Alexandra Dalechina
Neglectable effect of brain MRI data prepreprocessing for tumor segmentation
Apr 11, 2022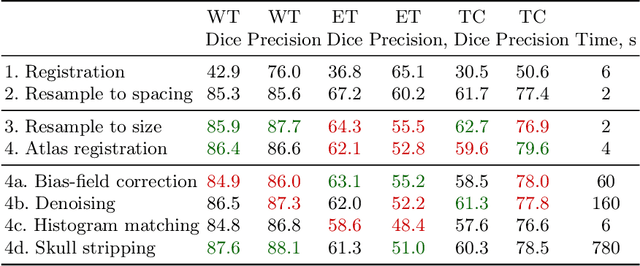
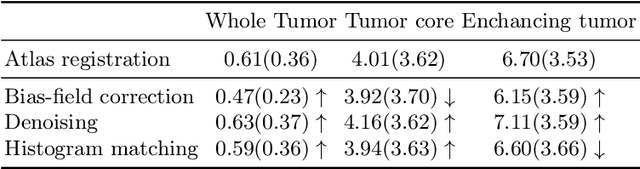
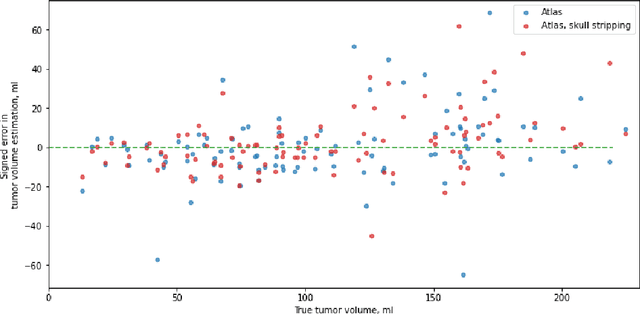

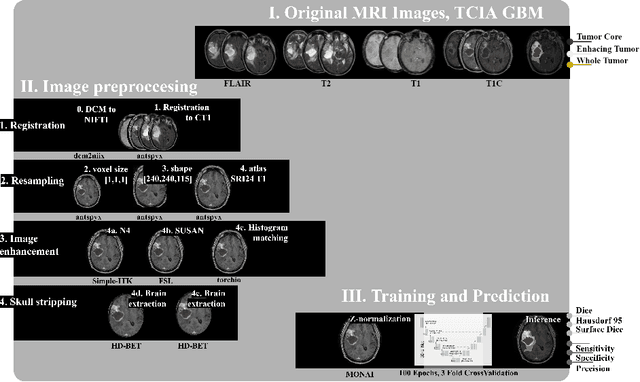
Abstract:Magnetic resonance imaging (MRI) data is heterogeneous due to the differences in device manufacturers, scanning protocols, and inter-subject variability. A conventional way to mitigate MR image heterogeneity is to apply preprocessing transformations, such as anatomy alignment, voxel resampling, signal intensity equalization, image denoising, and localization of regions of interest (ROI). Although preprocessing pipeline standardizes image appearance, its influence on the quality of image segmentation and other downstream tasks on deep neural networks (DNN) has never been rigorously studied. Here we report a comprehensive study of multimodal MRI brain cancer image segmentation on TCIA-GBM open-source dataset. Our results demonstrate that most popular standardization steps add no value to artificial neural network performance; moreover, preprocessing can hamper model performance. We suggest that image intensity normalization approaches do not contribute to model accuracy because of the reduction of signal variance with image standardization. Finally, we show the contribution of scull-stripping in data preprocessing is almost negligible if measured in terms of clinically relevant metrics. We show that the only essential transformation for accurate analysis is the unification of voxel spacing across the dataset. In contrast, anatomy alignment in form of non-rigid atlas registration is not necessary and most intensity equalization steps do not improve model productiveness.
Systematic Clinical Evaluation of A Deep Learning Method for Medical Image Segmentation: Radiosurgery Application
Aug 21, 2021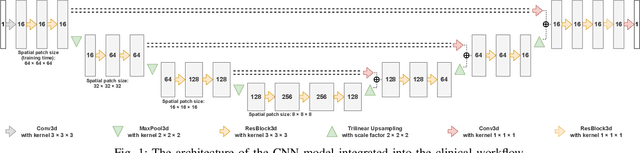
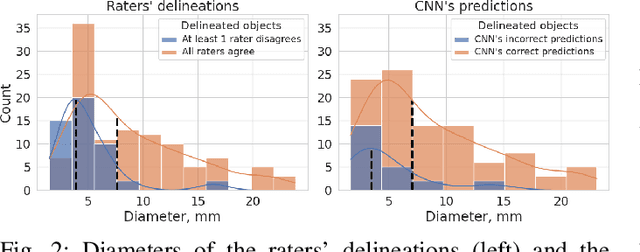
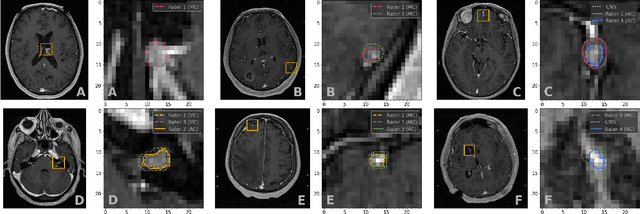
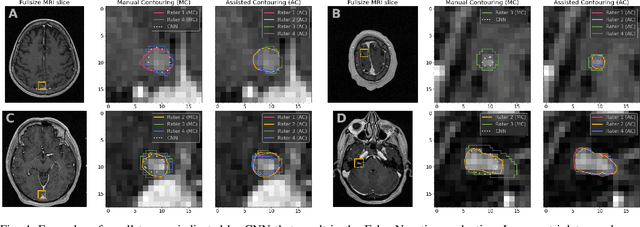
Abstract:We systematically evaluate a Deep Learning (DL) method in a 3D medical image segmentation task. Our segmentation method is integrated into the radiosurgery treatment process and directly impacts the clinical workflow. With our method, we address the relative drawbacks of manual segmentation: high inter-rater contouring variability and high time consumption of the contouring process. The main extension over the existing evaluations is the careful and detailed analysis that could be further generalized on other medical image segmentation tasks. Firstly, we analyze the changes in the inter-rater detection agreement. We show that the segmentation model reduces the ratio of detection disagreements from 0.162 to 0.085 (p < 0.05). Secondly, we show that the model improves the inter-rater contouring agreement from 0.845 to 0.871 surface Dice Score (p < 0.05). Thirdly, we show that the model accelerates the delineation process in between 1.6 and 2.0 times (p < 0.05). Finally, we design the setup of the clinical experiment to either exclude or estimate the evaluation biases, thus preserve the significance of the results. Besides the clinical evaluation, we also summarize the intuitions and practical ideas for building an efficient DL-based model for 3D medical image segmentation.
Universal Loss Reweighting to Balance Lesion Size Inequality in 3D Medical Image Segmentation
Jul 20, 2020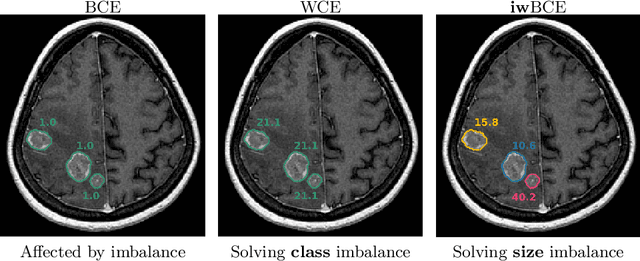
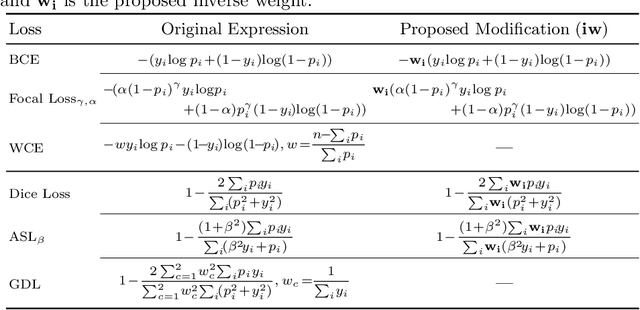

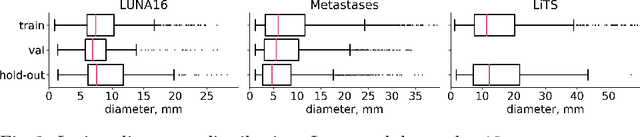
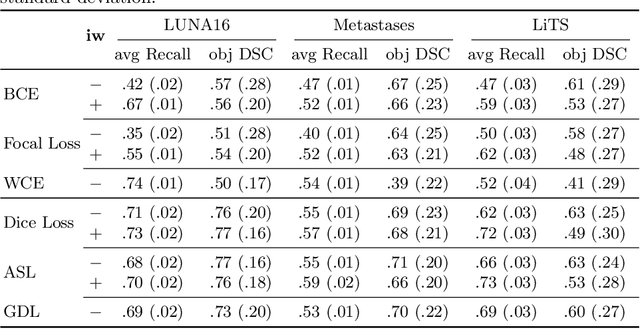
Abstract:Target imbalance affects the performance of recent deep learning methods in many medical image segmentation tasks. It is a twofold problem: class imbalance - positive class (lesion) size compared to negative class (non-lesion) size; lesion size imbalance - large lesions overshadows small ones (in the case of multiple lesions per image). While the former was addressed in multiple works, the latter lacks investigation. We propose a loss reweighting approach to increase the ability of the network to detect small lesions. During the learning process, we assign a weight to every image voxel. The assigned weights are inversely proportional to the lesion volume, thus smaller lesions get larger weights. We report the benefit from our method for well-known loss functions, including Dice Loss, Focal Loss, and Asymmetric Similarity Loss. Additionally, we compare our results with other reweighting techniques: Weighted Cross-Entropy and Generalized Dice Loss. Our experiments show that inverse weighting considerably increases the detection quality, while preserves the delineation quality on a state-of-the-art level. We publish a complete experimental pipeline for two publicly available datasets of CT images: LiTS and LUNA16 (https://github.com/neuro-ml/inverse_weighting). We also show results on a private database of MR images for the task of multiple brain metastases delineation.
Multi-domain CT metal artifacts reduction using partial convolution based inpainting
Nov 13, 2019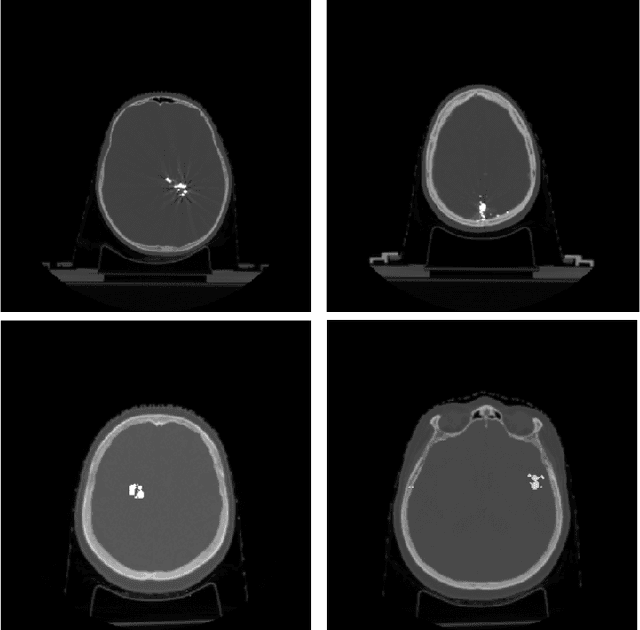
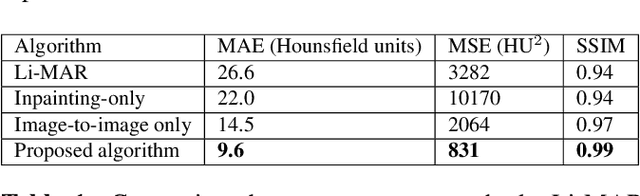

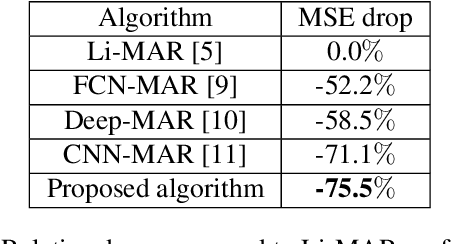
Abstract:Recent CT Metal Artifacts Reduction (MAR) methods are often based on image-to-image convolutional neural networks for adjustment of corrupted sinograms or images themselves. In this paper, we are exploring the capabilities of a multi-domain method which consists of both sinogram correction (projection domain step) and restored image correction (image-domain step). Moreover, we propose a formulation of the first step problem as sinogram inpainting which allows us to use methods of this specific field such as partial convolutions. The proposed method allows to achieve state-of-the-art (-75% MSE) improvement in comparison with a classic benchmark - Li-MAR.
Deep Learning for Brain Tumor Segmentation in Radiosurgery: Prospective Clinical Evaluation
Sep 06, 2019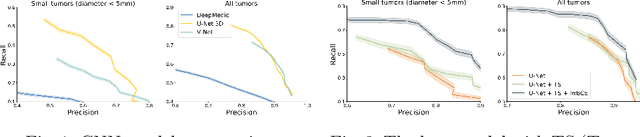
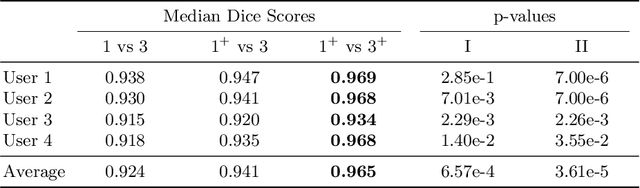
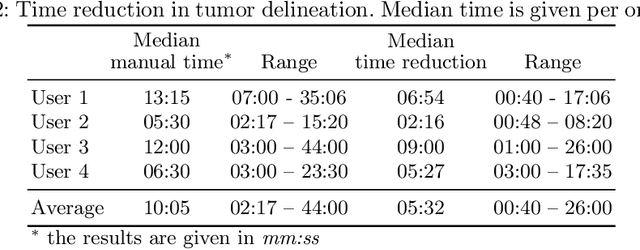
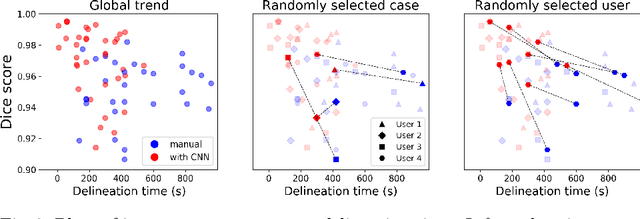
Abstract:Stereotactic radiosurgery is a minimally-invasive treatment option for a large number of patients with intracranial tumors. As part of the therapy treatment, accurate delineation of brain tumors is of great importance. However, slice-by-slice manual segmentation on T1c MRI could be time-consuming (especially for multiple metastases) and subjective (especially for meningiomas). In our work, we compared several deep convolutional networks architectures and training procedures and evaluated the best model in a radiation therapy department for three types of brain tumors: meningiomas, schwannomas and multiple brain metastases. The developed semiautomatic segmentation system accelerates the contouring process by 2.2 times on average and increases inter-rater agreement from 92% to 96.5%.
Brain Tumor Image Retrieval via Multitask Learning
Oct 22, 2018
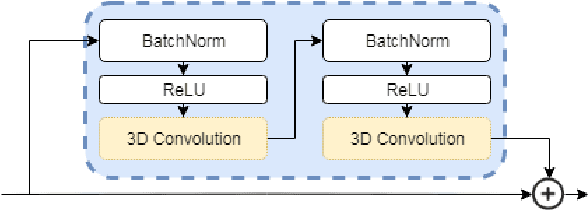
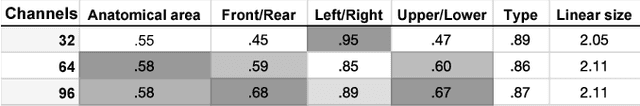
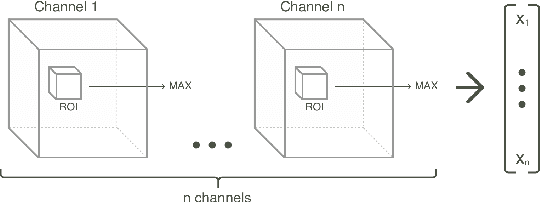
Abstract:Classification-based image retrieval systems are built by training convolutional neural networks (CNNs) on a relevant classification problem and using the distance in the resulting feature space as a similarity metric. However, in practical applications, it is often desirable to have representations which take into account several aspects of the data (e.g., brain tumor type and its localization). In our work, we extend the classification-based approach with multitask learning: we train a CNN on brain MRI scans with heterogeneous labels and implement a corresponding tumor image retrieval system. We validate our approach on brain tumor data which contains information about tumor types, shapes and localization. We show that our method allows us to build representations that contain more relevant information about tumors than single-task classification-based approaches.
Tumor Delineation For Brain Radiosurgery by a ConvNet and Non-Uniform Patch Generation
Aug 01, 2018

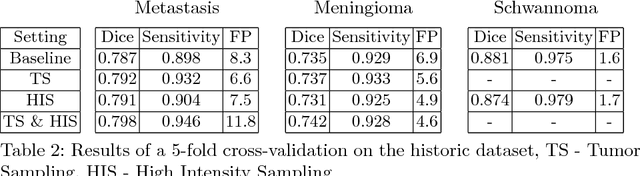

Abstract:Deep learning methods are actively used for brain lesion segmentation. One of the most popular models is DeepMedic, which was developed for segmentation of relatively large lesions like glioma and ischemic stroke. In our work, we consider segmentation of brain tumors appropriate to stereotactic radiosurgery which limits typical lesion sizes. These differences in target volumes lead to a large number of false negatives (especially for small lesions) as well as to an increased number of false positives for DeepMedic. We propose a new patch-sampling procedure to increase network performance for small lesions. We used a 6-year dataset from a stereotactic radiosurgery center. To evaluate our approach, we conducted experiments with the three most frequent brain tumors: metastasis, meningioma, schwannoma. In addition to cross-validation, we estimated quality on a hold-out test set which was collected several years later than the train one. The experimental results show solid improvements in both cases.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge