deep learning-based brain tumor segmentation models have demonstrated clear clinical utility, they typically require MR scans from various modalities (T1, T1ce, T2, FLAIR) as input. However, due to time constraints or imaging artifacts, some of these modalities may be missing, hindering the application of well-performing segmentation algorithms in clinical routine. To address this issue, we propose a method that synthesizes one missing modality image conditioned on three available images, enabling the application of downstream segmentation models. We treat this paired image-to-image translation task as a conditional generation problem and solve it by combining a Wavelet Diffusion Model for high-resolution 3D image synthesis with a simple conditioning strategy. This approach allows us to directly apply our model to full-resolution volumes, avoiding artifacts caused by slice- or patch-wise data processing. While this work focuses on a specific application, the presented method can be applied to all kinds of paired image-to-image translation problems, such as CT $\leftrightarrow$ MR and MR $\leftrightarrow$ PET translation, or mask-conditioned anatomically guided image generation.
Modeling the Neonatal Brain Development Using Implicit Neural Representations
Aug 16, 2024Abstract:The human brain undergoes rapid development during the third trimester of pregnancy. In this work, we model the neonatal development of the infant brain in this age range. As a basis, we use MR images of preterm- and term-birth neonates from the developing human connectome project (dHCP). We propose a neural network, specifically an implicit neural representation (INR), to predict 2D- and 3D images of varying time points. In order to model a subject-specific development process, it is necessary to disentangle the age from the subjects' identity in the latent space of the INR. We propose two methods, Subject Specific Latent Vectors (SSL) and Stochastic Global Latent Augmentation (SGLA), enabling this disentanglement. We perform an analysis of the results and compare our proposed model to an age-conditioned denoising diffusion model as a baseline. We also show that our method can be applied in a memory-efficient way, which is especially important for 3D data.
Denoising Diffusion Models for 3D Healthy Brain Tissue Inpainting
Mar 21, 2024



Abstract:Monitoring diseases that affect the brain's structural integrity requires automated analysis of magnetic resonance (MR) images, e.g., for the evaluation of volumetric changes. However, many of the evaluation tools are optimized for analyzing healthy tissue. To enable the evaluation of scans containing pathological tissue, it is therefore required to restore healthy tissue in the pathological areas. In this work, we explore and extend denoising diffusion models for consistent inpainting of healthy 3D brain tissue. We modify state-of-the-art 2D, pseudo-3D, and 3D methods working in the image space, as well as 3D latent and 3D wavelet diffusion models, and train them to synthesize healthy brain tissue. Our evaluation shows that the pseudo-3D model performs best regarding the structural-similarity index, peak signal-to-noise ratio, and mean squared error. To emphasize the clinical relevance, we fine-tune this model on data containing synthetic MS lesions and evaluate it on a downstream brain tissue segmentation task, whereby it outperforms the established FMRIB Software Library (FSL) lesion-filling method.
Binary Noise for Binary Tasks: Masked Bernoulli Diffusion for Unsupervised Anomaly Detection
Mar 18, 2024

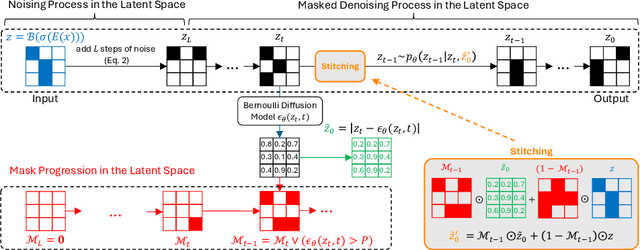

Abstract:The high performance of denoising diffusion models for image generation has paved the way for their application in unsupervised medical anomaly detection. As diffusion-based methods require a lot of GPU memory and have long sampling times, we present a novel and fast unsupervised anomaly detection approach based on latent Bernoulli diffusion models. We first apply an autoencoder to compress the input images into a binary latent representation. Next, a diffusion model that follows a Bernoulli noise schedule is employed to this latent space and trained to restore binary latent representations from perturbed ones. The binary nature of this diffusion model allows us to identify entries in the latent space that have a high probability of flipping their binary code during the denoising process, which indicates out-of-distribution data. We propose a masking algorithm based on these probabilities, which improves the anomaly detection scores. We achieve state-of-the-art performance compared to other diffusion-based unsupervised anomaly detection algorithms while significantly reducing sampling time and memory consumption. The code is available at https://github.com/JuliaWolleb/Anomaly_berdiff.
WDM: 3D Wavelet Diffusion Models for High-Resolution Medical Image Synthesis
Feb 29, 2024Abstract:Due to the three-dimensional nature of CT- or MR-scans, generative modeling of medical images is a particularly challenging task. Existing approaches mostly apply patch-wise, slice-wise, or cascaded generation techniques to fit the high-dimensional data into the limited GPU memory. However, these approaches may introduce artifacts and potentially restrict the model's applicability for certain downstream tasks. This work presents WDM, a wavelet-based medical image synthesis framework that applies a diffusion model on wavelet decomposed images. The presented approach is a simple yet effective way of scaling diffusion models to high resolutions and can be trained on a single 40 GB GPU. Experimental results on BraTS and LIDC-IDRI unconditional image generation at a resolution of $128 \times 128 \times 128$ show state-of-the-art image fidelity (FID) and sample diversity (MS-SSIM) scores compared to GANs, Diffusion Models, and Latent Diffusion Models. Our proposed method is the only one capable of generating high-quality images at a resolution of $256 \times 256 \times 256$.
Denoising Diffusion Models for Inpainting of Healthy Brain Tissue
Feb 27, 2024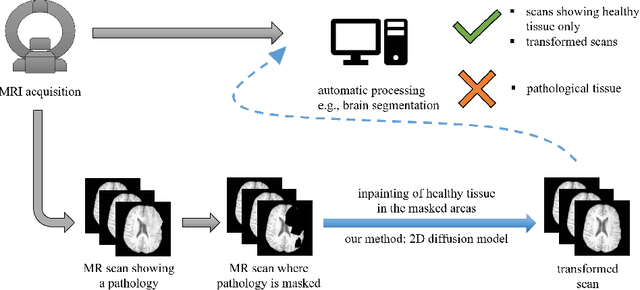
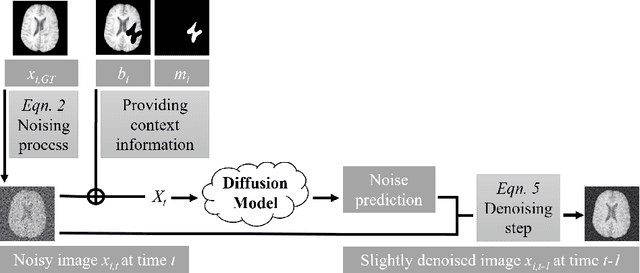
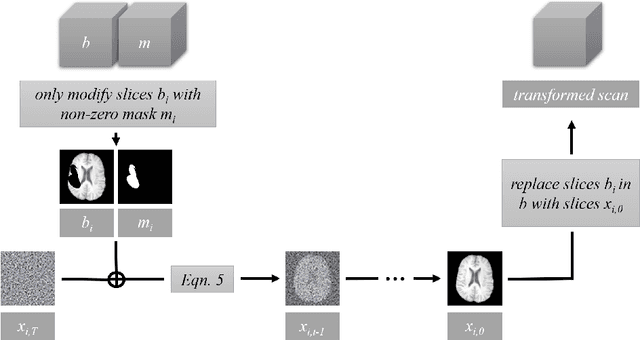
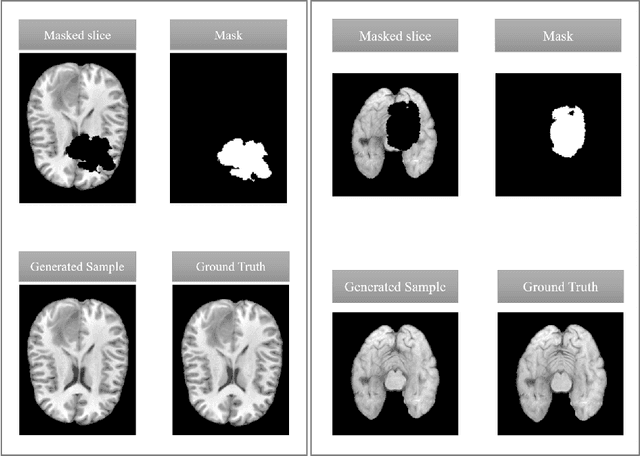
Abstract:This paper is a contribution to the "BraTS 2023 Local Synthesis of Healthy Brain Tissue via Inpainting Challenge". The task of this challenge is to transform tumor tissue into healthy tissue in brain magnetic resonance (MR) images. This idea originates from the problem that MR images can be evaluated using automatic processing tools, however, many of these tools are optimized for the analysis of healthy tissue. By solving the given inpainting task, we enable the automatic analysis of images featuring lesions, and further downstream tasks. Our approach builds on denoising diffusion probabilistic models. We use a 2D model that is trained using slices in which healthy tissue was cropped out and is learned to be inpainted again. This allows us to use the ground truth healthy tissue during training. In the sampling stage, we replace the slices containing diseased tissue in the original 3D volume with the slices containing the healthy tissue inpainting. With our approach, we achieve comparable results to the competing methods. On the validation set our model achieves a mean SSIM of 0.7804, a PSNR of 20.3525 and a MSE of 0.0113. In future we plan to extend our 2D model to a 3D model, allowing to inpaint the region of interest as a whole without losing context information of neighboring slices.
Diffusion Models for Memory-efficient Processing of 3D Medical Images
Mar 27, 2023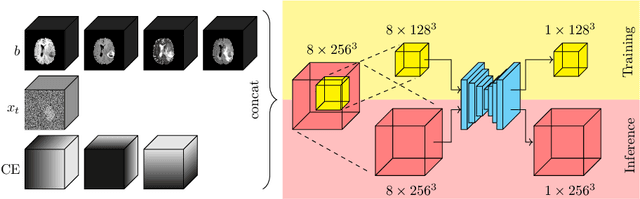

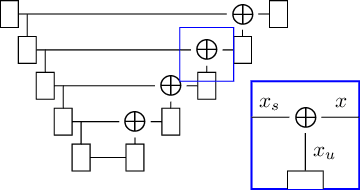
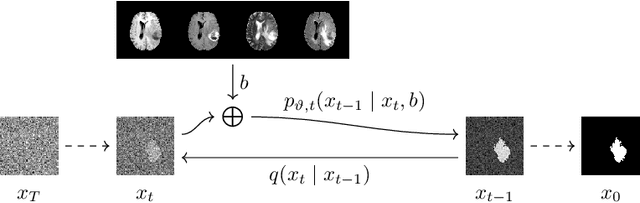
Abstract:Denoising diffusion models have recently achieved state-of-the-art performance in many image-generation tasks. They do, however, require a large amount of computational resources. This limits their application to medical tasks, where we often deal with large 3D volumes, like high-resolution three-dimensional data. In this work, we present a number of different ways to reduce the resource consumption for 3D diffusion models and apply them to a dataset of 3D images. The main contribution of this paper is the memory-efficient patch-based diffusion model \textit{PatchDDM}, which can be applied to the total volume during inference while the training is performed only on patches. While the proposed diffusion model can be applied to any image generation tasks, we evaluate the method on the tumor segmentation task of the BraTS2020 dataset and demonstrate that we can generate meaningful three-dimensional segmentations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge