Thomas Kroencke
Detecting Unforeseen Data Properties with Diffusion Autoencoder Embeddings using Spine MRI data
Oct 14, 2024

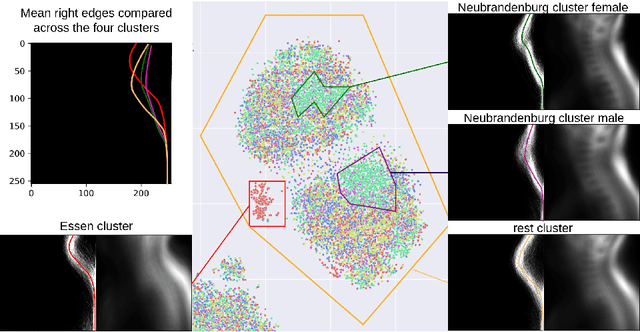
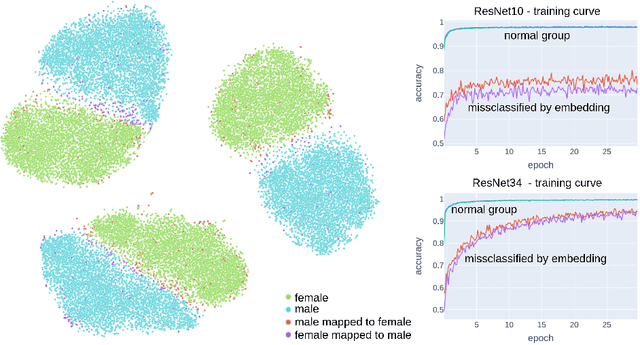
Abstract:Deep learning has made significant strides in medical imaging, leveraging the use of large datasets to improve diagnostics and prognostics. However, large datasets often come with inherent errors through subject selection and acquisition. In this paper, we investigate the use of Diffusion Autoencoder (DAE) embeddings for uncovering and understanding data characteristics and biases, including biases for protected variables like sex and data abnormalities indicative of unwanted protocol variations. We use sagittal T2-weighted magnetic resonance (MR) images of the neck, chest, and lumbar region from 11186 German National Cohort (NAKO) participants. We compare DAE embeddings with existing generative models like StyleGAN and Variational Autoencoder. Evaluations on a large-scale dataset consisting of sagittal T2-weighted MR images of three spine regions show that DAE embeddings effectively separate protected variables such as sex and age. Furthermore, we used t-SNE visualization to identify unwanted variations in imaging protocols, revealing differences in head positioning. Our embedding can identify samples where a sex predictor will have issues learning the correct sex. Our findings highlight the potential of using advanced embedding techniques like DAEs to detect data quality issues and biases in medical imaging datasets. Identifying such hidden relations can enhance the reliability and fairness of deep learning models in healthcare applications, ultimately improving patient care and outcomes.
SPINEPS -- Automatic Whole Spine Segmentation of T2-weighted MR images using a Two-Phase Approach to Multi-class Semantic and Instance Segmentation
Feb 26, 2024
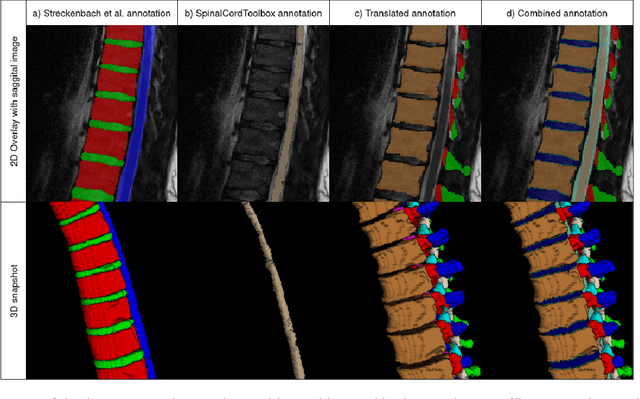
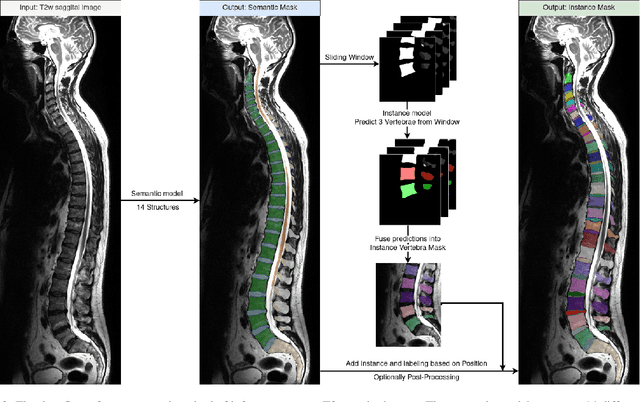
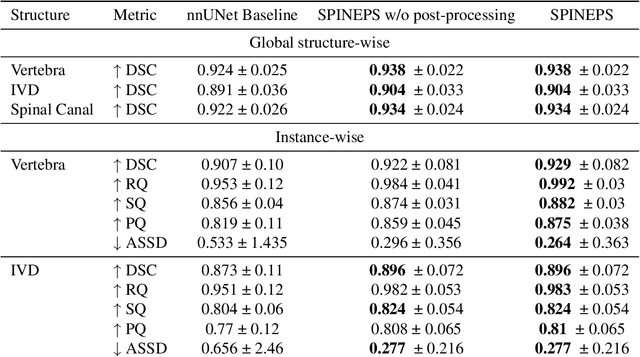
Abstract:Purpose. To present SPINEPS, an open-source deep learning approach for semantic and instance segmentation of 14 spinal structures (ten vertebra substructures, intervertebral discs, spinal cord, spinal canal, and sacrum) in whole body T2w MRI. Methods. During this HIPPA-compliant, retrospective study, we utilized the public SPIDER dataset (218 subjects, 63% female) and a subset of the German National Cohort (1423 subjects, mean age 53, 49% female) for training and evaluation. We combined CT and T2w segmentations to train models that segment 14 spinal structures in T2w sagittal scans both semantically and instance-wise. Performance evaluation metrics included Dice similarity coefficient, average symmetrical surface distance, panoptic quality, segmentation quality, and recognition quality. Statistical significance was assessed using the Wilcoxon signed-rank test. An in-house dataset was used to qualitatively evaluate out-of-distribution samples. Results. On the public dataset, our approach outperformed the baseline (instance-wise vertebra dice score 0.929 vs. 0.907, p-value<0.001). Training on auto-generated annotations and evaluating on manually corrected test data from the GNC yielded global dice scores of 0.900 for vertebrae, 0.960 for intervertebral discs, and 0.947 for the spinal canal. Incorporating the SPIDER dataset during training increased these scores to 0.920, 0.967, 0.958, respectively. Conclusions. The proposed segmentation approach offers robust segmentation of 14 spinal structures in T2w sagittal images, including the spinal cord, spinal canal, intervertebral discs, endplate, sacrum, and vertebrae. The approach yields both a semantic and instance mask as output, thus being easy to utilize. This marks the first publicly available algorithm for whole spine segmentation in sagittal T2w MR imaging.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge