Stephanie Hong
Analyzing historical diagnosis code data from NIH N3C and RECOVER Programs using deep learning to determine risk factors for Long Covid
Oct 05, 2022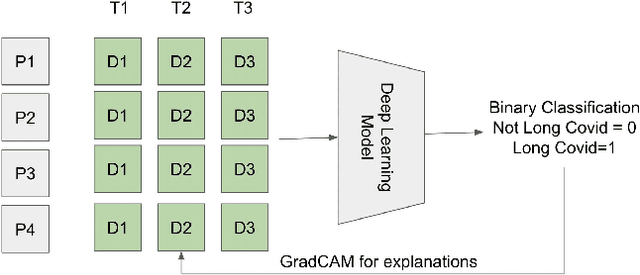
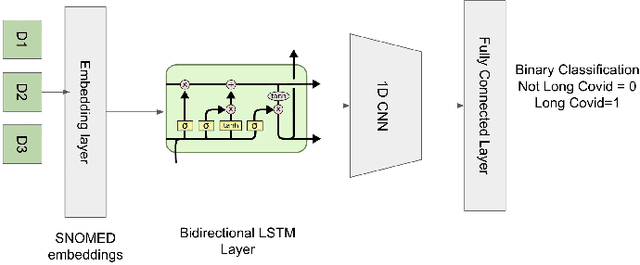
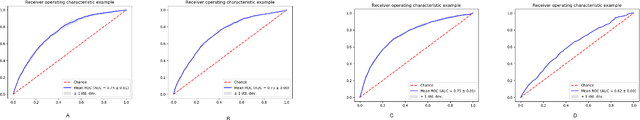
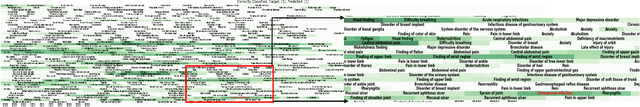
Abstract:Post-acute sequelae of SARS-CoV-2 infection (PASC) or Long COVID is an emerging medical condition that has been observed in several patients with a positive diagnosis for COVID-19. Historical Electronic Health Records (EHR) like diagnosis codes, lab results and clinical notes have been analyzed using deep learning and have been used to predict future clinical events. In this paper, we propose an interpretable deep learning approach to analyze historical diagnosis code data from the National COVID Cohort Collective (N3C) to find the risk factors contributing to developing Long COVID. Using our deep learning approach, we are able to predict if a patient is suffering from Long COVID from a temporally ordered list of diagnosis codes up to 45 days post the first COVID positive test or diagnosis for each patient, with an accuracy of 70.48\%. We are then able to examine the trained model using Gradient-weighted Class Activation Mapping (GradCAM) to give each input diagnoses a score. The highest scored diagnosis were deemed to be the most important for making the correct prediction for a patient. We also propose a way to summarize these top diagnoses for each patient in our cohort and look at their temporal trends to determine which codes contribute towards a positive Long COVID diagnosis.
A Methodological Framework for the Comparative Evaluation of Multiple Imputation Methods: Multiple Imputation of Race, Ethnicity and Body Mass Index in the U.S. National COVID Cohort Collaborative
Jun 13, 2022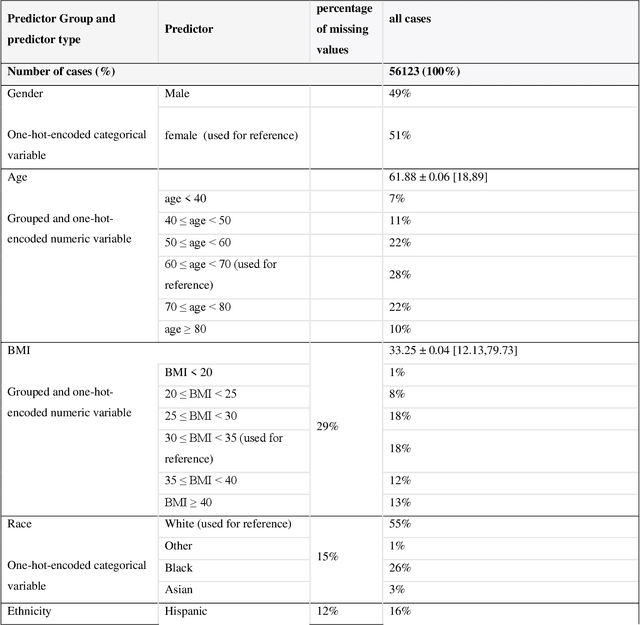
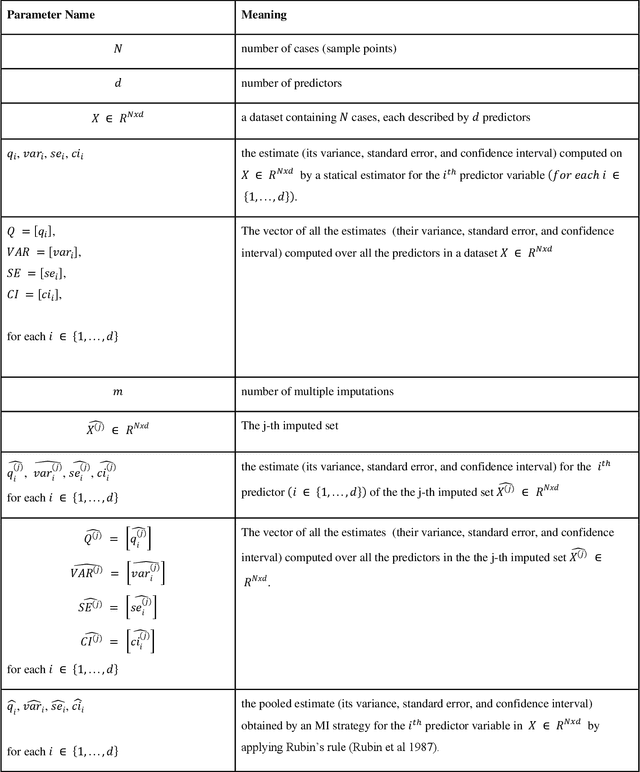
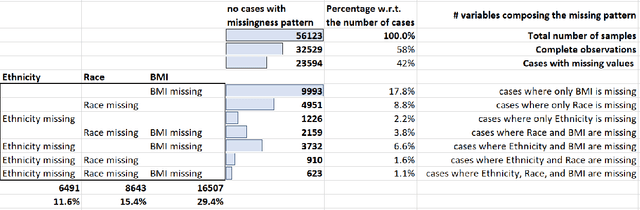

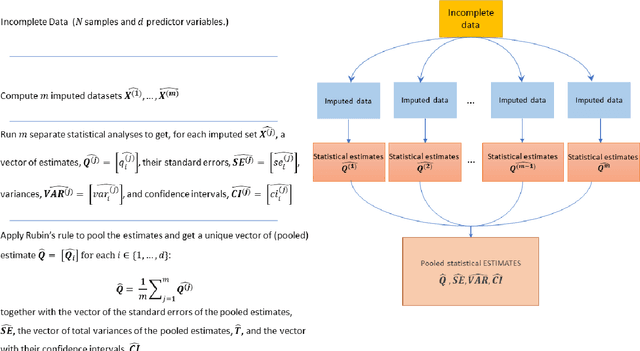
Abstract:While electronic health records are a rich data source for biomedical research, these systems are not implemented uniformly across healthcare settings and significant data may be missing due to healthcare fragmentation and lack of interoperability between siloed electronic health records. Considering that the deletion of cases with missing data may introduce severe bias in the subsequent analysis, several authors prefer applying a multiple imputation strategy to recover the missing information. Unfortunately, although several literature works have documented promising results by using any of the different multiple imputation algorithms that are now freely available for research, there is no consensus on which MI algorithm works best. Beside the choice of the MI strategy, the choice of the imputation algorithm and its application settings are also both crucial and challenging. In this paper, inspired by the seminal works of Rubin and van Buuren, we propose a methodological framework that may be applied to evaluate and compare several multiple imputation techniques, with the aim to choose the most valid for computing inferences in a clinical research work. Our framework has been applied to validate, and extend on a larger cohort, the results we presented in a previous literature study, where we evaluated the influence of crucial patients' descriptors and COVID-19 severity in patients with type 2 diabetes mellitus whose data is provided by the National COVID Cohort Collaborative Enclave.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge