Xuanzhong Chen
EvoCode-Bench: Evaluating Coding Agents in Multi-Turn Iterative Interactions
May 22, 2026Abstract:Coding agents are increasingly used as iterative development partners, but most benchmarks still evaluate one specification followed by one final assessment. This leaves out a basic question: can an agent keep its own codebase working as requirements change? We introduce EvoCode-Bench, a benchmark of 26 stateful coding tasks and 227 evaluated rounds. Each task preserves the agent's workspace for 5-15 rounds, states requirements through observable behavior, and uses cumulative executable tests to check new requirements and still-active prior ones. We evaluate 13 coding agents with two metrics: MT@4, a four-attempt fail-stop multi-round score, and SR, a single-round score from a reference-completed prior state. For most agents, SR exceeds MT@4 by 22-40 points. The gap also changes rankings: the highest-SR agent (78.9) ranks only third in persistent execution (44.0 MT@4). Even the strongest agents achieve only about 50% success on multi-turn metrics, and aggregate pass rate drops below half of round-1 performance by round 5. Failure analysis shows tier-dependent behavior: weaker agents fail early, while stronger agents survive long enough to expose specification-tracking and regression failures. We release the benchmark data and Harbor multi-turn infrastructure.
MindLoom: Composing Thought Modes for Frontier-Level Reasoning Data Synthesis
May 20, 2026Abstract:Although LLMs have made substantial progress in reasoning, systematically producing frontier-level reasoning data remains difficult. Existing synthesis methods often have limited visibility into the structural factors that govern problem difficulty, which can result in narrow diversity and unstable difficulty control. In this work, we view the difficulty of a reasoning problem as arising from the accumulation of atomic knowledge-reasoning transformations, which we term thought modes. Building on this perspective, we propose MindLoom, a framework for synthesizing frontier-level reasoning data through compositional thought mode engineering. Given a collection of hard problems with verified solutions, MindLoom first decomposes those solutions into thought mode chains that reveal each problem's construction logic. It then trains a retrieval model that matches problem states to compatible thought modes, providing guidance on which reasoning challenges to introduce during synthesis. New problems are composed by iteratively applying retrieved thought modes to seed questions, with distribution-aligned sampling to encourage diverse reasoning coverage. Finally, a rollout-based judging stage labels generated questions by difficulty and supplies judged-correct responses for supervised fine-tuning. We evaluate MindLoom on nine benchmarks covering five STEM disciplines and four mathematical reasoning tasks across multiple model families and sizes. Models fine-tuned on MindLoom-generated data achieves favorable performances over base models, distillation, and external-data baselines across the reported benchmarks. Ablation studies indicate the contribution of each component, and further analysis suggests that MindLoom covers a broad range of reasoning patterns while maintaining useful difficulty control. We have open-sourced our implementation at https://github.com/EachSheep/MindLoom.
BabyVision: Visual Reasoning Beyond Language
Jan 10, 2026Abstract:While humans develop core visual skills long before acquiring language, contemporary Multimodal LLMs (MLLMs) still rely heavily on linguistic priors to compensate for their fragile visual understanding. We uncovered a crucial fact: state-of-the-art MLLMs consistently fail on basic visual tasks that humans, even 3-year-olds, can solve effortlessly. To systematically investigate this gap, we introduce BabyVision, a benchmark designed to assess core visual abilities independent of linguistic knowledge for MLLMs. BabyVision spans a wide range of tasks, with 388 items divided into 22 subclasses across four key categories. Empirical results and human evaluation reveal that leading MLLMs perform significantly below human baselines. Gemini3-Pro-Preview scores 49.7, lagging behind 6-year-old humans and falling well behind the average adult score of 94.1. These results show despite excelling in knowledge-heavy evaluations, current MLLMs still lack fundamental visual primitives. Progress in BabyVision represents a step toward human-level visual perception and reasoning capabilities. We also explore solving visual reasoning with generation models by proposing BabyVision-Gen and automatic evaluation toolkit. Our code and benchmark data are released at https://github.com/UniPat-AI/BabyVision for reproduction.
EcomBench: Towards Holistic Evaluation of Foundation Agents in E-commerce
Dec 11, 2025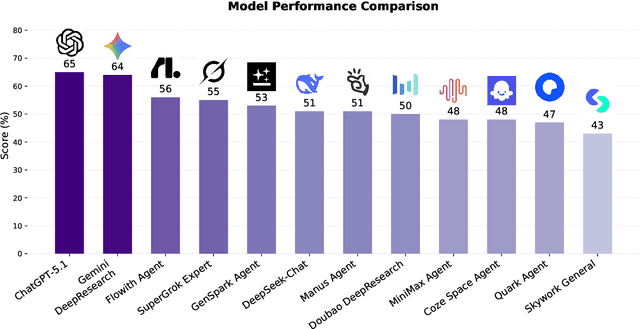
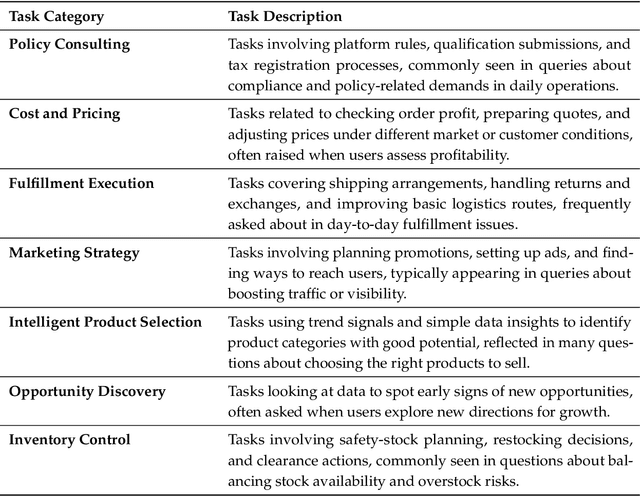

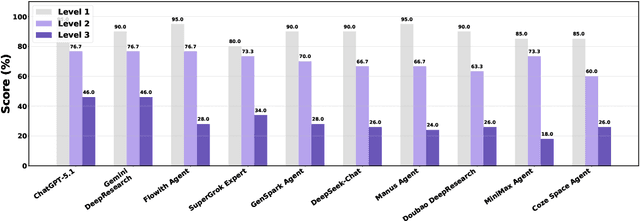
Abstract:Foundation agents have rapidly advanced in their ability to reason and interact with real environments, making the evaluation of their core capabilities increasingly important. While many benchmarks have been developed to assess agent performance, most concentrate on academic settings or artificially designed scenarios while overlooking the challenges that arise in real applications. To address this issue, we focus on a highly practical real-world setting, the e-commerce domain, which involves a large volume of diverse user interactions, dynamic market conditions, and tasks directly tied to real decision-making processes. To this end, we introduce EcomBench, a holistic E-commerce Benchmark designed to evaluate agent performance in realistic e-commerce environments. EcomBench is built from genuine user demands embedded in leading global e-commerce ecosystems and is carefully curated and annotated through human experts to ensure clarity, accuracy, and domain relevance. It covers multiple task categories within e-commerce scenarios and defines three difficulty levels that evaluate agents on key capabilities such as deep information retrieval, multi-step reasoning, and cross-source knowledge integration. By grounding evaluation in real e-commerce contexts, EcomBench provides a rigorous and dynamic testbed for measuring the practical capabilities of agents in modern e-commerce.
IterResearch: Rethinking Long-Horizon Agents via Markovian State Reconstruction
Nov 10, 2025Abstract:Recent advances in deep-research agents have shown promise for autonomous knowledge construction through dynamic reasoning over external sources. However, existing approaches rely on a mono-contextual paradigm that accumulates all information in a single, expanding context window, leading to context suffocation and noise contamination that limit their effectiveness on long-horizon tasks. We introduce IterResearch, a novel iterative deep-research paradigm that reformulates long-horizon research as a Markov Decision Process with strategic workspace reconstruction. By maintaining an evolving report as memory and periodically synthesizing insights, our approach preserves consistent reasoning capacity across arbitrary exploration depths. We further develop Efficiency-Aware Policy Optimization (EAPO), a reinforcement learning framework that incentivizes efficient exploration through geometric reward discounting and enables stable distributed training via adaptive downsampling. Extensive experiments demonstrate that IterResearch achieves substantial improvements over existing open-source agents with average +14.5pp across six benchmarks and narrows the gap with frontier proprietary systems. Remarkably, our paradigm exhibits unprecedented interaction scaling, extending to 2048 interactions with dramatic performance gains (from 3.5\% to 42.5\%), and serves as an effective prompting strategy, improving frontier models by up to 19.2pp over ReAct on long-horizon tasks. These findings position IterResearch as a versatile solution for long-horizon reasoning, effective both as a trained agent and as a prompting paradigm for frontier models.
MARS: Optimizing Dual-System Deep Research via Multi-Agent Reinforcement Learning
Oct 06, 2025Abstract:Large Reasoning Models (LRMs) often exhibit a tendency for overanalysis in simple tasks, where the models excessively utilize System 2-type, deliberate reasoning, leading to inefficient token generation. Furthermore, these models face challenges in adapting their reasoning capabilities to rapidly changing environments due to the static nature of their pretraining data. To address these issues, advancing Large Language Models (LLMs) for complex reasoning tasks requires innovative approaches that bridge intuitive and deliberate cognitive processes, akin to human cognition's dual-system dynamic. This paper introduces a Multi-Agent System for Deep ReSearch (MARS) enabling seamless integration of System 1's fast, intuitive thinking with System 2's deliberate reasoning within LLMs. MARS strategically integrates multiple external tools, such as Google Search, Google Scholar, and Python Interpreter, to access up-to-date information and execute complex computations, while creating a specialized division of labor where System 1 efficiently processes and summarizes high-volume external information, providing distilled insights that expand System 2's reasoning context without overwhelming its capacity. Furthermore, we propose a multi-agent reinforcement learning framework extending Group Relative Policy Optimization to simultaneously optimize both systems with multi-turn tool interactions, bin-packing optimization, and sample balancing strategies that enhance collaborative efficiency. Extensive experiments demonstrate MARS achieves substantial improvements of 3.86% on the challenging Humanity's Last Exam (HLE) benchmark and an average gain of 8.9% across 7 knowledge-intensive tasks, validating the effectiveness of our dual-system paradigm for complex reasoning in dynamic information environments.
Scaling Agents via Continual Pre-training
Sep 16, 2025Abstract:Large language models (LLMs) have evolved into agentic systems capable of autonomous tool use and multi-step reasoning for complex problem-solving. However, post-training approaches building upon general-purpose foundation models consistently underperform in agentic tasks, particularly in open-source implementations. We identify the root cause: the absence of robust agentic foundation models forces models during post-training to simultaneously learn diverse agentic behaviors while aligning them to expert demonstrations, thereby creating fundamental optimization tensions. To this end, we are the first to propose incorporating Agentic Continual Pre-training (Agentic CPT) into the deep research agents training pipeline to build powerful agentic foundational models. Based on this approach, we develop a deep research agent model named AgentFounder. We evaluate our AgentFounder-30B on 10 benchmarks and achieve state-of-the-art performance while retains strong tool-use ability, notably 39.9% on BrowseComp-en, 43.3% on BrowseComp-zh, and 31.5% Pass@1 on HLE.
WebResearcher: Unleashing unbounded reasoning capability in Long-Horizon Agents
Sep 16, 2025



Abstract:Recent advances in deep-research systems have demonstrated the potential for AI agents to autonomously discover and synthesize knowledge from external sources. In this paper, we introduce WebResearcher, a novel framework for building such agents through two key components: (1) WebResearcher, an iterative deep-research paradigm that reformulates deep research as a Markov Decision Process, where agents periodically consolidate findings into evolving reports while maintaining focused workspaces, overcoming the context suffocation and noise contamination that plague existing mono-contextual approaches; and (2) WebFrontier, a scalable data synthesis engine that generates high-quality training data through tool-augmented complexity escalation, enabling systematic creation of research tasks that bridge the gap between passive knowledge recall and active knowledge construction. Notably, we find that the training data from our paradigm significantly enhances tool-use capabilities even for traditional mono-contextual methods. Furthermore, our paradigm naturally scales through parallel thinking, enabling concurrent multi-agent exploration for more comprehensive conclusions. Extensive experiments across 6 challenging benchmarks demonstrate that WebResearcher achieves state-of-the-art performance, even surpassing frontier proprietary systems.
RareAgents: Autonomous Multi-disciplinary Team for Rare Disease Diagnosis and Treatment
Dec 17, 2024



Abstract:Rare diseases, despite their low individual incidence, collectively impact around 300 million people worldwide due to the huge number of diseases. The complexity of symptoms and the shortage of specialized doctors with relevant experience make diagnosing and treating rare diseases more challenging than common diseases. Recently, agents powered by large language models (LLMs) have demonstrated notable improvements across various domains. In the medical field, some agent methods have outperformed direct prompts in question-answering tasks from medical exams. However, current agent frameworks lack adaptation for real-world clinical scenarios, especially those involving the intricate demands of rare diseases. To address these challenges, we present RareAgents, the first multi-disciplinary team of LLM-based agents tailored to the complex clinical context of rare diseases. RareAgents integrates advanced planning capabilities, memory mechanisms, and medical tools utilization, leveraging Llama-3.1-8B/70B as the base model. Experimental results show that RareAgents surpasses state-of-the-art domain-specific models, GPT-4o, and existing agent frameworks in both differential diagnosis and medication recommendation for rare diseases. Furthermore, we contribute a novel dataset, MIMIC-IV-Ext-Rare, derived from MIMIC-IV, to support further advancements in this field.
VividMed: Vision Language Model with Versatile Visual Grounding for Medicine
Oct 16, 2024



Abstract:Recent advancements in Vision Language Models (VLMs) have demonstrated remarkable promise in generating visually grounded responses. However, their application in the medical domain is hindered by unique challenges. For instance, most VLMs rely on a single method of visual grounding, whereas complex medical tasks demand more versatile approaches. Additionally, while most VLMs process only 2D images, a large portion of medical images are 3D. The lack of medical data further compounds these obstacles. To address these challenges, we present VividMed, a vision language model with versatile visual grounding for medicine. Our model supports generating both semantic segmentation masks and instance-level bounding boxes, and accommodates various imaging modalities, including both 2D and 3D data. We design a three-stage training procedure and an automatic data synthesis pipeline based on open datasets and models. Besides visual grounding tasks, VividMed also excels in other common downstream tasks, including Visual Question Answering (VQA) and report generation. Ablation studies empirically show that the integration of visual grounding ability leads to improved performance on these tasks. Our code is publicly available at https://github.com/function2-llx/MMMM.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge