Zachary Freyberg
Deep learning based supervised semantic segmentation of Electron Cryo-Subtomograms
Feb 12, 2018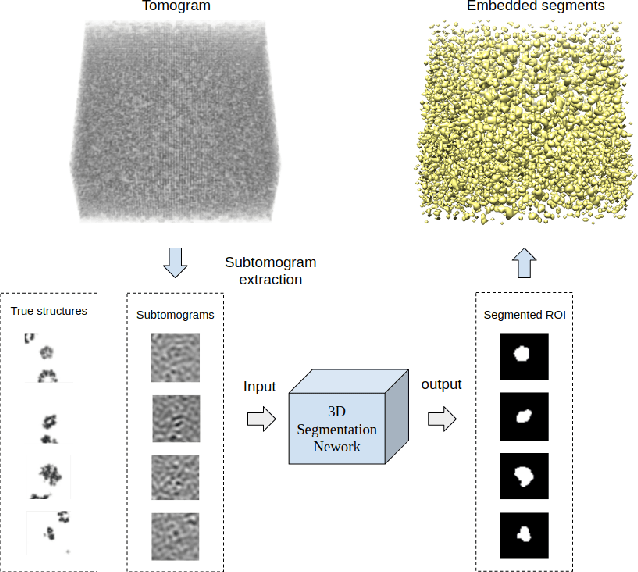
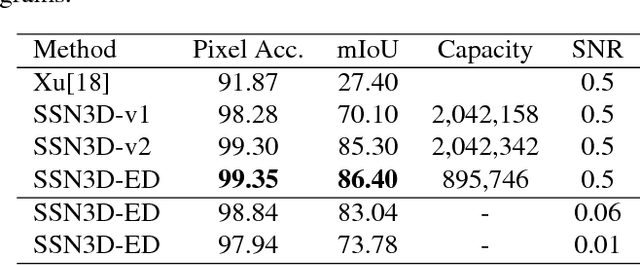
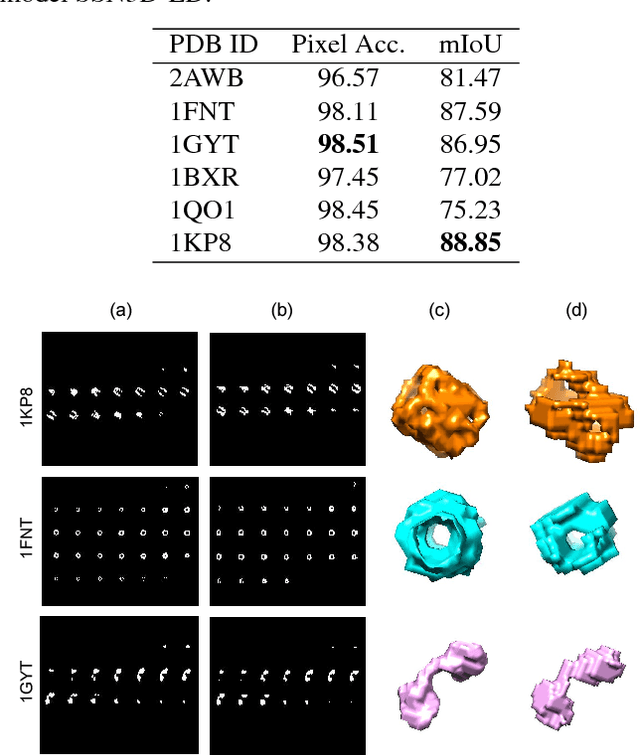
Abstract:Cellular Electron Cryo-Tomography (CECT) is a powerful imaging technique for the 3D visualization of cellular structure and organization at submolecular resolution. It enables analyzing the native structures of macromolecular complexes and their spatial organization inside single cells. However, due to the high degree of structural complexity and practical imaging limitations, systematic macromolecular structural recovery inside CECT images remains challenging. Particularly, the recovery of a macromolecule is likely to be biased by its neighbor structures due to the high molecular crowding. To reduce the bias, here we introduce a novel 3D convolutional neural network inspired by Fully Convolutional Network and Encoder-Decoder Architecture for the supervised segmentation of macromolecules of interest in subtomograms. The tests of our models on realistically simulated CECT data demonstrate that our new approach has significantly improved segmentation performance compared to our baseline approach. Also, we demonstrate that the proposed model has generalization ability to segment new structures that do not exist in training data.
* 9 pages
Model compression for faster structural separation of macromolecules captured by Cellular Electron Cryo-Tomography
Jan 31, 2018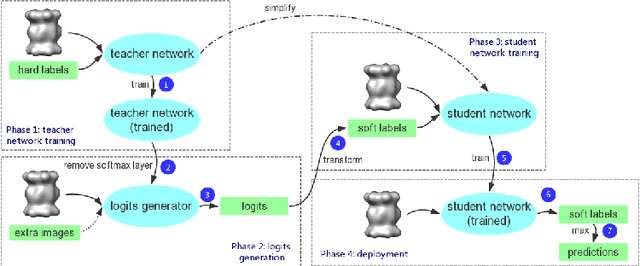
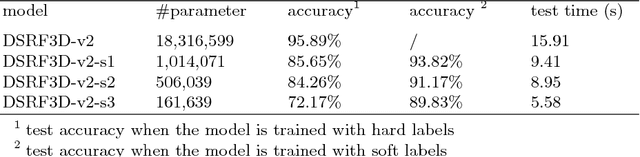

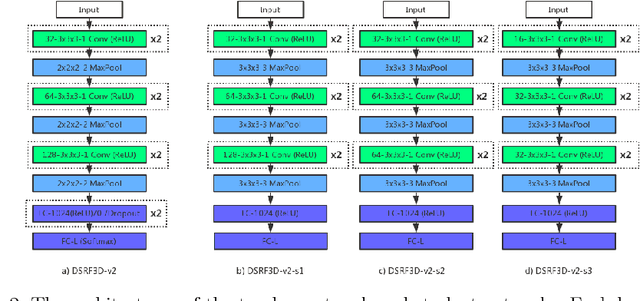
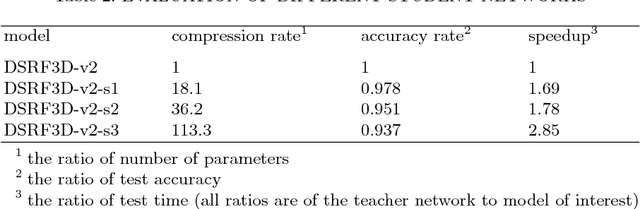
Abstract:Electron Cryo-Tomography (ECT) enables 3D visualization of macromolecule structure inside single cells. Macromolecule classification approaches based on convolutional neural networks (CNN) were developed to separate millions of macromolecules captured from ECT systematically. However, given the fast accumulation of ECT data, it will soon become necessary to use CNN models to efficiently and accurately separate substantially more macromolecules at the prediction stage, which requires additional computational costs. To speed up the prediction, we compress classification models into compact neural networks with little in accuracy for deployment. Specifically, we propose to perform model compression through knowledge distillation. Firstly, a complex teacher network is trained to generate soft labels with better classification feasibility followed by training of customized student networks with simple architectures using the soft label to compress model complexity. Our tests demonstrate that our compressed models significantly reduce the number of parameters and time cost while maintaining similar classification accuracy.
* 8 pages
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge