Ross T. Whitaker
Learning Deep Features for Shape Correspondence with Domain Invariance
Feb 21, 2021
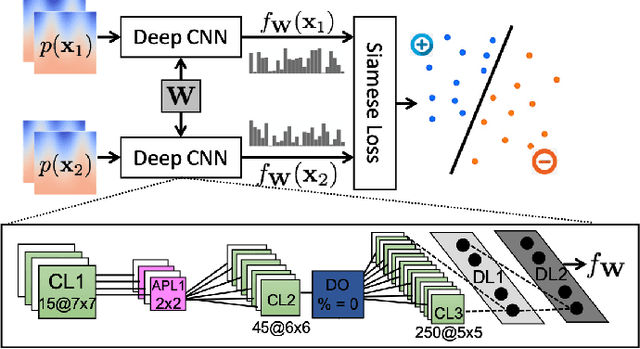
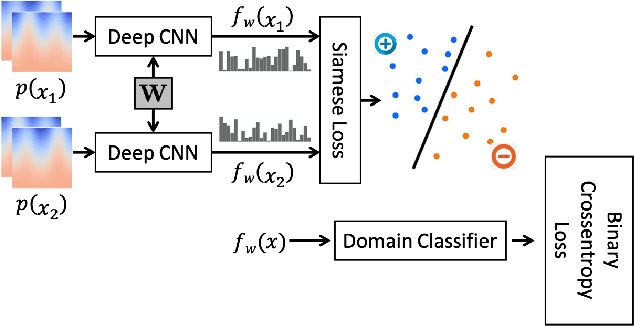
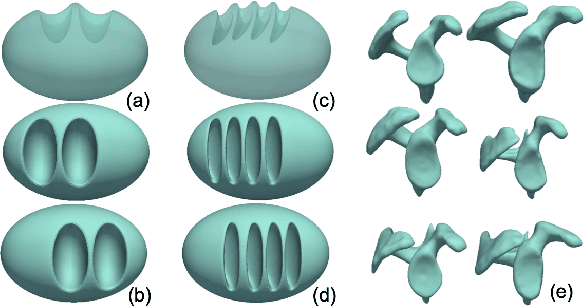
Abstract:Correspondence-based shape models are key to various medical imaging applications that rely on a statistical analysis of anatomies. Such shape models are expected to represent consistent anatomical features across the population for population-specific shape statistics. Early approaches for correspondence placement rely on nearest neighbor search for simpler anatomies. Coordinate transformations for shape correspondence hold promise to address the increasing anatomical complexities. Nonetheless, due to the inherent shape-level geometric complexity and population-level shape variation, the coordinate-wise correspondence often does not translate to the anatomical correspondence. An alternative, group-wise approach for correspondence placement explicitly models the trade-off between geometric description and the population's statistical compactness. However, these models achieve limited success in resolving nonlinear shape correspondence. Recent works have addressed this limitation by adopting an application-specific notion of correspondence through lifting positional data to a higher dimensional feature space. However, they heavily rely on manual expertise to create domain-specific features and consistent landmarks. This paper proposes an automated feature learning approach, using deep convolutional neural networks to extract correspondence-friendly features from shape ensembles. Further, an unsupervised domain adaptation scheme is introduced to augment the pretrained geometric features with new anatomies. Results on anatomical datasets of human scapula, femur, and pelvis bones demonstrate that features learned in supervised fashion show improved performance for correspondence estimation compared to the manual features. Further, unsupervised learning is demonstrated to learn complex anatomy features using the supervised domain adaptation from features learned on simpler anatomy.
GENs: Generative Encoding Networks
Oct 28, 2020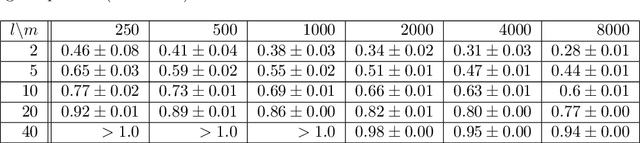
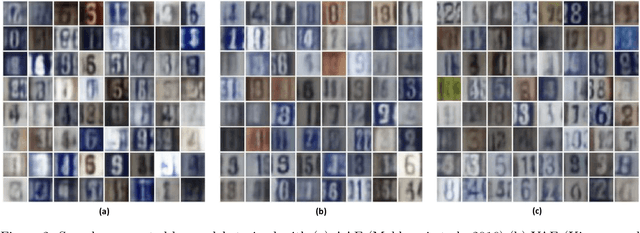

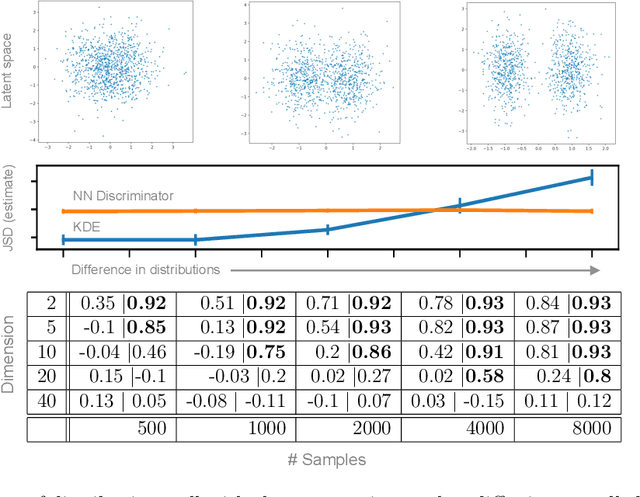

Abstract:Mapping data from and/or onto a known family of distributions has become an important topic in machine learning and data analysis. Deep generative models (e.g., generative adversarial networks ) have been used effectively to match known and unknown distributions. Nonetheless, when the form of the target distribution is known, analytical methods are advantageous in providing robust results with provable properties. In this paper, we propose and analyze the use of nonparametric density methods to estimate the Jensen-Shannon divergence for matching unknown data distributions to known target distributions, such Gaussian or mixtures of Gaussians, in latent spaces. This analytical method has several advantages: better behavior when training sample quantity is low, provable convergence properties, and relatively few parameters, which can be derived analytically. Using the proposed method, we enforce the latent representation of an autoencoder to match a target distribution in a learning framework that we call a {\em generative encoding network}. Here, we present the numerical methods; derive the expected distribution of the data in the latent space; evaluate the properties of the latent space, sample reconstruction, and generated samples; show the advantages over the adversarial counterpart; and demonstrate the application of the method in real world.
Unsupervised Shape Normality Metric for Severity Quantification
Jul 18, 2020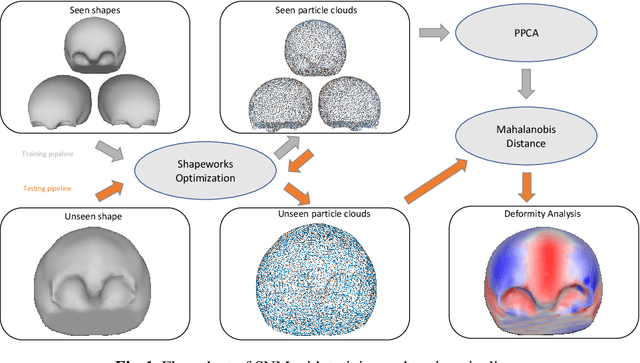
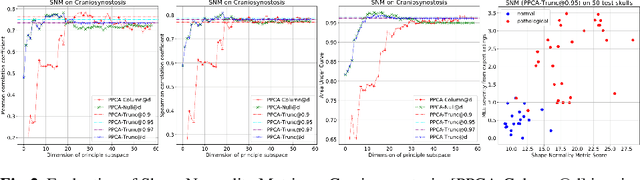

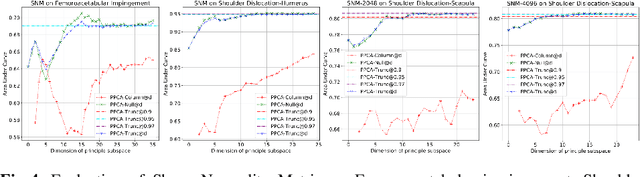
Abstract:This work describes an unsupervised method to objectively quantify the abnormality of general anatomical shapes. The severity of an anatomical deformity often serves as a determinant in the clinical management of patients. However, experiential bias and distinctive random residuals among specialist individuals bring variability in diagnosis and patient management decisions, irrespective of the objective deformity degree. Therefore, supervised methods are prone to be misled given insufficient labeling of pathological samples that inevitably preserve human bias and inconsistency. Furthermore, subjects demonstrating a specific pathology are naturally rare relative to the normal population. To avoid relying on sufficient pathological samples by fully utilizing the power of normal samples, we propose the shape normality metric (SNM), which requires learning only from normal samples and zero knowledge about the pathology. We represent shapes by landmarks automatically inferred from the data and model the normal group by a multivariate Gaussian distribution. Extensive experiments on different anatomical datasets, including skulls, femurs, scapulae, and humeri, demonstrate that SNM can provide an effective normality measurement, which can significantly detect and indicate pathology. Therefore, SNM offers promising value in a variety of clinical applications.
A Cooperative Autoencoder for Population-Based Regularization of CNN Image Registration
Aug 19, 2019

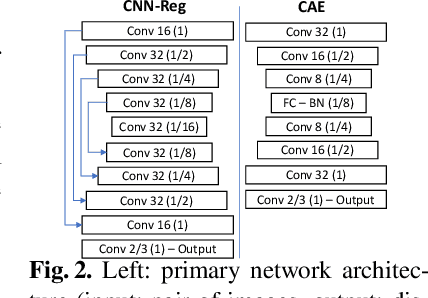
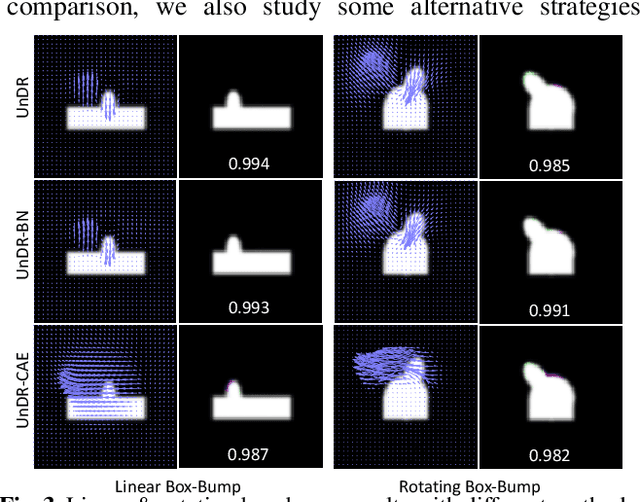
Abstract:Spatial transformations are enablers in a variety of medical image analysis applications that entail aligning images to a common coordinate systems. Population analysis of such transformations is expected to capture the underlying image and shape variations, and hence these transformations are required to produce anatomically feasible correspondences. This is usually enforced through some smoothness-based generic regularization on deformation field. Alternatively, population-based regularization has been shown to produce anatomically accurate correspondences in cases where anatomically unaware (i.e., data independent) fail. Recently, deep networks have been for unsupervised image registration, these methods are computationally faster and maintains the accuracy of state of the art methods. However, these networks use smoothness penalty on deformation fields and ignores population-level statistics of the transformations. We propose a novel neural network architecture that simultaneously learns and uses the population-level statistics of the spatial transformations to regularize the neural networks for unsupervised image registration. This regularization is in the form of a bottleneck autoencoder, which encodes the population level information of the deformation fields in a low-dimensional manifold. The proposed architecture produces deformation fields that describe the population-level features and associated correspondences in an anatomically relevant manner and are statistically compact relative to the state-of-the-art approaches while maintaining computational efficiency. We demonstrate the efficacy of the proposed architecture on synthetic data sets, as well as 2D and 3D medical data.
DeepSSM: A Deep Learning Framework for Statistical Shape Modeling from Raw Images
Sep 28, 2018



Abstract:Statistical shape modeling is an important tool to characterize variation in anatomical morphology. Typical shapes of interest are measured using 3D imaging and a subsequent pipeline of registration, segmentation, and some extraction of shape features or projections onto some lower-dimensional shape space, which facilitates subsequent statistical analysis. Many methods for constructing compact shape representations have been proposed, but are often impractical due to the sequence of image preprocessing operations, which involve significant parameter tuning, manual delineation, and/or quality control by the users. We propose DeepSSM: a deep learning approach to extract a low-dimensional shape representation directly from 3D images, requiring virtually no parameter tuning or user assistance. DeepSSM uses a convolutional neural network (CNN) that simultaneously localizes the biological structure of interest, establishes correspondences, and projects these points onto a low-dimensional shape representation in the form of PCA loadings within a point distribution model. To overcome the challenge of the limited availability of training images, we present a novel data augmentation procedure that uses existing correspondences on a relatively small set of processed images with shape statistics to create plausible training samples with known shape parameters. Hence, we leverage the limited CT/MRI scans (40-50) into thousands of images needed to train a CNN. After the training, the CNN automatically produces accurate low-dimensional shape representations for unseen images. We validate DeepSSM for three different applications pertaining to modeling pediatric cranial CT for characterization of metopic craniosynostosis, femur CT scans identifying morphologic deformities of the hip due to femoroacetabular impingement, and left atrium MRI scans for atrial fibrillation recurrence prediction.
Clustering With Pairwise Relationships: A Generative Approach
May 06, 2018
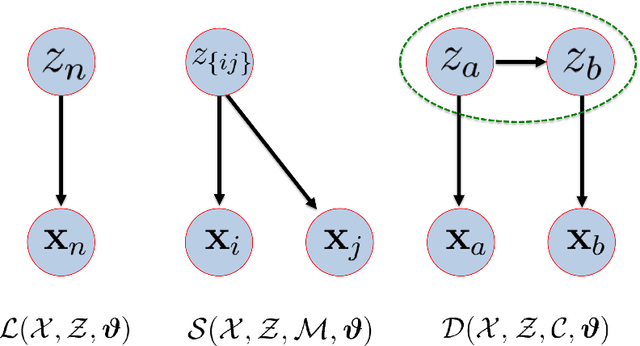
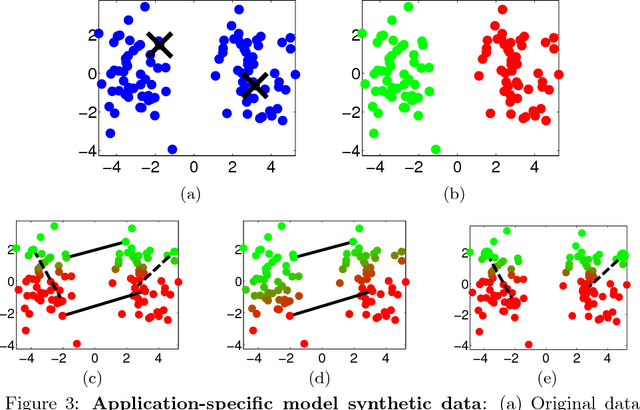

Abstract:Semi-supervised learning (SSL) has become important in current data analysis applications, where the amount of unlabeled data is growing exponentially and user input remains limited by logistics and expense. Constrained clustering, as a subclass of SSL, makes use of user input in the form of relationships between data points (e.g., pairs of data points belonging to the same class or different classes) and can remarkably improve the performance of unsupervised clustering in order to reflect user-defined knowledge of the relationships between particular data points. Existing algorithms incorporate such user input, heuristically, as either hard constraints or soft penalties, which are separate from any generative or statistical aspect of the clustering model; this results in formulations that are suboptimal and not sufficiently general. In this paper, we propose a principled, generative approach to probabilistically model, without ad hoc penalties, the joint distribution given by user-defined pairwise relations. The proposed model accounts for general underlying distributions without assuming a specific form and relies on expectation-maximization for model fitting. For distributions in a standard form, the proposed approach results in a closed-form solution for updated parameters.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge