The Shaky Foundations of Clinical Foundation Models: A Survey of Large Language Models and Foundation Models for EMRs
Paper and Code
Mar 24, 2023
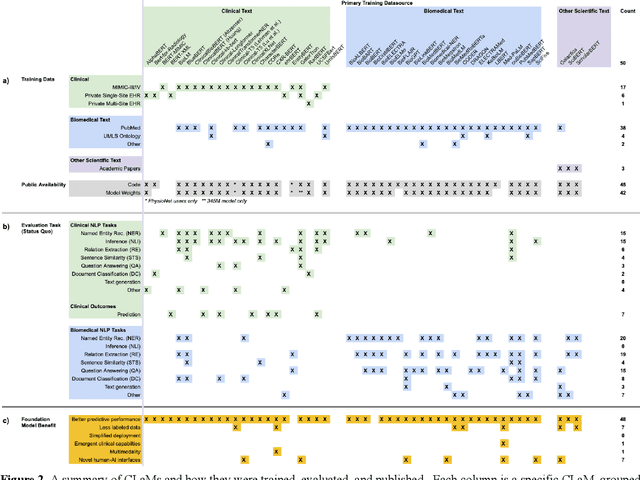
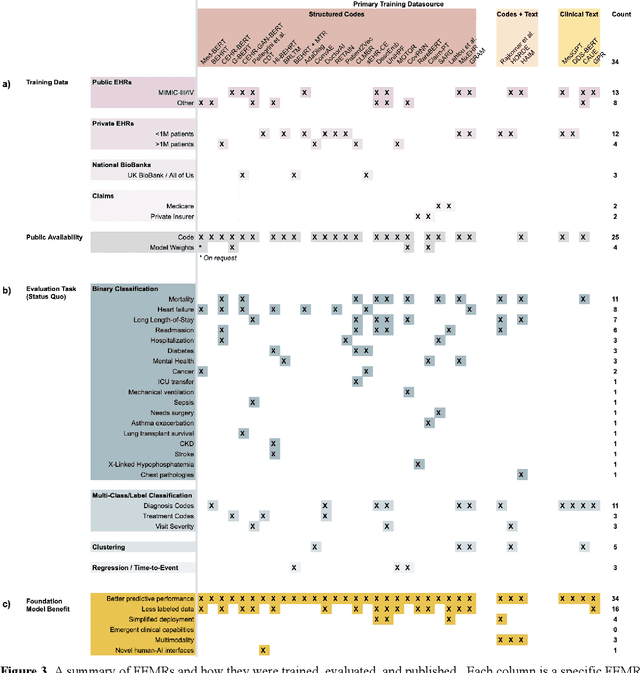

The successes of foundation models such as ChatGPT and AlphaFold have spurred significant interest in building similar models for electronic medical records (EMRs) to improve patient care and hospital operations. However, recent hype has obscured critical gaps in our understanding of these models' capabilities. We review over 80 foundation models trained on non-imaging EMR data (i.e. clinical text and/or structured data) and create a taxonomy delineating their architectures, training data, and potential use cases. We find that most models are trained on small, narrowly-scoped clinical datasets (e.g. MIMIC-III) or broad, public biomedical corpora (e.g. PubMed) and are evaluated on tasks that do not provide meaningful insights on their usefulness to health systems. In light of these findings, we propose an improved evaluation framework for measuring the benefits of clinical foundation models that is more closely grounded to metrics that matter in healthcare.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge