Yiping Jiao
Pathological Primitive Segmentation Based on Visual Foundation Model with Zero-Shot Mask Generation
Apr 12, 2024Abstract:Medical image processing usually requires a model trained with carefully crafted datasets due to unique image characteristics and domain-specific challenges, especially in pathology. Primitive detection and segmentation in digitized tissue samples are essential for objective and automated diagnosis and prognosis of cancer. SAM (Segment Anything Model) has recently been developed to segment general objects from natural images with high accuracy, but it requires human prompts to generate masks. In this work, we present a novel approach that adapts pre-trained natural image encoders of SAM for detection-based region proposals. Regions proposed by a pre-trained encoder are sent to cascaded feature propagation layers for projection. Then, local semantic and global context is aggregated from multi-scale for bounding box localization and classification. Finally, the SAM decoder uses the identified bounding boxes as essential prompts to generate a comprehensive primitive segmentation map. The entire base framework, SAM, requires no additional training or fine-tuning but could produce an end-to-end result for two fundamental segmentation tasks in pathology. Our method compares with state-of-the-art models in F1 score for nuclei detection and binary/multiclass panoptic(bPQ/mPQ) and mask quality(dice) for segmentation quality on the PanNuke dataset while offering end-to-end efficiency. Our model also achieves remarkable Average Precision (+4.5%) on the secondary dataset (HuBMAP Kidney) compared to Faster RCNN. The code is publicly available at https://github.com/learner-codec/autoprom_sam.
LYSTO: The Lymphocyte Assessment Hackathon and Benchmark Dataset
Jan 16, 2023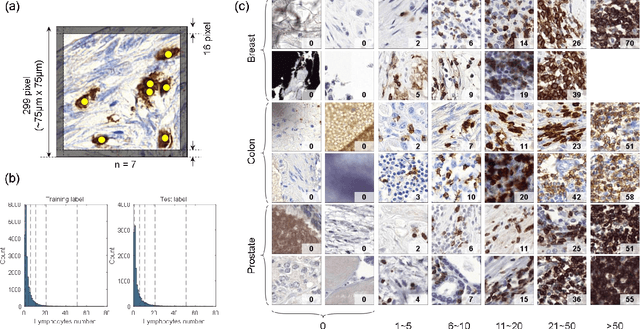
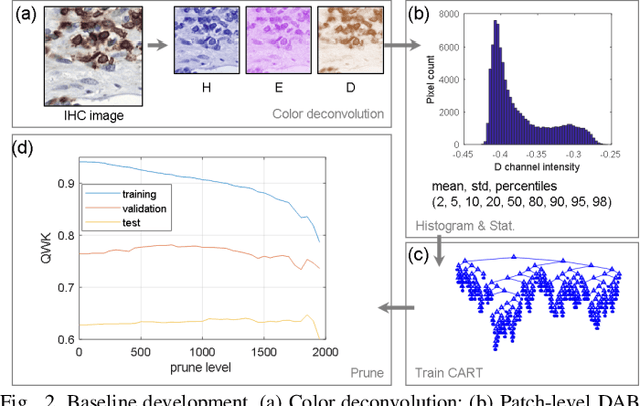
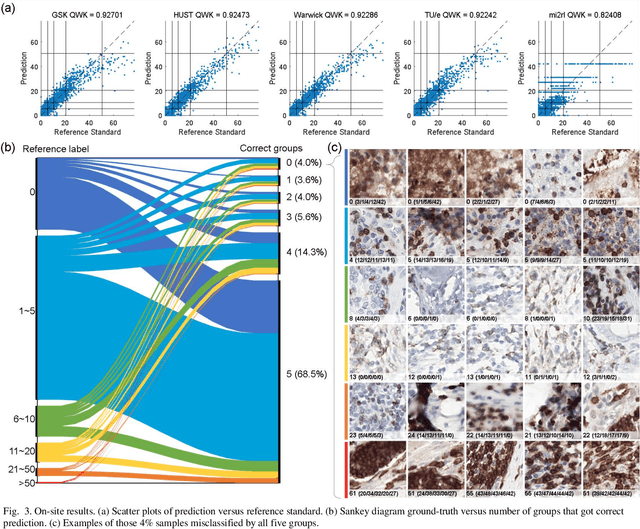
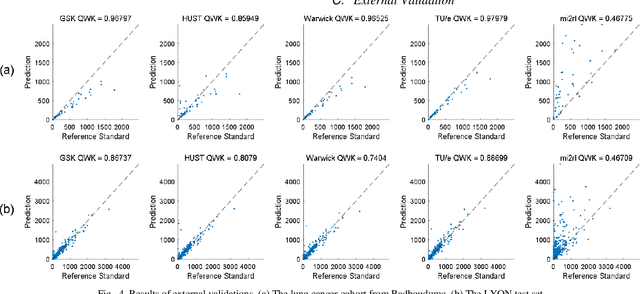
Abstract:We introduce LYSTO, the Lymphocyte Assessment Hackathon, which was held in conjunction with the MICCAI 2019 Conference in Shenzen (China). The competition required participants to automatically assess the number of lymphocytes, in particular T-cells, in histopathological images of colon, breast, and prostate cancer stained with CD3 and CD8 immunohistochemistry. Differently from other challenges setup in medical image analysis, LYSTO participants were solely given a few hours to address this problem. In this paper, we describe the goal and the multi-phase organization of the hackathon; we describe the proposed methods and the on-site results. Additionally, we present post-competition results where we show how the presented methods perform on an independent set of lung cancer slides, which was not part of the initial competition, as well as a comparison on lymphocyte assessment between presented methods and a panel of pathologists. We show that some of the participants were capable to achieve pathologist-level performance at lymphocyte assessment. After the hackathon, LYSTO was left as a lightweight plug-and-play benchmark dataset on grand-challenge website, together with an automatic evaluation platform. LYSTO has supported a number of research in lymphocyte assessment in oncology. LYSTO will be a long-lasting educational challenge for deep learning and digital pathology, it is available at https://lysto.grand-challenge.org/.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge