Rob Brennan
Retrieval-aligned Tabular Foundation Models Enable Robust Clinical Risk Prediction in Electronic Health Records Under Real-world Constraints
Apr 02, 2026Abstract:Clinical prediction from structured electronic health records (EHRs) is challenging due to high dimensionality, heterogeneity, class imbalance, and distribution shift. While tabular in-context learning (TICL) and retrieval-augmented methods perform well on generic benchmarks, their behavior in clinical settings remains unclear. We present a multi-cohort EHR benchmark comparing classical, deep tabular, and TICL models across varying data scale, feature dimensionality, outcome rarity, and cross-cohort generalization. PFN-based TICL models are sample-efficient in low-data regimes but degrade under naive distance-based retrieval as heterogeneity and imbalance increase. We propose AWARE, a task-aligned retrieval framework using supervised embedding learning and lightweight adapters. AWARE improves AUPRC by up to 12.2% under extreme imbalance, with gains increasing with data complexity. Our results identify retrieval quality and retrieval-inference alignment as key bottlenecks for deploying tabular in-context learning in clinical prediction.
KGroups: A Versatile Univariate Max-Relevance Min-Redundancy Feature Selection Algorithm for High-dimensional Biological Data
Mar 30, 2026Abstract:This paper proposes a new univariate filter feature selection (FFS) algorithm called KGroups. The majority of work in the literature focuses on investigating the relevance or redundancy estimations of feature selection (FS) methods. This has shown promising results and a real improvement of FFS methods' predictive performance. However, limited efforts have been made to investigate alternative FFS algorithms. This raises the following question: how much of the FFS methods' predictive performance depends on the selection algorithm rather than the relevance or the redundancy estimations? The majority of FFS methods fall into two categories: relevance maximisation (Max-Rel, also known as KBest) or simultaneous relevance maximisation and redundancy minimisation (mRMR). KBest is a univariate FFS algorithm that employs sorting (descending) for selection. mRMR is a multivariate FFS algorithm that employs an incremental search algorithm for selection. In this paper, we propose a new univariate mRMR called KGroups that employs clustering for selection. Extensive experiments on 14 high-dimensional biological benchmark datasets showed that KGroups achieves similar predictive performance compared to multivariate mRMR while being up to 821 times faster. KGroups is parameterisable, which leaves room for further predictive performance improvement through hyperparameter finetuning, unlike mRMR and KBest. KGroups outperforms KBest.
Explainable AI for Infection Prevention and Control: Modeling CPE Acquisition and Patient Outcomes in an Irish Hospital with Transformers
Sep 18, 2025Abstract:Carbapenemase-Producing Enterobacteriace poses a critical concern for infection prevention and control in hospitals. However, predictive modeling of previously highlighted CPE-associated risks such as readmission, mortality, and extended length of stay (LOS) remains underexplored, particularly with modern deep learning approaches. This study introduces an eXplainable AI modeling framework to investigate CPE impact on patient outcomes from Electronic Medical Records data of an Irish hospital. We analyzed an inpatient dataset from an Irish acute hospital, incorporating diagnostic codes, ward transitions, patient demographics, infection-related variables and contact network features. Several Transformer-based architectures were benchmarked alongside traditional machine learning models. Clinical outcomes were predicted, and XAI techniques were applied to interpret model decisions. Our framework successfully demonstrated the utility of Transformer-based models, with TabTransformer consistently outperforming baselines across multiple clinical prediction tasks, especially for CPE acquisition (AUROC and sensitivity). We found infection-related features, including historical hospital exposure, admission context, and network centrality measures, to be highly influential in predicting patient outcomes and CPE acquisition risk. Explainability analyses revealed that features like "Area of Residence", "Admission Ward" and prior admissions are key risk factors. Network variables like "Ward PageRank" also ranked highly, reflecting the potential value of structural exposure information. This study presents a robust and explainable AI framework for analyzing complex EMR data to identify key risk factors and predict CPE-related outcomes. Our findings underscore the superior performance of the Transformer models and highlight the importance of diverse clinical and network features.
Personalization of Dataset Retrieval Results using a Metadata-based Data Valuation Method
Jul 22, 2024Abstract:In this paper, we propose a novel data valuation method for a Dataset Retrieval (DR) use case in Ireland's National mapping agency. To the best of our knowledge, data valuation has not yet been applied to Dataset Retrieval. By leveraging metadata and a user's preferences, we estimate the personal value of each dataset to facilitate dataset retrieval and filtering. We then validated the data value-based ranking against the stakeholders' ranking of the datasets. The proposed data valuation method and use case demonstrated that data valuation is promising for dataset retrieval. For instance, the outperforming dataset retrieval based on our approach obtained 0.8207 in terms of NDCG@5 (the truncated Normalized Discounted Cumulative Gain at 5). This study is unique in its exploration of a data valuation-based approach to dataset retrieval and stands out because, unlike most existing methods, our approach is validated using the stakeholders ranking of the datasets.
Advanced Data Augmentation Approaches: A Comprehensive Survey and Future directions
Jan 14, 2023



Abstract:Deep learning (DL) algorithms have shown significant performance in various computer vision tasks. However, having limited labelled data lead to a network overfitting problem, where network performance is bad on unseen data as compared to training data. Consequently, it limits performance improvement. To cope with this problem, various techniques have been proposed such as dropout, normalization and advanced data augmentation. Among these, data augmentation, which aims to enlarge the dataset size by including sample diversity, has been a hot topic in recent times. In this article, we focus on advanced data augmentation techniques. we provide a background of data augmentation, a novel and comprehensive taxonomy of reviewed data augmentation techniques, and the strengths and weaknesses (wherever possible) of each technique. We also provide comprehensive results of the data augmentation effect on three popular computer vision tasks, such as image classification, object detection and semantic segmentation. For results reproducibility, we compiled available codes of all data augmentation techniques. Finally, we discuss the challenges and difficulties, and possible future direction for the research community. We believe, this survey provides several benefits i) readers will understand the data augmentation working mechanism to fix overfitting problems ii) results will save the searching time of the researcher for comparison purposes. iii) Codes of the mentioned data augmentation techniques are available at https://github.com/kmr2017/Advanced-Data-augmentation-codes iv) Future work will spark interest in research community.
Random Data Augmentation based Enhancement: A Generalized Enhancement Approach for Medical Datasets
Oct 03, 2022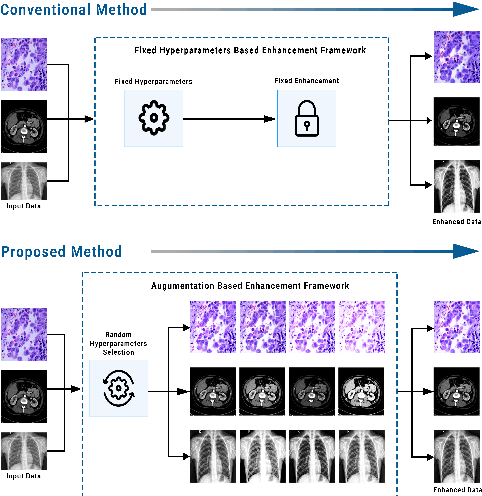
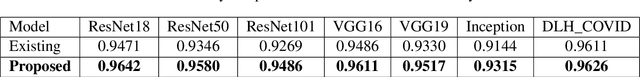
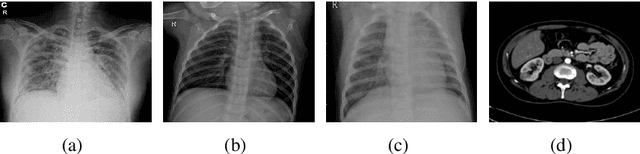
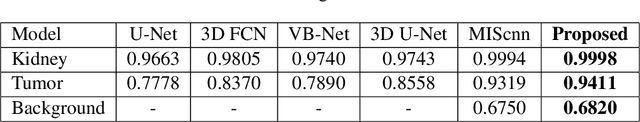
Abstract:Over the years, the paradigm of medical image analysis has shifted from manual expertise to automated systems, often using deep learning (DL) systems. The performance of deep learning algorithms is highly dependent on data quality. Particularly for the medical domain, it is an important aspect as medical data is very sensitive to quality and poor quality can lead to misdiagnosis. To improve the diagnostic performance, research has been done both in complex DL architectures and in improving data quality using dataset dependent static hyperparameters. However, the performance is still constrained due to data quality and overfitting of hyperparameters to a specific dataset. To overcome these issues, this paper proposes random data augmentation based enhancement. The main objective is to develop a generalized, data-independent and computationally efficient enhancement approach to improve medical data quality for DL. The quality is enhanced by improving the brightness and contrast of images. In contrast to the existing methods, our method generates enhancement hyperparameters randomly within a defined range, which makes it robust and prevents overfitting to a specific dataset. To evaluate the generalization of the proposed method, we use four medical datasets and compare its performance with state-of-the-art methods for both classification and segmentation tasks. For grayscale imagery, experiments have been performed with: COVID-19 chest X-ray, KiTS19, and for RGB imagery with: LC25000 datasets. Experimental results demonstrate that with the proposed enhancement methodology, DL architectures outperform other existing methods. Our code is publicly available at: https://github.com/aleemsidra/Augmentation-Based-Generalized-Enhancement
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge