Saverio Vadacchino
BigDocs: An Open and Permissively-Licensed Dataset for Training Multimodal Models on Document and Code Tasks
Dec 05, 2024



Abstract:Multimodal AI has the potential to significantly enhance document-understanding tasks, such as processing receipts, understanding workflows, extracting data from documents, and summarizing reports. Code generation tasks that require long-structured outputs can also be enhanced by multimodality. Despite this, their use in commercial applications is often limited due to limited access to training data and restrictive licensing, which hinders open access. To address these limitations, we introduce BigDocs-7.5M, a high-quality, open-access dataset comprising 7.5 million multimodal documents across 30 tasks. We use an efficient data curation process to ensure our data is high-quality and license-permissive. Our process emphasizes accountability, responsibility, and transparency through filtering rules, traceable metadata, and careful content analysis. Additionally, we introduce BigDocs-Bench, a benchmark suite with 10 novel tasks where we create datasets that reflect real-world use cases involving reasoning over Graphical User Interfaces (GUI) and code generation from images. Our experiments show that training with BigDocs-Bench improves average performance up to 25.8% over closed-source GPT-4o in document reasoning and structured output tasks such as Screenshot2HTML or Image2Latex generation. Finally, human evaluations showed a preference for outputs from models trained on BigDocs over GPT-4o. This suggests that BigDocs can help both academics and the open-source community utilize and improve AI tools to enhance multimodal capabilities and document reasoning. The project is hosted at https://bigdocs.github.io .
HAD-Net: A Hierarchical Adversarial Knowledge Distillation Network for Improved Enhanced Tumour Segmentation Without Post-Contrast Images
Apr 09, 2021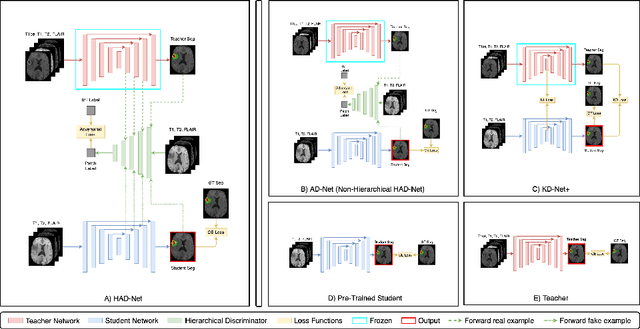



Abstract:Segmentation of enhancing tumours or lesions from MRI is important for detecting new disease activity in many clinical contexts. However, accurate segmentation requires the inclusion of medical images (e.g., T1 post contrast MRI) acquired after injecting patients with a contrast agent (e.g., Gadolinium), a process no longer thought to be safe. Although a number of modality-agnostic segmentation networks have been developed over the past few years, they have been met with limited success in the context of enhancing pathology segmentation. In this work, we present HAD-Net, a novel offline adversarial knowledge distillation (KD) technique, whereby a pre-trained teacher segmentation network, with access to all MRI sequences, teaches a student network, via hierarchical adversarial training, to better overcome the large domain shift presented when crucial images are absent during inference. In particular, we apply HAD-Net to the challenging task of enhancing tumour segmentation when access to post-contrast imaging is not available. The proposed network is trained and tested on the BraTS 2019 brain tumour segmentation challenge dataset, where it achieves performance improvements in the ranges of 16% - 26% over (a) recent modality-agnostic segmentation methods (U-HeMIS, U-HVED), (b) KD-Net adapted to this problem, (c) the pre-trained student network and (d) a non-hierarchical version of the network (AD-Net), in terms of Dice scores for enhancing tumour (ET). The network also shows improvements in tumour core (TC) Dice scores. Finally, the network outperforms both the baseline student network and AD-Net in terms of uncertainty quantification for enhancing tumour segmentation based on the BraTs 2019 uncertainty challenge metrics. Our code is publicly available at: https://github.com/SaverioVad/HAD_Net
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge