Vijayalakshmi Senthil
The Three-Dimensional Structural Configuration of the Central Retinal Vessel Trunk and Branches as a Glaucoma Biomarker
Nov 09, 2021
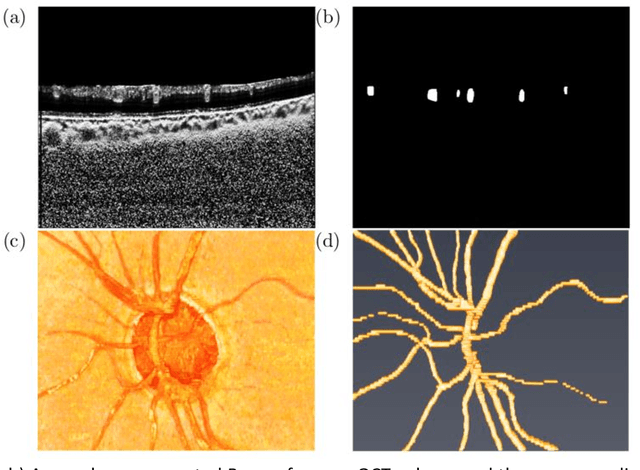
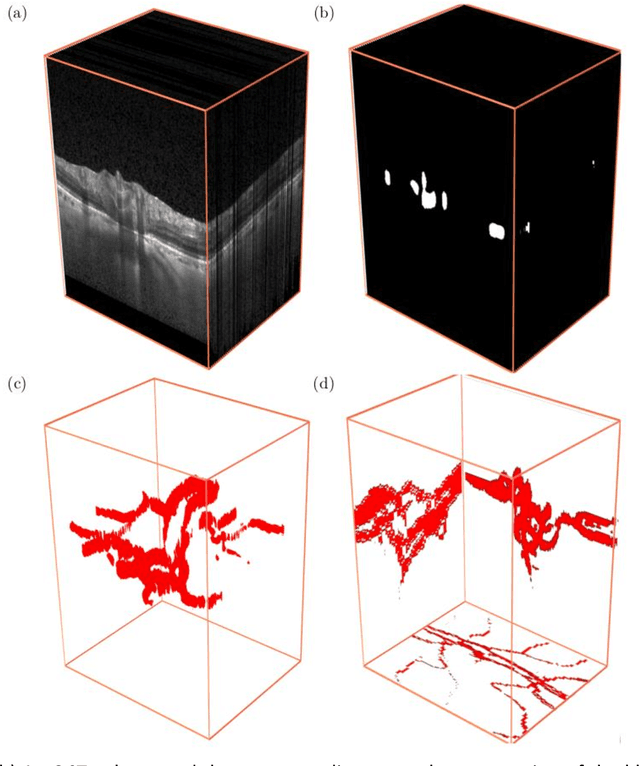

Abstract:Purpose: To assess whether the three-dimensional (3D) structural configuration of the central retinal vessel trunk and its branches (CRVT&B) could be used as a diagnostic marker for glaucoma. Method: We trained a deep learning network to automatically segment the CRVT&B from the B-scans of the optical coherence tomography (OCT) volume of the optic nerve head (ONH). Subsequently, two different approaches were used for glaucoma diagnosis using the structural configuration of the CRVT&B as extracted from the OCT volumes. In the first approach, we aimed to provide a diagnosis using only 3D CNN and the 3D structure of the CRVT&B. For the second approach, we projected the 3D structure of the CRVT&B orthographically onto three planes to obtain 2D images, and then a 2D CNN was used for diagnosis. The segmentation accuracy was evaluated using the Dice coefficient, whereas the diagnostic accuracy was assessed using the area under the receiver operating characteristic curves (AUC). The diagnostic performance of the CRVT&B was also compared with that of retinal nerve fiber layer (RNFL) thickness. Results: Our segmentation network was able to efficiently segment retinal blood vessels from OCT scans. On a test set, we achieved a Dice coefficient of 0.81\pm0.07. The 3D and 2D diagnostic networks were able to differentiate glaucoma from non-glaucoma subjects with accuracies of 82.7% and 83.3%, respectively. The corresponding AUCs for CRVT&B were 0.89 and 0.90, higher than those obtained with RNFL thickness alone. Conclusions: Our work demonstrated that the diagnostic power of the CRVT&B is superior to that of a gold-standard glaucoma parameter, i.e., RNFL thickness. Our work also suggested that the major retinal blood vessels form a skeleton -- the configuration of which may be representative of major ONH structural changes as typically observed with the development and progression of glaucoma.
Towards Label-Free 3D Segmentation of Optical Coherence Tomography Images of the Optic Nerve Head Using Deep Learning
Feb 22, 2020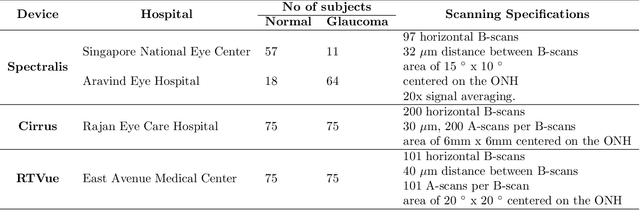
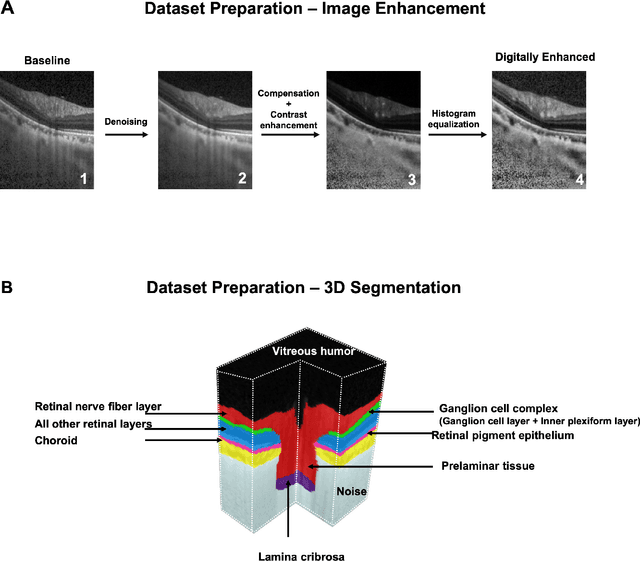
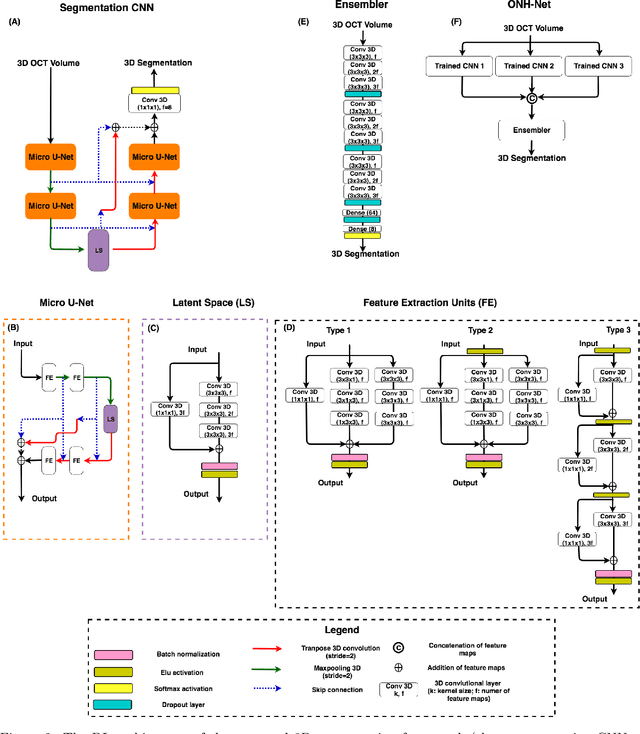
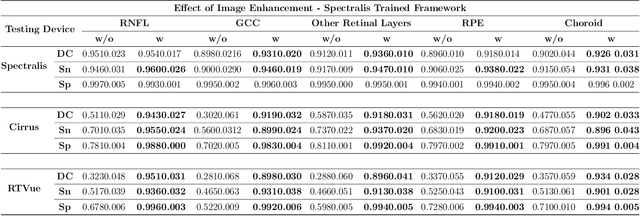
Abstract:Since the introduction of optical coherence tomography (OCT), it has been possible to study the complex 3D morphological changes of the optic nerve head (ONH) tissues that occur along with the progression of glaucoma. Although several deep learning (DL) techniques have been recently proposed for the automated extraction (segmentation) and quantification of these morphological changes, the device specific nature and the difficulty in preparing manual segmentations (training data) limit their clinical adoption. With several new manufacturers and next-generation OCT devices entering the market, the complexity in deploying DL algorithms clinically is only increasing. To address this, we propose a DL based 3D segmentation framework that is easily translatable across OCT devices in a label-free manner (i.e. without the need to manually re-segment data for each device). Specifically, we developed 2 sets of DL networks. The first (referred to as the enhancer) was able to enhance OCT image quality from 3 OCT devices, and harmonized image-characteristics across these devices. The second performed 3D segmentation of 6 important ONH tissue layers. We found that the use of the enhancer was critical for our segmentation network to achieve device independency. In other words, our 3D segmentation network trained on any of 3 devices successfully segmented ONH tissue layers from the other two devices with high performance (Dice coefficients > 0.92). With such an approach, we could automatically segment images from new OCT devices without ever needing manual segmentation data from such devices.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge