Sarwan Ali
DPSR: Differentially Private Sparse Reconstruction via Multi-Stage Denoising for Recommender Systems
Dec 22, 2025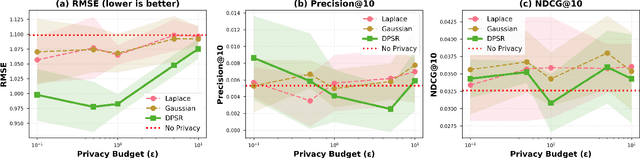



Abstract:Differential privacy (DP) has emerged as the gold standard for protecting user data in recommender systems, but existing privacy-preserving mechanisms face a fundamental challenge: the privacy-utility tradeoff inevitably degrades recommendation quality as privacy budgets tighten. We introduce DPSR (Differentially Private Sparse Reconstruction), a novel three-stage denoising framework that fundamentally addresses this limitation by exploiting the inherent structure of rating matrices -- sparsity, low-rank properties, and collaborative patterns. DPSR consists of three synergistic stages: (1) \textit{information-theoretic noise calibration} that adaptively reduces noise for high-information ratings, (2) \textit{collaborative filtering-based denoising} that leverages item-item similarities to remove privacy noise, and (3) \textit{low-rank matrix completion} that exploits latent structure for signal recovery. Critically, all denoising operations occur \textit{after} noise injection, preserving differential privacy through the post-processing immunity theorem while removing both privacy-induced and inherent data noise. Through extensive experiments on synthetic datasets with controlled ground truth, we demonstrate that DPSR achieves 5.57\% to 9.23\% RMSE improvement over state-of-the-art Laplace and Gaussian mechanisms across privacy budgets ranging from $\varepsilon=0.1$ to $\varepsilon=10.0$ (all improvements statistically significant with $p < 0.05$, most $p < 0.001$). Remarkably, at $\varepsilon=1.0$, DPSR achieves RMSE of 0.9823, \textit{outperforming even the non-private baseline} (1.0983), demonstrating that our denoising pipeline acts as an effective regularizer that removes data noise in addition to privacy noise.
Boosting t-SNE Efficiency for Sequencing Data: Insights from Kernel Selection
Dec 17, 2025



Abstract:Dimensionality reduction techniques are essential for visualizing and analyzing high-dimensional biological sequencing data. t-distributed Stochastic Neighbor Embedding (t-SNE) is widely used for this purpose, traditionally employing the Gaussian kernel to compute pairwise similarities. However, the Gaussian kernel's lack of data-dependence and computational overhead limit its scalability and effectiveness for categorical biological sequences. Recent work proposed the isolation kernel as an alternative, yet it may not optimally capture sequence similarities. In this study, we comprehensively evaluate nine different kernel functions for t-SNE applied to molecular sequences, using three embedding methods: One-Hot Encoding, Spike2Vec, and minimizers. Through both subjective visualization and objective metrics (including neighborhood preservation scores), we demonstrate that the cosine similarity kernel in general outperforms other kernels, including Gaussian and isolation kernels, achieving superior runtime efficiency and better preservation of pairwise distances in low-dimensional space. We further validate our findings through extensive classification and clustering experiments across six diverse biological datasets (Spike7k, Host, ShortRead, Rabies, Genome, and Breast Cancer), employing multiple machine learning algorithms and evaluation metrics. Our results show that kernel selection significantly impacts not only visualization quality but also downstream analytical tasks, with the cosine similarity kernel providing the most robust performance across different data types and embedding strategies, making it particularly suitable for large-scale biological sequence analysis.
Robust Gradient Descent via Heavy-Ball Momentum with Predictive Extrapolation
Dec 10, 2025Abstract:Accelerated gradient methods like Nesterov's Accelerated Gradient (NAG) achieve faster convergence on well-conditioned problems but often diverge on ill-conditioned or non-convex landscapes due to aggressive momentum accumulation. We propose Heavy-Ball Synthetic Gradient Extrapolation (HB-SGE), a robust first-order method that combines heavy-ball momentum with predictive gradient extrapolation. Unlike classical momentum methods that accumulate historical gradients, HB-SGE estimates future gradient directions using local Taylor approximations, providing adaptive acceleration while maintaining stability. We prove convergence guarantees for strongly convex functions and demonstrate empirically that HB-SGE prevents divergence on problems where NAG and standard momentum fail. On ill-conditioned quadratics (condition number $κ=50$), HB-SGE converges in 119 iterations while both SGD and NAG diverge. On the non-convex Rosenbrock function, HB-SGE achieves convergence in 2,718 iterations where classical momentum methods diverge within 10 steps. While NAG remains faster on well-conditioned problems, HB-SGE provides a robust alternative with speedup over SGD across diverse landscapes, requiring only $O(d)$ memory overhead and the same hyperparameters as standard momentum.
Sequence-to-Image Transformation for Sequence Classification Using Rips Complex Construction and Chaos Game Representation
Dec 10, 2025Abstract:Traditional feature engineering approaches for molecular sequence classification suffer from sparsity issues and computational complexity, while deep learning models often underperform on tabular biological data. This paper introduces a novel topological approach that transforms molecular sequences into images by combining Chaos Game Representation (CGR) with Rips complex construction from algebraic topology. Our method maps sequence elements to 2D coordinates via CGR, computes pairwise distances, and constructs Rips complexes to capture both local structural and global topological features. We provide formal guarantees on representation uniqueness, topological stability, and information preservation. Extensive experiments on anticancer peptide datasets demonstrate superior performance over vector-based, sequence language models, and existing image-based methods, achieving 86.8\% and 94.5\% accuracy on breast and lung cancer datasets, respectively. The topological representation preserves critical sequence information while enabling effective utilization of vision-based deep learning architectures for molecular sequence analysis.
Murmur2Vec: A Hashing Based Solution For Embedding Generation Of COVID-19 Spike Sequences
Dec 10, 2025Abstract:Early detection and characterization of coronavirus disease (COVID-19), caused by SARS-CoV-2, remain critical for effective clinical response and public-health planning. The global availability of large-scale viral sequence data presents significant opportunities for computational analysis; however, existing approaches face notable limitations. Phylogenetic tree-based methods are computationally intensive and do not scale efficiently to today's multi-million-sequence datasets. Similarly, current embedding-based techniques often rely on aligned sequences or exhibit suboptimal predictive performance and high runtime costs, creating barriers to practical large-scale analysis. In this study, we focus on the most prevalent SARS-CoV-2 lineages associated with the spike protein region and introduce a scalable embedding method that leverages hashing to generate compact, low-dimensional representations of spike sequences. These embeddings are subsequently used to train a variety of machine learning models for supervised lineage classification. We conduct an extensive evaluation comparing our approach with multiple baseline and state-of-the-art biological sequence embedding methods across diverse metrics. Our results demonstrate that the proposed embeddings offer substantial improvements in efficiency, achieving up to 86.4\% classification accuracy while reducing embedding generation time by as much as 99.81\%. This highlights the method's potential as a fast, effective, and scalable solution for large-scale viral sequence analysis.
Sequence Analysis Using the Bezier Curve
Mar 18, 2025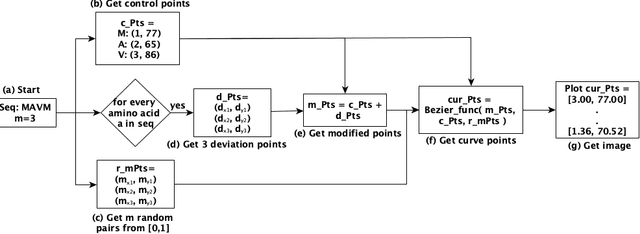
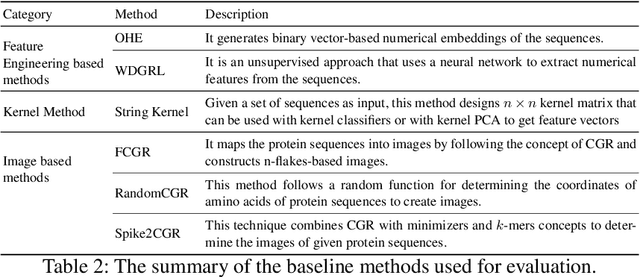
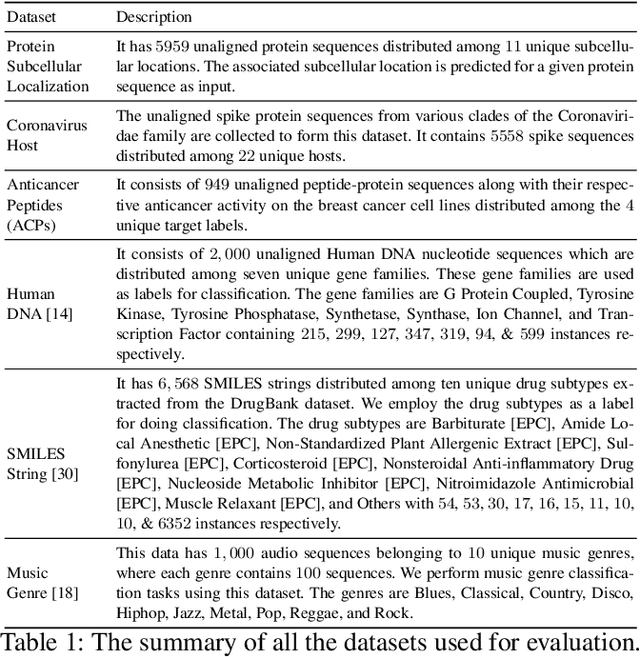

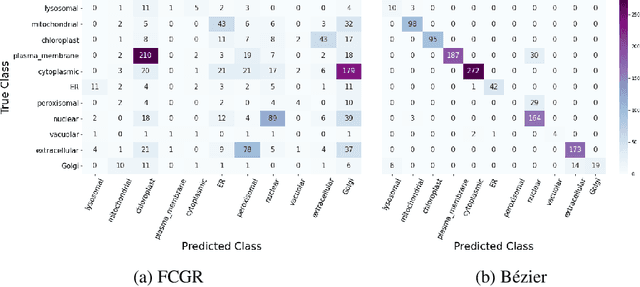
Abstract:The analysis of sequences (e.g., protein, DNA, and SMILES string) is essential for disease diagnosis, biomaterial engineering, genetic engineering, and drug discovery domains. Conventional analytical methods focus on transforming sequences into numerical representations for applying machine learning/deep learning-based sequence characterization. However, their efficacy is constrained by the intrinsic nature of deep learning (DL) models, which tend to exhibit suboptimal performance when applied to tabular data. An alternative group of methodologies endeavors to convert biological sequences into image forms by applying the concept of Chaos Game Representation (CGR). However, a noteworthy drawback of these methods lies in their tendency to map individual elements of the sequence onto a relatively small subset of designated pixels within the generated image. The resulting sparse image representation may not adequately encapsulate the comprehensive sequence information, potentially resulting in suboptimal predictions. In this study, we introduce a novel approach to transform sequences into images using the B\'ezier curve concept for element mapping. Mapping the elements onto a curve enhances the sequence information representation in the respective images, hence yielding better DL-based classification performance. We employed different sequence datasets to validate our system by using different classification tasks, and the results illustrate that our B\'ezier curve method is able to achieve good performance for all the tasks.
Hilbert Curve Based Molecular Sequence Analysis
Dec 29, 2024Abstract:Accurate molecular sequence analysis is a key task in the field of bioinformatics. To apply molecular sequence classification algorithms, we first need to generate the appropriate representations of the sequences. Traditional numeric sequence representation techniques are mostly based on sequence alignment that faces limitations in the form of lack of accuracy. Although several alignment-free techniques have also been introduced, their tabular data form results in low performance when used with Deep Learning (DL) models compared to the competitive performance observed in the case of image-based data. To find a solution to this problem and to make Deep Learning (DL) models function to their maximum potential while capturing the important spatial information in the sequence data, we propose a universal Hibert curve-based Chaos Game Representation (CGR) method. This method is a transformative function that involves a novel Alphabetic index mapping technique used in constructing Hilbert curve-based image representation from molecular sequences. Our method can be globally applied to any type of molecular sequence data. The Hilbert curve-based image representations can be used as input to sophisticated vision DL models for sequence classification. The proposed method shows promising results as it outperforms current state-of-the-art methods by achieving a high accuracy of $94.5$\% and an F1 score of $93.9\%$ when tested with the CNN model on the lung cancer dataset. This approach opens up a new horizon for exploring molecular sequence analysis using image classification methods.
Converting Time Series Data to Numeric Representations Using Alphabetic Mapping and k-mer strategy
Dec 29, 2024Abstract:In the realm of data analysis and bioinformatics, representing time series data in a manner akin to biological sequences offers a novel approach to leverage sequence analysis techniques. Transforming time series signals into molecular sequence-type representations allows us to enhance pattern recognition by applying sophisticated sequence analysis techniques (e.g. $k$-mers based representation) developed in bioinformatics, uncovering hidden patterns and relationships in complex, non-linear time series data. This paper proposes a method to transform time series signals into biological/molecular sequence-type representations using a unique alphabetic mapping technique. By generating 26 ranges corresponding to the 26 letters of the English alphabet, each value within the time series is mapped to a specific character based on its range. This conversion facilitates the application of sequence analysis algorithms, typically used in bioinformatics, to analyze time series data. We demonstrate the effectiveness of this approach by converting real-world time series signals into character sequences and performing sequence classification. The resulting sequences can be utilized for various sequence-based analysis techniques, offering a new perspective on time series data representation and analysis.
Computing Gram Matrix for SMILES Strings using RDKFingerprint and Sinkhorn-Knopp Algorithm
Dec 19, 2024Abstract:In molecular structure data, SMILES (Simplified Molecular Input Line Entry System) strings are used to analyze molecular structure design. Numerical feature representation of SMILES strings is a challenging task. This work proposes a kernel-based approach for encoding and analyzing molecular structures from SMILES strings. The proposed approach involves computing a kernel matrix using the Sinkhorn-Knopp algorithm while using kernel principal component analysis (PCA) for dimensionality reduction. The resulting low-dimensional embeddings are then used for classification and regression analysis. The kernel matrix is computed by converting the SMILES strings into molecular structures using the Morgan Fingerprint, which computes a fingerprint for each molecule. The distance matrix is computed using the pairwise kernels function. The Sinkhorn-Knopp algorithm is used to compute the final kernel matrix that satisfies the constraints of a probability distribution. This is achieved by iteratively adjusting the kernel matrix until the marginal distributions of the rows and columns match the desired marginal distributions. We provided a comprehensive empirical analysis of the proposed kernel method to evaluate its goodness with greater depth. The suggested method is assessed for drug subcategory prediction (classification task) and solubility AlogPS ``Aqueous solubility and Octanol/Water partition coefficient" (regression task) using the benchmark SMILES string dataset. The outcomes show the proposed method outperforms several baseline methods in terms of supervised analysis and has potential uses in molecular design and drug discovery. Overall, the suggested method is a promising avenue for kernel methods-based molecular structure analysis and design.
DWFL: Enhancing Federated Learning through Dynamic Weighted Averaging
Nov 07, 2024Abstract:Federated Learning (FL) is a distributed learning technique that maintains data privacy by providing a decentralized training method for machine learning models using distributed big data. This promising Federated Learning approach has also gained popularity in bioinformatics, where the privacy of biomedical data holds immense importance, especially when patient data is involved. Despite the successful implementation of Federated learning in biological sequence analysis, rigorous consideration is still required to improve accuracy in a way that data privacy should not be compromised. Additionally, the optimal integration of federated learning, especially in protein sequence analysis, has not been fully explored. We propose a deep feed-forward neural network-based enhanced federated learning method for protein sequence classification to overcome these challenges. Our method introduces novel enhancements to improve classification accuracy. We introduce dynamic weighted federated learning (DWFL) which is a federated learning-based approach, where local model weights are adjusted using weighted averaging based on their performance metrics. By assigning higher weights to well-performing models, we aim to create a more potent initial global model for the federated learning process, leading to improved accuracy. We conduct experiments using real-world protein sequence datasets to assess the effectiveness of DWFL. The results obtained using our proposed approach demonstrate significant improvements in model accuracy, making federated learning a preferred, more robust, and privacy-preserving approach for collaborative machine-learning tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge