Adrien Bazoge
MediQAl: A French Medical Question Answering Dataset for Knowledge and Reasoning Evaluation
Jul 28, 2025Abstract:This work introduces MediQAl, a French medical question answering dataset designed to evaluate the capabilities of language models in factual medical recall and reasoning over real-world clinical scenarios. MediQAl contains 32,603 questions sourced from French medical examinations across 41 medical subjects. The dataset includes three tasks: (i) Multiple-Choice Question with Unique answer, (ii) Multiple-Choice Question with Multiple answer, and (iii) Open-Ended Question with Short-Answer. Each question is labeled as Understanding or Reasoning, enabling a detailed analysis of models' cognitive capabilities. We validate the MediQAl dataset through extensive evaluation with 14 large language models, including recent reasoning-augmented models, and observe a significant performance gap between factual recall and reasoning tasks. Our evaluation provides a comprehensive benchmark for assessing language models' performance on French medical question answering, addressing a crucial gap in multilingual resources for the medical domain.
Adaptation of Biomedical and Clinical Pretrained Models to French Long Documents: A Comparative Study
Feb 26, 2024



Abstract:Recently, pretrained language models based on BERT have been introduced for the French biomedical domain. Although these models have achieved state-of-the-art results on biomedical and clinical NLP tasks, they are constrained by a limited input sequence length of 512 tokens, which poses challenges when applied to clinical notes. In this paper, we present a comparative study of three adaptation strategies for long-sequence models, leveraging the Longformer architecture. We conducted evaluations of these models on 16 downstream tasks spanning both biomedical and clinical domains. Our findings reveal that further pre-training an English clinical model with French biomedical texts can outperform both converting a French biomedical BERT to the Longformer architecture and pre-training a French biomedical Longformer from scratch. The results underscore that long-sequence French biomedical models improve performance across most downstream tasks regardless of sequence length, but BERT based models remain the most efficient for named entity recognition tasks.
How Important Is Tokenization in French Medical Masked Language Models?
Feb 22, 2024Abstract:Subword tokenization has become the prevailing standard in the field of natural language processing (NLP) over recent years, primarily due to the widespread utilization of pre-trained language models. This shift began with Byte-Pair Encoding (BPE) and was later followed by the adoption of SentencePiece and WordPiece. While subword tokenization consistently outperforms character and word-level tokenization, the precise factors contributing to its success remain unclear. Key aspects such as the optimal segmentation granularity for diverse tasks and languages, the influence of data sources on tokenizers, and the role of morphological information in Indo-European languages remain insufficiently explored. This is particularly pertinent for biomedical terminology, characterized by specific rules governing morpheme combinations. Despite the agglutinative nature of biomedical terminology, existing language models do not explicitly incorporate this knowledge, leading to inconsistent tokenization strategies for common terms. In this paper, we seek to delve into the complexities of subword tokenization in French biomedical domain across a variety of NLP tasks and pinpoint areas where further enhancements can be made. We analyze classical tokenization algorithms, including BPE and SentencePiece, and introduce an original tokenization strategy that integrates morpheme-enriched word segmentation into existing tokenization methods.
DrBenchmark: A Large Language Understanding Evaluation Benchmark for French Biomedical Domain
Feb 20, 2024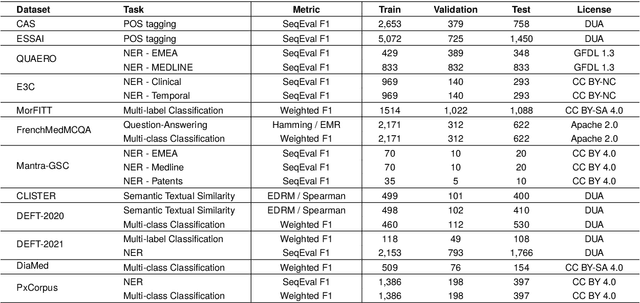
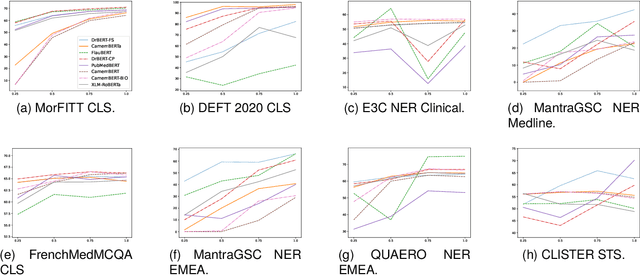
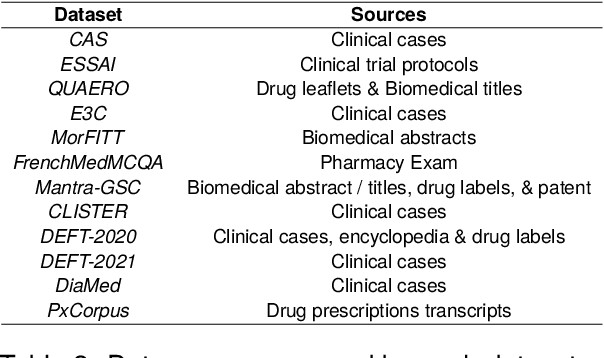
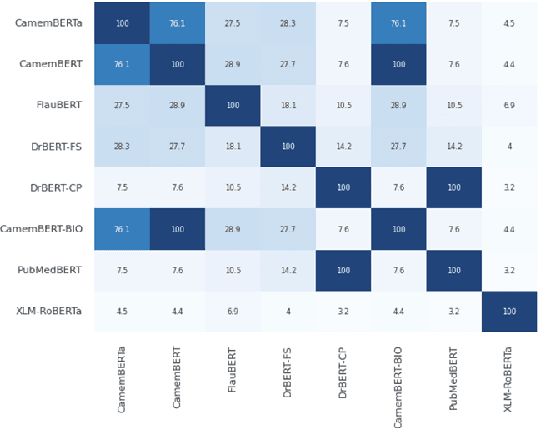
Abstract:The biomedical domain has sparked a significant interest in the field of Natural Language Processing (NLP), which has seen substantial advancements with pre-trained language models (PLMs). However, comparing these models has proven challenging due to variations in evaluation protocols across different models. A fair solution is to aggregate diverse downstream tasks into a benchmark, allowing for the assessment of intrinsic PLMs qualities from various perspectives. Although still limited to few languages, this initiative has been undertaken in the biomedical field, notably English and Chinese. This limitation hampers the evaluation of the latest French biomedical models, as they are either assessed on a minimal number of tasks with non-standardized protocols or evaluated using general downstream tasks. To bridge this research gap and account for the unique sensitivities of French, we present the first-ever publicly available French biomedical language understanding benchmark called DrBenchmark. It encompasses 20 diversified tasks, including named-entity recognition, part-of-speech tagging, question-answering, semantic textual similarity, and classification. We evaluate 8 state-of-the-art pre-trained masked language models (MLMs) on general and biomedical-specific data, as well as English specific MLMs to assess their cross-lingual capabilities. Our experiments reveal that no single model excels across all tasks, while generalist models are sometimes still competitive.
BioMistral: A Collection of Open-Source Pretrained Large Language Models for Medical Domains
Feb 15, 2024



Abstract:Large Language Models (LLMs) have demonstrated remarkable versatility in recent years, offering potential applications across specialized domains such as healthcare and medicine. Despite the availability of various open-source LLMs tailored for health contexts, adapting general-purpose LLMs to the medical domain presents significant challenges. In this paper, we introduce BioMistral, an open-source LLM tailored for the biomedical domain, utilizing Mistral as its foundation model and further pre-trained on PubMed Central. We conduct a comprehensive evaluation of BioMistral on a benchmark comprising 10 established medical question-answering (QA) tasks in English. We also explore lightweight models obtained through quantization and model merging approaches. Our results demonstrate BioMistral's superior performance compared to existing open-source medical models and its competitive edge against proprietary counterparts. Finally, to address the limited availability of data beyond English and to assess the multilingual generalization of medical LLMs, we automatically translated and evaluated this benchmark into 7 other languages. This marks the first large-scale multilingual evaluation of LLMs in the medical domain. Datasets, multilingual evaluation benchmarks, scripts, and all the models obtained during our experiments are freely released.
FrenchMedMCQA: A French Multiple-Choice Question Answering Dataset for Medical domain
Apr 09, 2023


Abstract:This paper introduces FrenchMedMCQA, the first publicly available Multiple-Choice Question Answering (MCQA) dataset in French for medical domain. It is composed of 3,105 questions taken from real exams of the French medical specialization diploma in pharmacy, mixing single and multiple answers. Each instance of the dataset contains an identifier, a question, five possible answers and their manual correction(s). We also propose first baseline models to automatically process this MCQA task in order to report on the current performances and to highlight the difficulty of the task. A detailed analysis of the results showed that it is necessary to have representations adapted to the medical domain or to the MCQA task: in our case, English specialized models yielded better results than generic French ones, even though FrenchMedMCQA is in French. Corpus, models and tools are available online.
DrBERT: A Robust Pre-trained Model in French for Biomedical and Clinical domains
Apr 03, 2023Abstract:In recent years, pre-trained language models (PLMs) achieve the best performance on a wide range of natural language processing (NLP) tasks. While the first models were trained on general domain data, specialized ones have emerged to more effectively treat specific domains. In this paper, we propose an original study of PLMs in the medical domain on French language. We compare, for the first time, the performance of PLMs trained on both public data from the web and private data from healthcare establishments. We also evaluate different learning strategies on a set of biomedical tasks. In particular, we show that we can take advantage of already existing biomedical PLMs in a foreign language by further pre-train it on our targeted data. Finally, we release the first specialized PLMs for the biomedical field in French, called DrBERT, as well as the largest corpus of medical data under free license on which these models are trained.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge