R. Nick Bryan
Deep Learning Based Detection of Enlarged Perivascular Spaces on Brain MRI
Sep 27, 2022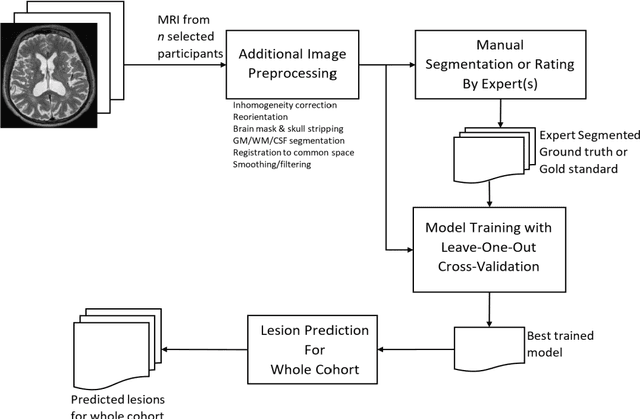
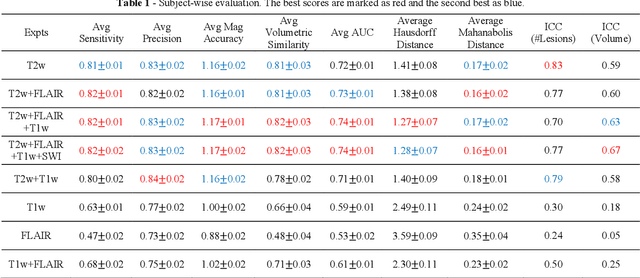
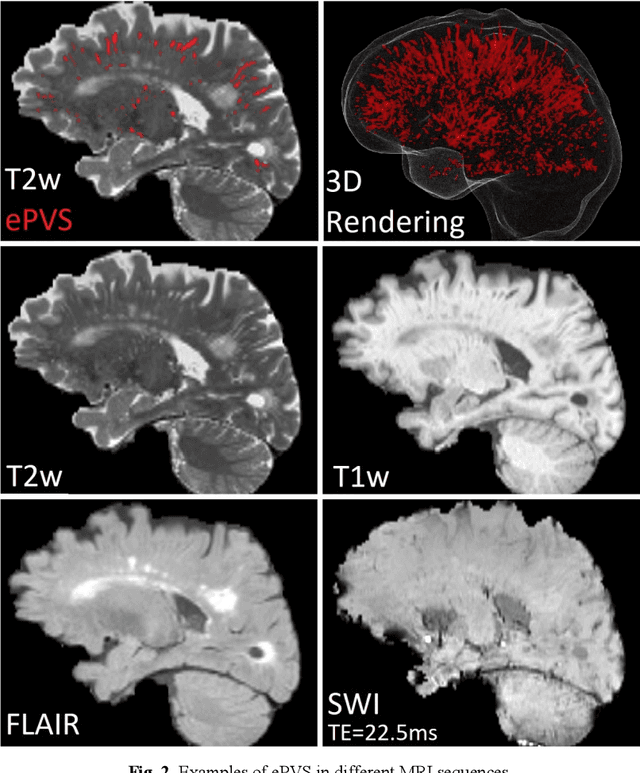
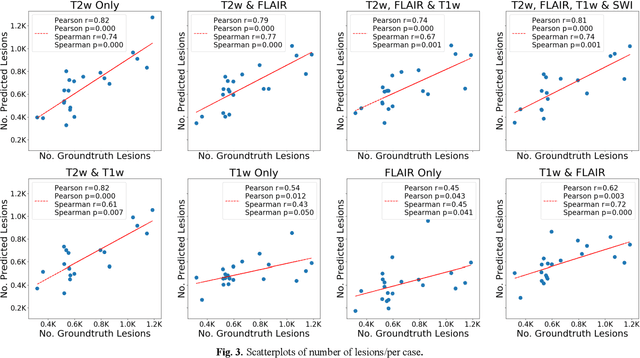
Abstract:Deep learning has been demonstrated effective in many neuroimaging applications. However, in many scenarios the number of imaging sequences capturing information related to small vessel disease lesions is insufficient to support data-driven techniques. Additionally, cohort-based studies may not always have the optimal or essential imaging sequences for accurate lesion detection. Therefore, it is necessary to determine which of these imaging sequences are essential for accurate detection. In this study we aimed to find the optimal combination of magnetic resonance imaging (MRI) sequences for deep learning-based detection of enlarged perivascular spaces (ePVS). To this end, we implemented an effective light-weight U-Net adapted for ePVS detection and comprehensively investigated different combinations of information from susceptibility weighted imaging (SWI), fluid-attenuated inversion recovery (FLAIR), T1-weighted (T1w) and T2-weighted (T2w) MRI sequences. We conclude that T2w MRI is the most important for accurate ePVS detection, and the incorporation of SWI, FLAIR and T1w MRI in the deep neural network could make insignificant improvements in accuracy.
Making Document-Level Information Extraction Right for the Right Reasons
Oct 14, 2021

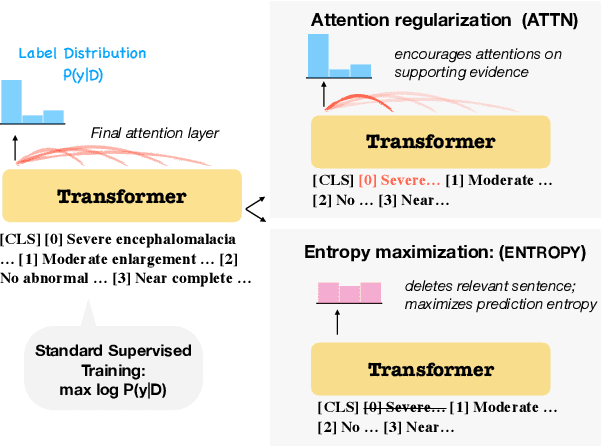
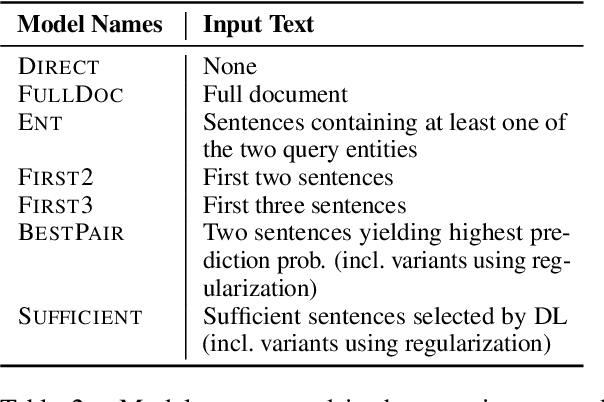
Abstract:Document-level information extraction is a flexible framework compatible with applications where information is not necessarily localized in a single sentence. For example, key features of a diagnosis in radiology a report may not be explicitly stated, but nevertheless can be inferred from the report's text. However, document-level neural models can easily learn spurious correlations from irrelevant information. This work studies how to ensure that these models make correct inferences from complex text and make those inferences in an auditable way: beyond just being right, are these models "right for the right reasons?" We experiment with post-hoc evidence extraction in a predict-select-verify framework using feature attribution techniques. While this basic approach can extract reasonable evidence, it can be regularized with small amounts of evidence supervision during training, which substantially improves the quality of extracted evidence. We evaluate on two domains: a small-scale labeled dataset of brain MRI reports and a large-scale modified version of DocRED (Yao et al., 2019) and show that models' plausibility can be improved with no loss in accuracy.
Medical Image Harmonization Using Deep Learning Based Canonical Mapping: Toward Robust and Generalizable Learning in Imaging
Oct 11, 2020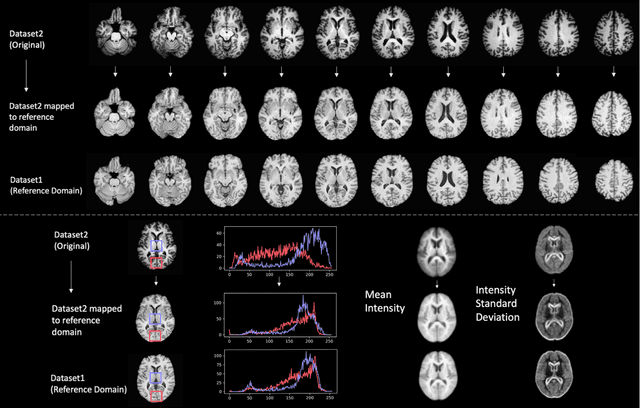
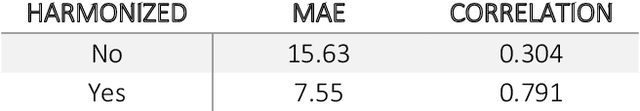
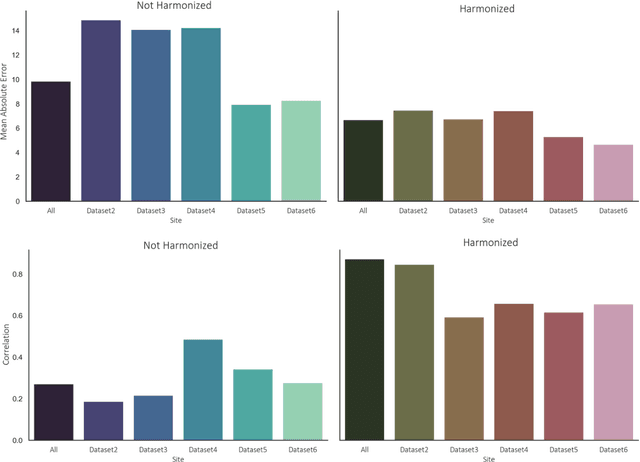
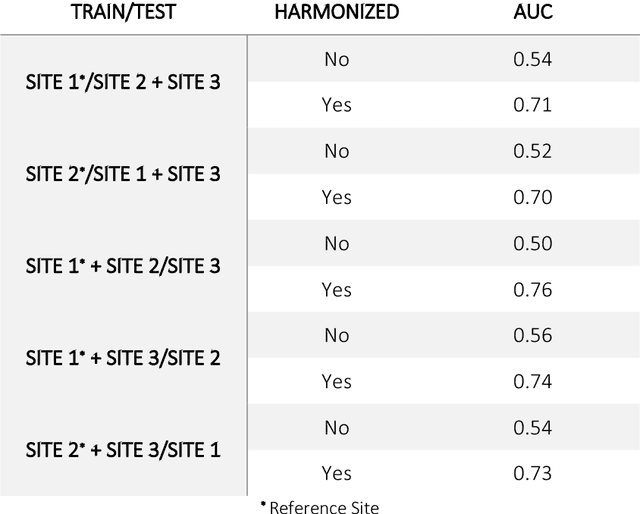
Abstract:Conventional and deep learning-based methods have shown great potential in the medical imaging domain, as means for deriving diagnostic, prognostic, and predictive biomarkers, and by contributing to precision medicine. However, these methods have yet to see widespread clinical adoption, in part due to limited generalization performance across various imaging devices, acquisition protocols, and patient populations. In this work, we propose a new paradigm in which data from a diverse range of acquisition conditions are "harmonized" to a common reference domain, where accurate model learning and prediction can take place. By learning an unsupervised image to image canonical mapping from diverse datasets to a reference domain using generative deep learning models, we aim to reduce confounding data variation while preserving semantic information, thereby rendering the learning task easier in the reference domain. We test this approach on two example problems, namely MRI-based brain age prediction and classification of schizophrenia, leveraging pooled cohorts of neuroimaging MRI data spanning 9 sites and 9701 subjects. Our results indicate a substantial improvement in these tasks in out-of-sample data, even when training is restricted to a single site.
DEEPMIR: A DEEP convolutional neural network for differential detection of cerebral Microbleeds and IRon deposits in MRI
Sep 30, 2020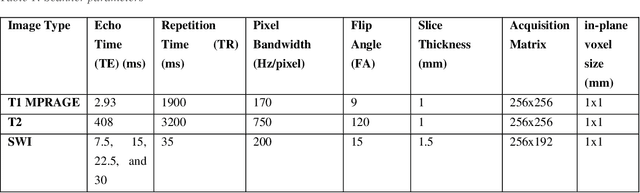
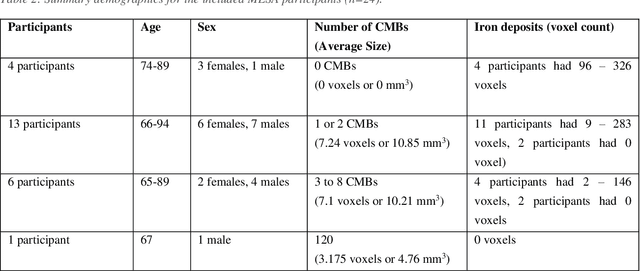
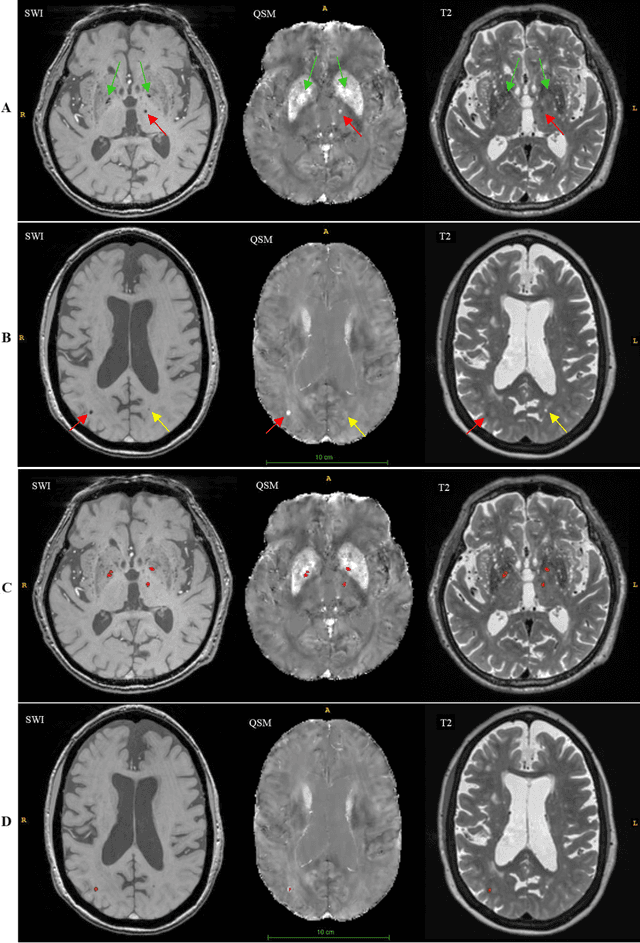

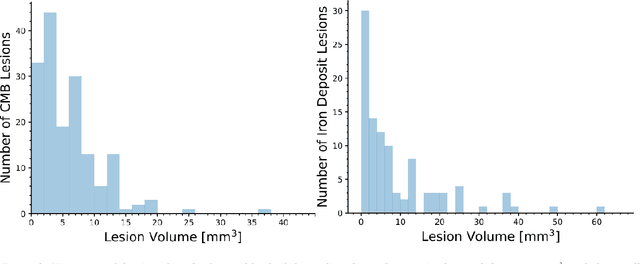
Abstract:Background: Cerebral microbleeds (CMBs) and non-hemorrhage iron deposits in the basal ganglia have been associated with brain aging, vascular disease and neurodegenerative disorders. Recent advances using quantitative susceptibility mapping (QSM) make it possible to differentiate iron content from mineralization in-vivo using magnetic resonance imaging (MRI). However, automated detection of such lesions is still challenging, making quantification in large cohort bases studies rather limited. Purpose: Development of a fully automated method using deep learning for detecting CMBs and basal ganglia iron deposits using multimodal MRI. Materials and Methods: We included a convenience sample of 24 participants from the MESA cohort and used T2-weighted images, susceptibility weighted imaging (SWI), and QSM to segment the lesions. We developed a protocol for simultaneous manual annotation of CMBs and non-hemorrhage iron deposits in the basal ganglia, which resulted in defining the gold standard. This gold standard was then used to train a deep convolution neural network (CNN) model. Specifically, we adapted the U-Net model with a higher number of resolution layers to be able to detect small lesions such as CMBs from standard resolution MRI which are used in cohort-based studies. The detection performance was then evaluated using the cross-validation principle in order to ensure generalization of the results. Results: With multi-class CNN models, we achieved an average sensitivity and precision of about 0.8 and 0.6, respectively for detecting CMBs. The same framework detected non-hemorrhage iron deposits reaching an average sensitivity and precision of about 0.8. Conclusions: Our results showed that deep learning could automate the detection of small vessel disease lesions and including multimodal MR data such as QSM can improve the detection of CMB and non-hemorrhage iron deposits.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge