Sadaf Khademi
AutoRad-Lung: A Radiomic-Guided Prompting Autoregressive Vision-Language Model for Lung Nodule Malignancy Prediction
Mar 26, 2025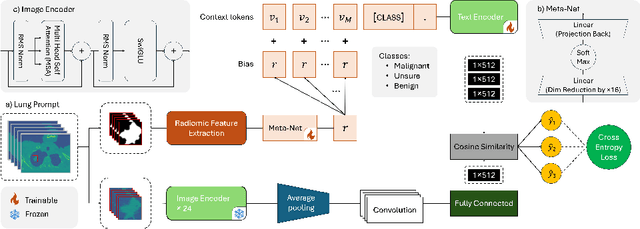
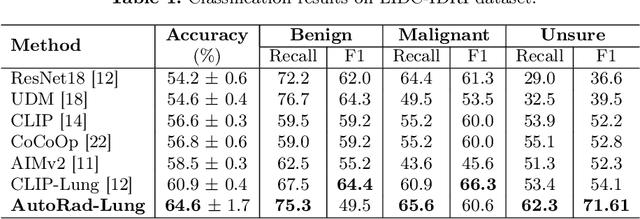
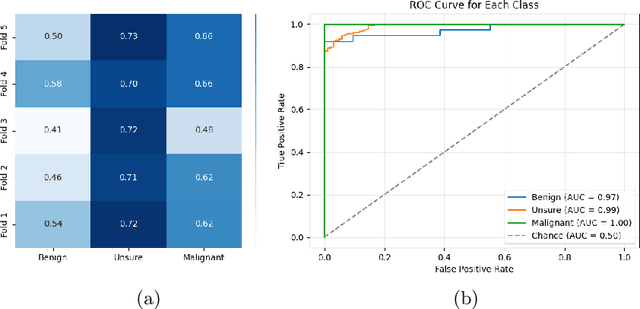
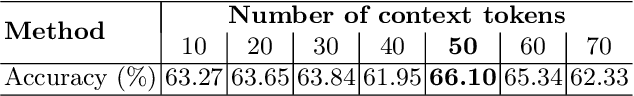
Abstract:Lung cancer remains one of the leading causes of cancer-related mortality worldwide. A crucial challenge for early diagnosis is differentiating uncertain cases with similar visual characteristics and closely annotation scores. In clinical practice, radiologists rely on quantitative, hand-crafted Radiomic features extracted from Computed Tomography (CT) images, while recent research has primarily focused on deep learning solutions. More recently, Vision-Language Models (VLMs), particularly Contrastive Language-Image Pre-Training (CLIP)-based models, have gained attention for their ability to integrate textual knowledge into lung cancer diagnosis. While CLIP-Lung models have shown promising results, we identified the following potential limitations: (a) dependence on radiologists' annotated attributes, which are inherently subjective and error-prone, (b) use of textual information only during training, limiting direct applicability at inference, and (c) Convolutional-based vision encoder with randomly initialized weights, which disregards prior knowledge. To address these limitations, we introduce AutoRad-Lung, which couples an autoregressively pre-trained VLM, with prompts generated from hand-crafted Radiomics. AutoRad-Lung uses the vision encoder of the Large-Scale Autoregressive Image Model (AIMv2), pre-trained using a multi-modal autoregressive objective. Given that lung tumors are typically small, irregularly shaped, and visually similar to healthy tissue, AutoRad-Lung offers significant advantages over its CLIP-based counterparts by capturing pixel-level differences. Additionally, we introduce conditional context optimization, which dynamically generates context-specific prompts based on input Radiomics, improving cross-modal alignment.
FH-TabNet: Multi-Class Familial Hypercholesterolemia Detection via a Multi-Stage Tabular Deep Learning
Mar 16, 2024Abstract:Familial Hypercholesterolemia (FH) is a genetic disorder characterized by elevated levels of Low-Density Lipoprotein (LDL) cholesterol or its associated genes. Early-stage and accurate categorization of FH is of significance allowing for timely interventions to mitigate the risk of life-threatening conditions. Conventional diagnosis approach, however, is complex, costly, and a challenging interpretation task even for experienced clinicians resulting in high underdiagnosis rates. Although there has been a recent surge of interest in using Machine Learning (ML) models for early FH detection, existing solutions only consider a binary classification task solely using classical ML models. Despite its significance, application of Deep Learning (DL) for FH detection is in its infancy, possibly, due to categorical nature of the underlying clinical data. The paper addresses this gap by introducing the FH-TabNet, which is a multi-stage tabular DL network for multi-class (Definite, Probable, Possible, and Unlikely) FH detection. The FH-TabNet initially involves applying a deep tabular data learning architecture (TabNet) for primary categorization into healthy (Possible/Unlikely) and patient (Probable/Definite) classes. Subsequently, independent TabNet classifiers are applied to each subgroup, enabling refined classification. The model's performance is evaluated through 5-fold cross-validation illustrating superior performance in categorizing FH patients, particularly in the challenging low-prevalence subcategories.
NYCTALE: Neuro-Evidence Transformer for Adaptive and Personalized Lung Nodule Invasiveness Prediction
Feb 15, 2024Abstract:Drawing inspiration from the primate brain's intriguing evidence accumulation process, and guided by models from cognitive psychology and neuroscience, the paper introduces the NYCTALE framework, a neuro-inspired and evidence accumulation-based Transformer architecture. The proposed neuro-inspired NYCTALE offers a novel pathway in the domain of Personalized Medicine (PM) for lung cancer diagnosis. In nature, Nyctales are small owls known for their nocturnal behavior, hunting primarily during the darkness of night. The NYCTALE operates in a similarly vigilant manner, i.e., processing data in an evidence-based fashion and making predictions dynamically/adaptively. Distinct from conventional Computed Tomography (CT)-based Deep Learning (DL) models, the NYCTALE performs predictions only when sufficient amount of evidence is accumulated. In other words, instead of processing all or a pre-defined subset of CT slices, for each person, slices are provided one at a time. The NYCTALE framework then computes an evidence vector associated with contribution of each new CT image. A decision is made once the total accumulated evidence surpasses a specific threshold. Preliminary experimental analyses conducted using a challenging in-house dataset comprising 114 subjects. The results are noteworthy, suggesting that NYCTALE outperforms the benchmark accuracy even with approximately 60% less training data on this demanding and small dataset.
Spatio-Temporal Hybrid Fusion of CAE and SWIn Transformers for Lung Cancer Malignancy Prediction
Oct 27, 2022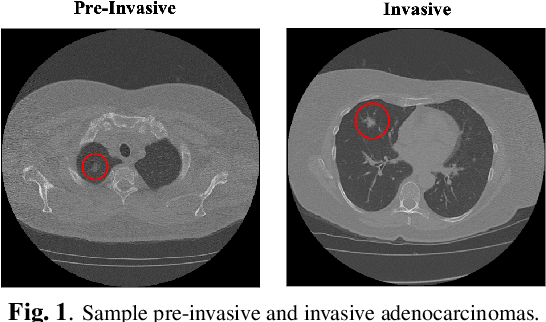
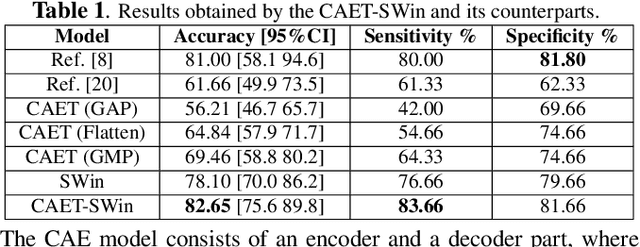
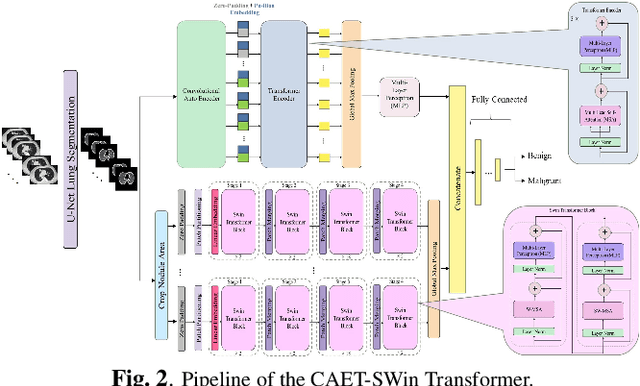
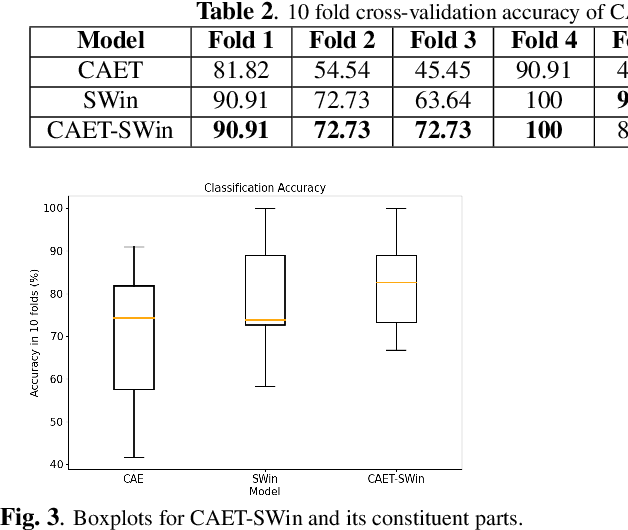
Abstract:The paper proposes a novel hybrid discovery Radiomics framework that simultaneously integrates temporal and spatial features extracted from non-thin chest Computed Tomography (CT) slices to predict Lung Adenocarcinoma (LUAC) malignancy with minimum expert involvement. Lung cancer is the leading cause of mortality from cancer worldwide and has various histologic types, among which LUAC has recently been the most prevalent. LUACs are classified as pre-invasive, minimally invasive, and invasive adenocarcinomas. Timely and accurate knowledge of the lung nodules malignancy leads to a proper treatment plan and reduces the risk of unnecessary or late surgeries. Currently, chest CT scan is the primary imaging modality to assess and predict the invasiveness of LUACs. However, the radiologists' analysis based on CT images is subjective and suffers from a low accuracy compared to the ground truth pathological reviews provided after surgical resections. The proposed hybrid framework, referred to as the CAET-SWin, consists of two parallel paths: (i) The Convolutional Auto-Encoder (CAE) Transformer path that extracts and captures informative features related to inter-slice relations via a modified Transformer architecture, and; (ii) The Shifted Window (SWin) Transformer path, which is a hierarchical vision transformer that extracts nodules' related spatial features from a volumetric CT scan. Extracted temporal (from the CAET-path) and spatial (from the Swin path) are then fused through a fusion path to classify LUACs. Experimental results on our in-house dataset of 114 pathologically proven Sub-Solid Nodules (SSNs) demonstrate that the CAET-SWin significantly improves reliability of the invasiveness prediction task while achieving an accuracy of 82.65%, sensitivity of 83.66%, and specificity of 81.66% using 10-fold cross-validation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge