Paul Fischer
CUTE-MRI: Conformalized Uncertainty-based framework for Time-adaptivE MRI
Aug 20, 2025Abstract:Magnetic Resonance Imaging (MRI) offers unparalleled soft-tissue contrast but is fundamentally limited by long acquisition times. While deep learning-based accelerated MRI can dramatically shorten scan times, the reconstruction from undersampled data introduces ambiguity resulting from an ill-posed problem with infinitely many possible solutions that propagates to downstream clinical tasks. This uncertainty is usually ignored during the acquisition process as acceleration factors are often fixed a priori, resulting in scans that are either unnecessarily long or of insufficient quality for a given clinical endpoint. This work introduces a dynamic, uncertainty-aware acquisition framework that adjusts scan time on a per-subject basis. Our method leverages a probabilistic reconstruction model to estimate image uncertainty, which is then propagated through a full analysis pipeline to a quantitative metric of interest (e.g., patellar cartilage volume or cardiac ejection fraction). We use conformal prediction to transform this uncertainty into a rigorous, calibrated confidence interval for the metric. During acquisition, the system iteratively samples k-space, updates the reconstruction, and evaluates the confidence interval. The scan terminates automatically once the uncertainty meets a user-predefined precision target. We validate our framework on both knee and cardiac MRI datasets. Our results demonstrate that this adaptive approach reduces scan times compared to fixed protocols while providing formal statistical guarantees on the precision of the final image. This framework moves beyond fixed acceleration factors, enabling patient-specific acquisitions that balance scan efficiency with diagnostic confidence, a critical step towards personalized and resource-efficient MRI.
Optimal Lattice Boltzmann Closures through Multi-Agent Reinforcement Learning
Apr 19, 2025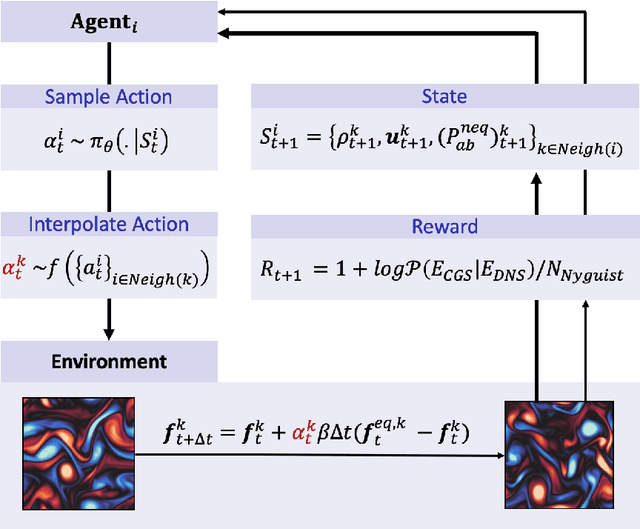
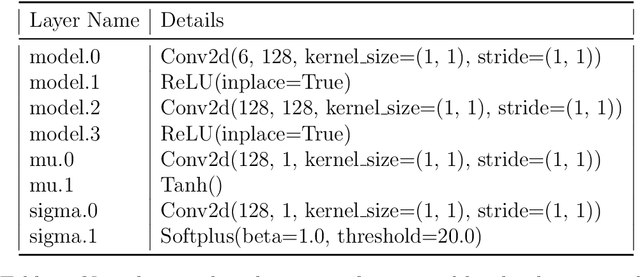
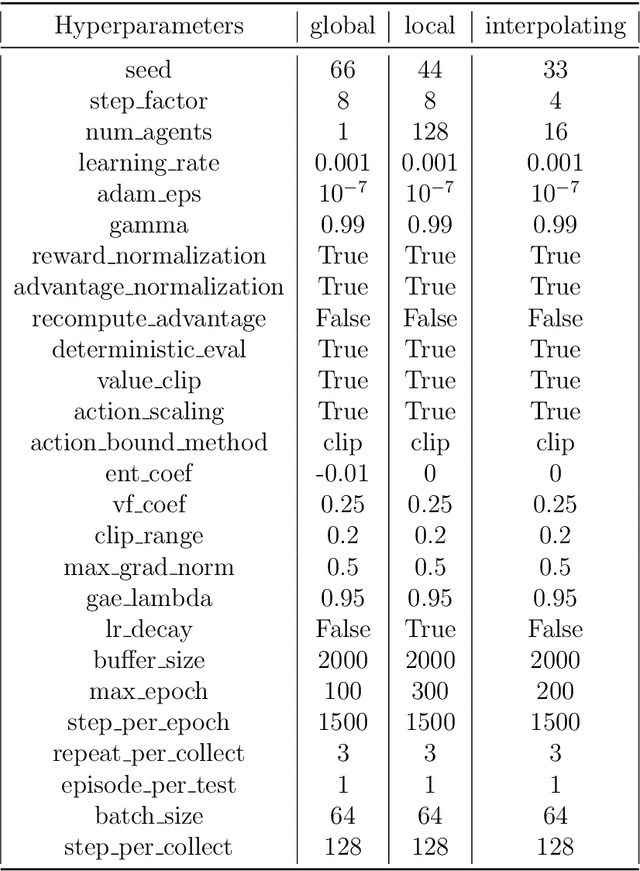
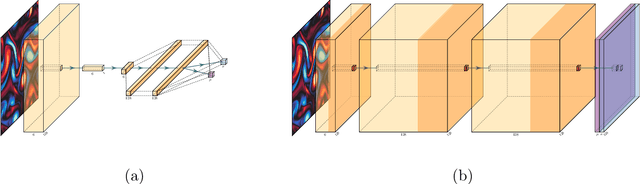

Abstract:The Lattice Boltzmann method (LBM) offers a powerful and versatile approach to simulating diverse hydrodynamic phenomena, spanning microfluidics to aerodynamics. The vast range of spatiotemporal scales inherent in these systems currently renders full resolution impractical, necessitating the development of effective closure models for under-resolved simulations. Under-resolved LBMs are unstable, and while there is a number of important efforts to stabilize them, they often face limitations in generalizing across scales and physical systems. We present a novel, data-driven, multiagent reinforcement learning (MARL) approach that drastically improves stability and accuracy of coarse-grained LBM simulations. The proposed method uses a convolutional neural network to dynamically control the local relaxation parameter for the LB across the simulation grid. The LB-MARL framework is showcased in turbulent Kolmogorov flows. We find that the MARL closures stabilize the simulations and recover the energy spectra of significantly more expensive fully resolved simulations while maintaining computational efficiency. The learned closure model can be transferred to flow scenarios unseen during training and has improved robustness and spectral accuracy compared to traditional LBM models. We believe that MARL closures open new frontiers for efficient and accurate simulations of a multitude of complex problems not accessible to present-day LB methods alone.
Conformal Performance Range Prediction for Segmentation Output Quality Control
Jul 18, 2024



Abstract:Recent works have introduced methods to estimate segmentation performance without ground truth, relying solely on neural network softmax outputs. These techniques hold potential for intuitive output quality control. However, such performance estimates rely on calibrated softmax outputs, which is often not the case in modern neural networks. Moreover, the estimates do not take into account inherent uncertainty in segmentation tasks. These limitations may render precise performance predictions unattainable, restricting the practical applicability of performance estimation methods. To address these challenges, we develop a novel approach for predicting performance ranges with statistical guarantees of containing the ground truth with a user specified probability. Our method leverages sampling-based segmentation uncertainty estimation to derive heuristic performance ranges, and applies split conformal prediction to transform these estimates into rigorous prediction ranges that meet the desired guarantees. We demonstrate our approach on the FIVES retinal vessel segmentation dataset and compare five commonly used sampling-based uncertainty estimation techniques. Our results show that it is possible to achieve the desired coverage with small prediction ranges, highlighting the potential of performance range prediction as a valuable tool for output quality control.
PULPo: Probabilistic Unsupervised Laplacian Pyramid Registration
Jul 15, 2024



Abstract:Deformable image registration is fundamental to many medical imaging applications. Registration is an inherently ambiguous task often admitting many viable solutions. While neural network-based registration techniques enable fast and accurate registration, the majority of existing approaches are not able to estimate uncertainty. Here, we present PULPo, a method for probabilistic deformable registration capable of uncertainty quantification. PULPo probabilistically models the distribution of deformation fields on different hierarchical levels combining them using Laplacian pyramids. This allows our method to model global as well as local aspects of the deformation field. We evaluate our method on two widely used neuroimaging datasets and find that it achieves high registration performance as well as substantially better calibrated uncertainty quantification compared to the current state-of-the-art.
Subgroup-Specific Risk-Controlled Dose Estimation in Radiotherapy
Jul 11, 2024



Abstract:Cancer remains a leading cause of death, highlighting the importance of effective radiotherapy (RT). Magnetic resonance-guided linear accelerators (MR-Linacs) enable imaging during RT, allowing for inter-fraction, and perhaps even intra-fraction, adjustments of treatment plans. However, achieving this requires fast and accurate dose calculations. While Monte Carlo simulations offer accuracy, they are computationally intensive. Deep learning frameworks show promise, yet lack uncertainty quantification crucial for high-risk applications like RT. Risk-controlling prediction sets (RCPS) offer model-agnostic uncertainty quantification with mathematical guarantees. However, we show that naive application of RCPS may lead to only certain subgroups such as the image background being risk-controlled. In this work, we extend RCPS to provide prediction intervals with coverage guarantees for multiple subgroups with unknown subgroup membership at test time. We evaluate our algorithm on real clinical planing volumes from five different anatomical regions and show that our novel subgroup RCPS (SG-RCPS) algorithm leads to prediction intervals that jointly control the risk for multiple subgroups. In particular, our method controls the risk of the crucial voxels along the radiation beam significantly better than conventional RCPS.
Uncertainty Estimation and Propagation in Accelerated MRI Reconstruction
Aug 04, 2023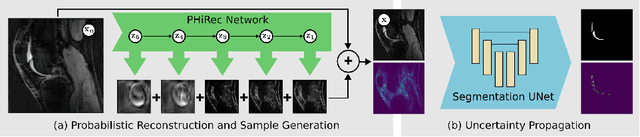



Abstract:MRI reconstruction techniques based on deep learning have led to unprecedented reconstruction quality especially in highly accelerated settings. However, deep learning techniques are also known to fail unexpectedly and hallucinate structures. This is particularly problematic if reconstructions are directly used for downstream tasks such as real-time treatment guidance or automated extraction of clinical paramters (e.g. via segmentation). Well-calibrated uncertainty quantification will be a key ingredient for safe use of this technology in clinical practice. In this paper we propose a novel probabilistic reconstruction technique (PHiRec) building on the idea of conditional hierarchical variational autoencoders. We demonstrate that our proposed method produces high-quality reconstructions as well as uncertainty quantification that is substantially better calibrated than several strong baselines. We furthermore demonstrate how uncertainties arising in the MR econstruction can be propagated to a downstream segmentation task, and show that PHiRec also allows well-calibrated estimation of segmentation uncertainties that originated in the MR reconstruction process.
First Steps towards a Runtime Analysis of Neuroevolution
Jul 03, 2023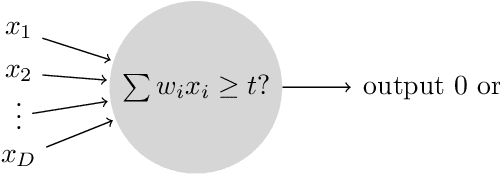
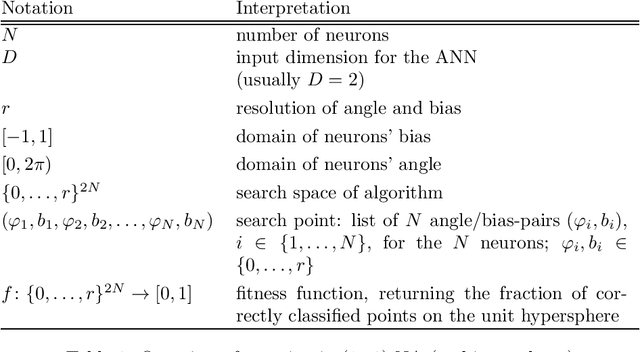

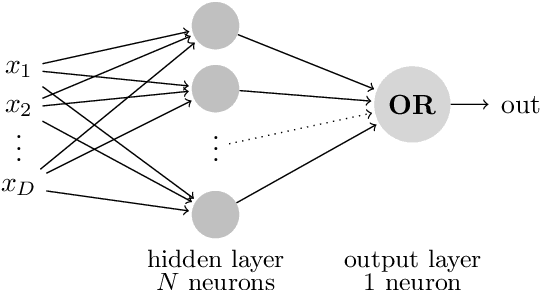
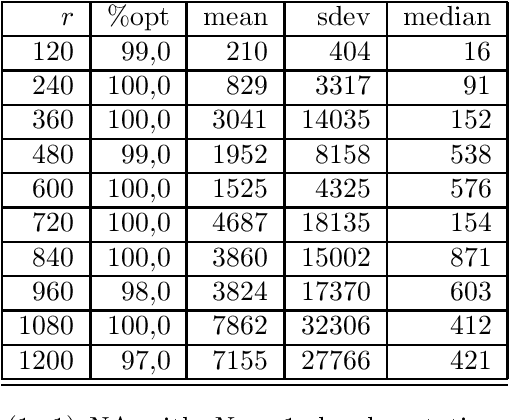
Abstract:We consider a simple setting in neuroevolution where an evolutionary algorithm optimizes the weights and activation functions of a simple artificial neural network. We then define simple example functions to be learned by the network and conduct rigorous runtime analyses for networks with a single neuron and for a more advanced structure with several neurons and two layers. Our results show that the proposed algorithm is generally efficient on two example problems designed for one neuron and efficient with at least constant probability on the example problem for a two-layer network. In particular, the so-called harmonic mutation operator choosing steps of size $j$ with probability proportional to $1/j$ turns out as a good choice for the underlying search space. However, for the case of one neuron, we also identify situations with hard-to-overcome local optima. Experimental investigations of our neuroevolutionary algorithm and a state-of-the-art CMA-ES support the theoretical findings.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge